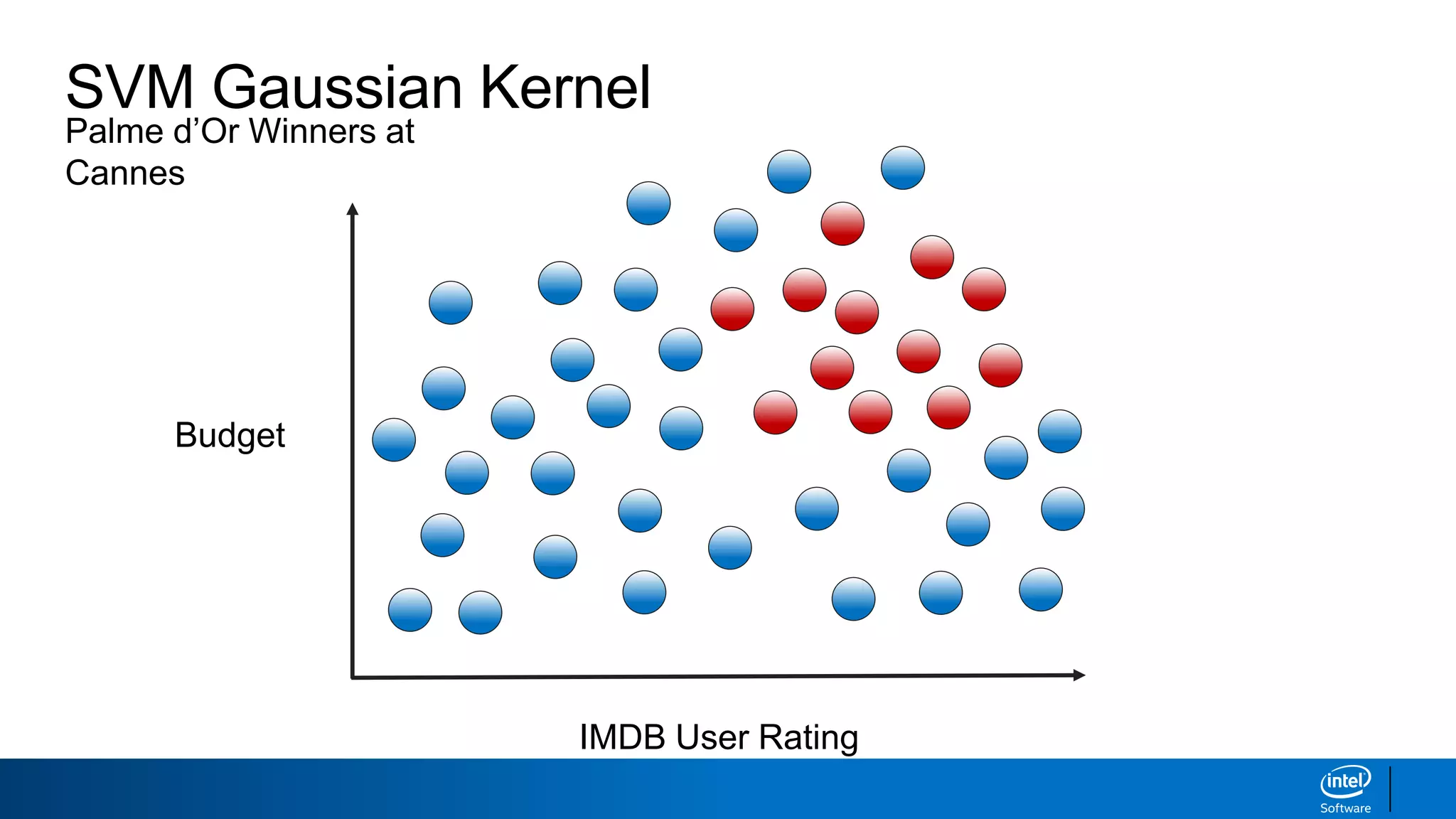
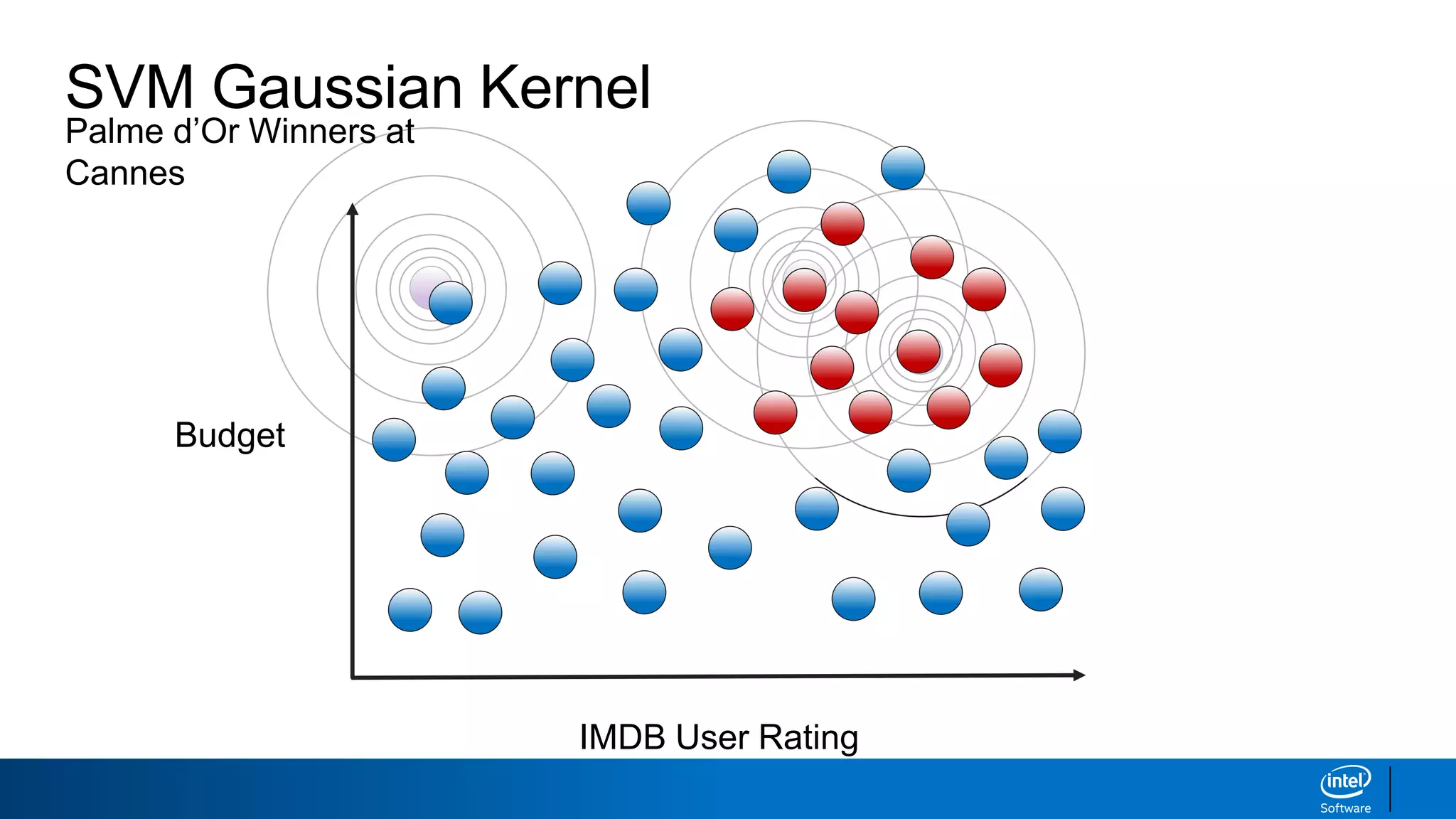
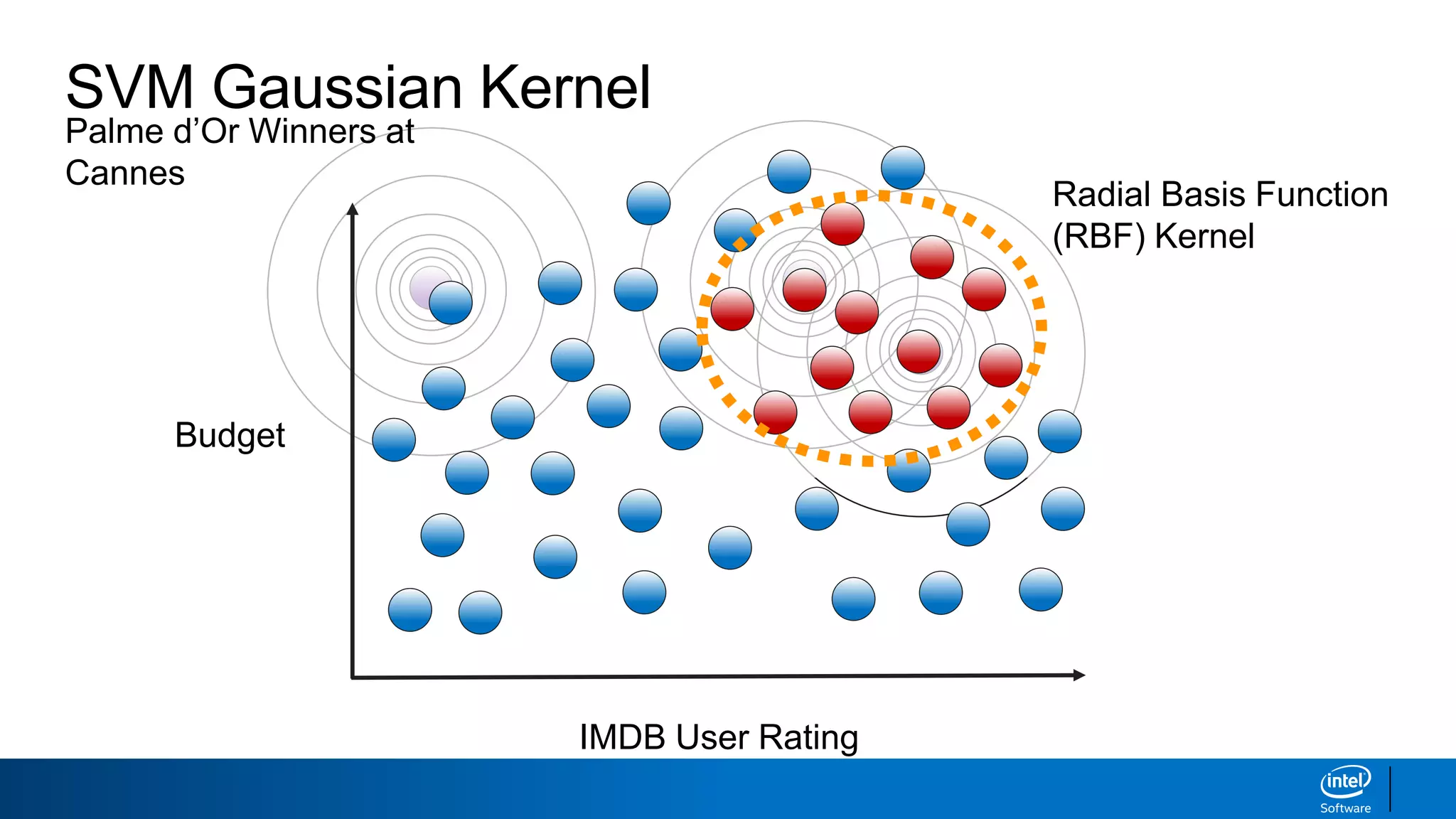
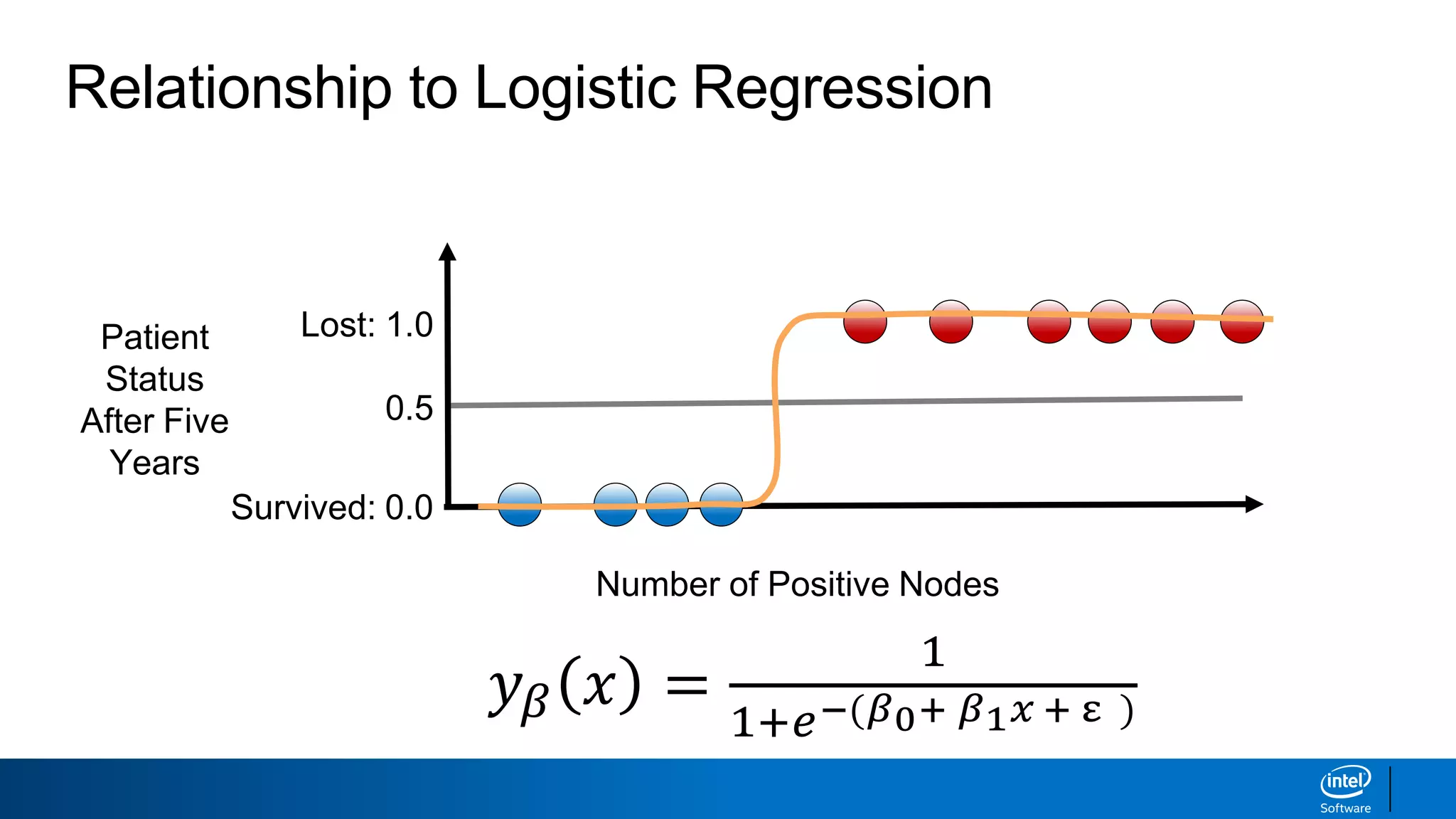
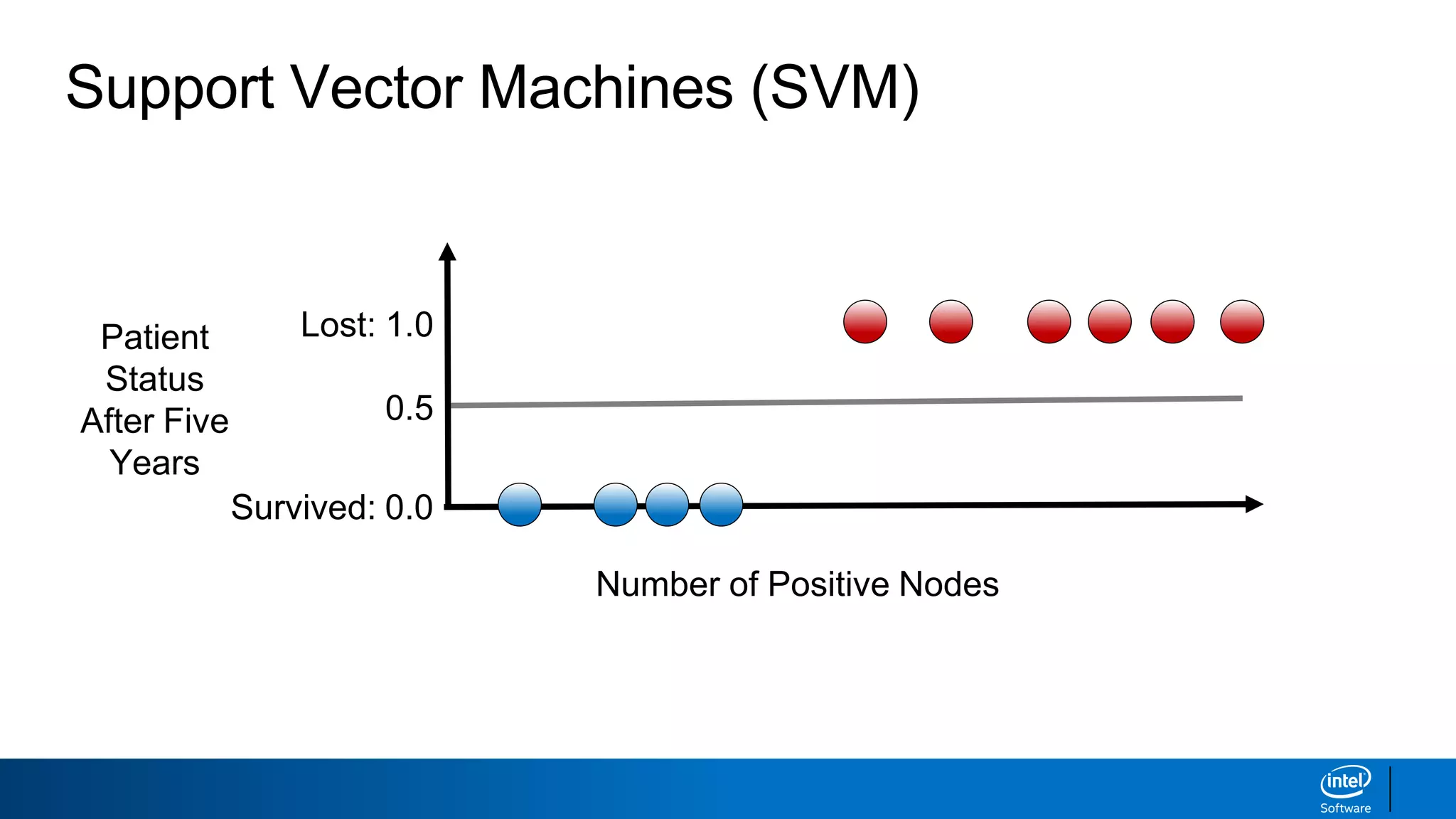
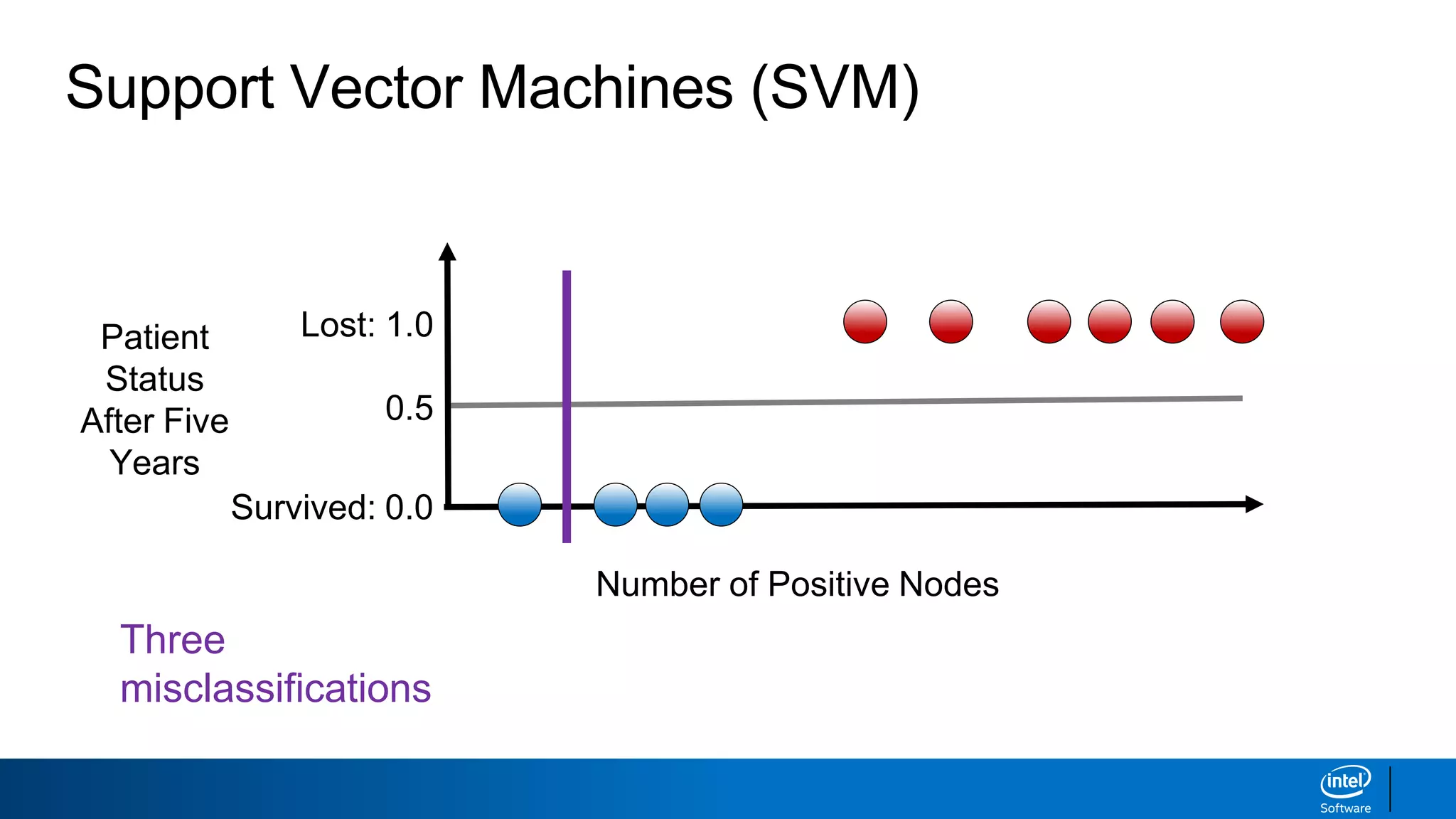
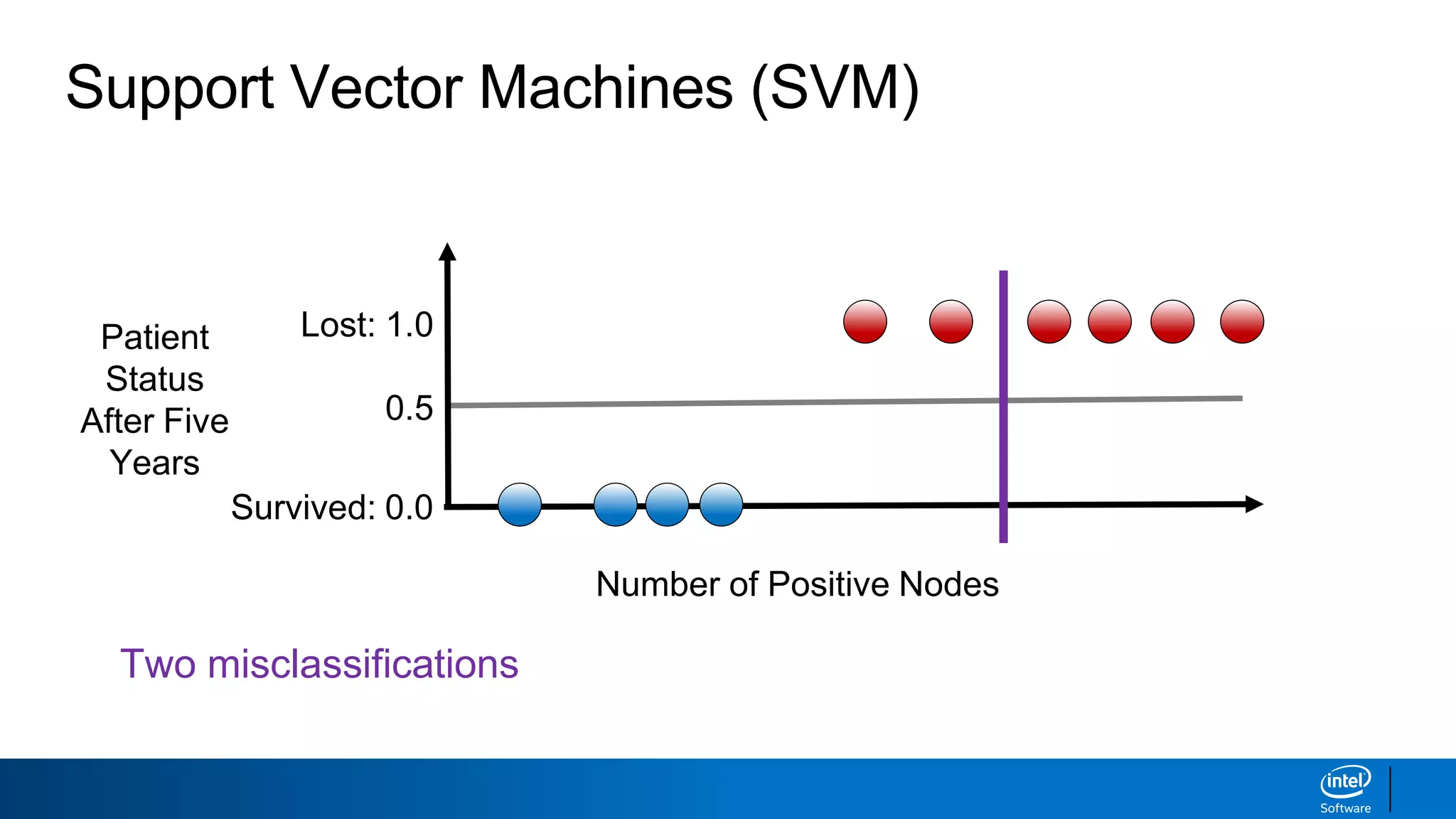
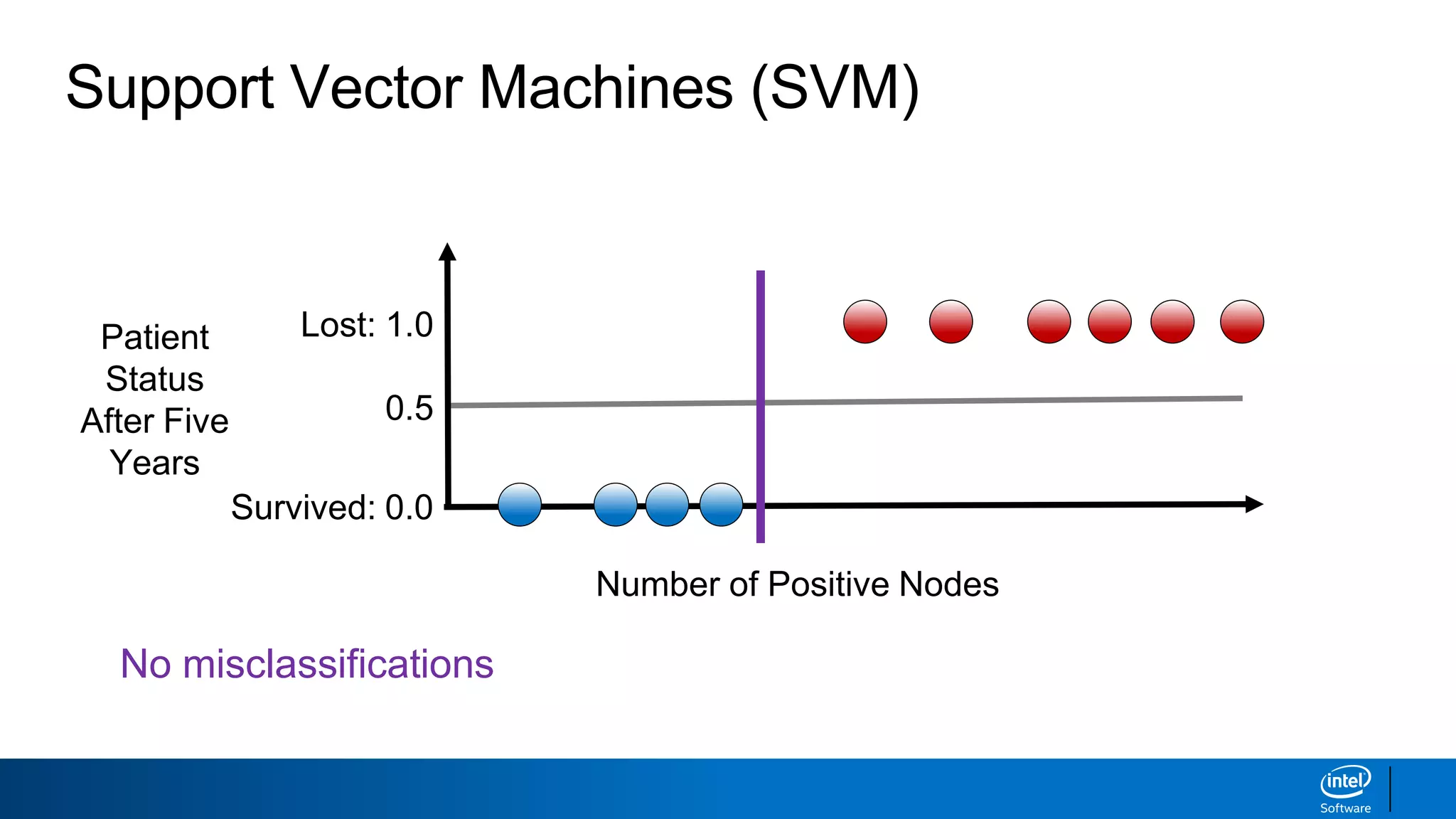
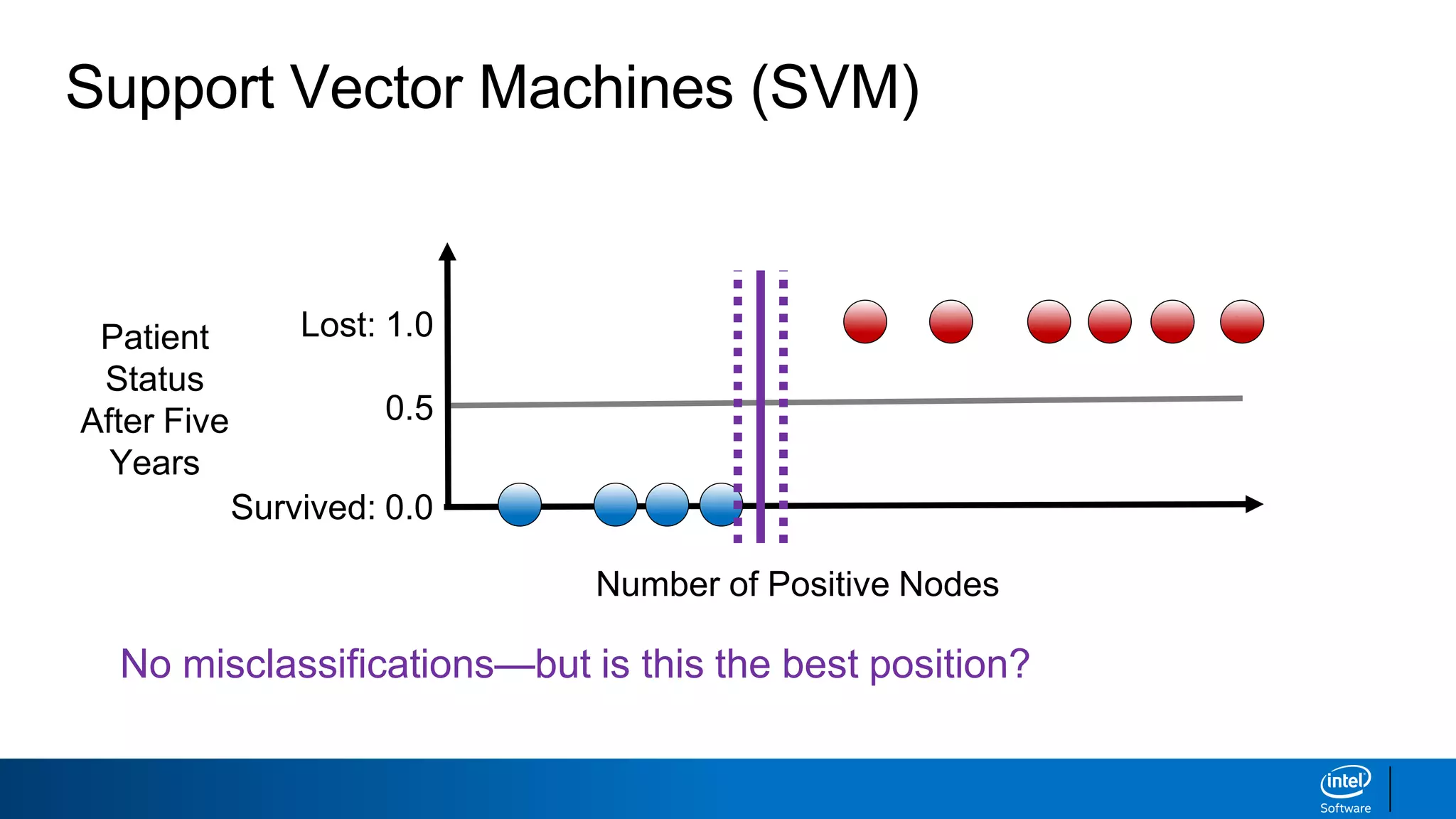
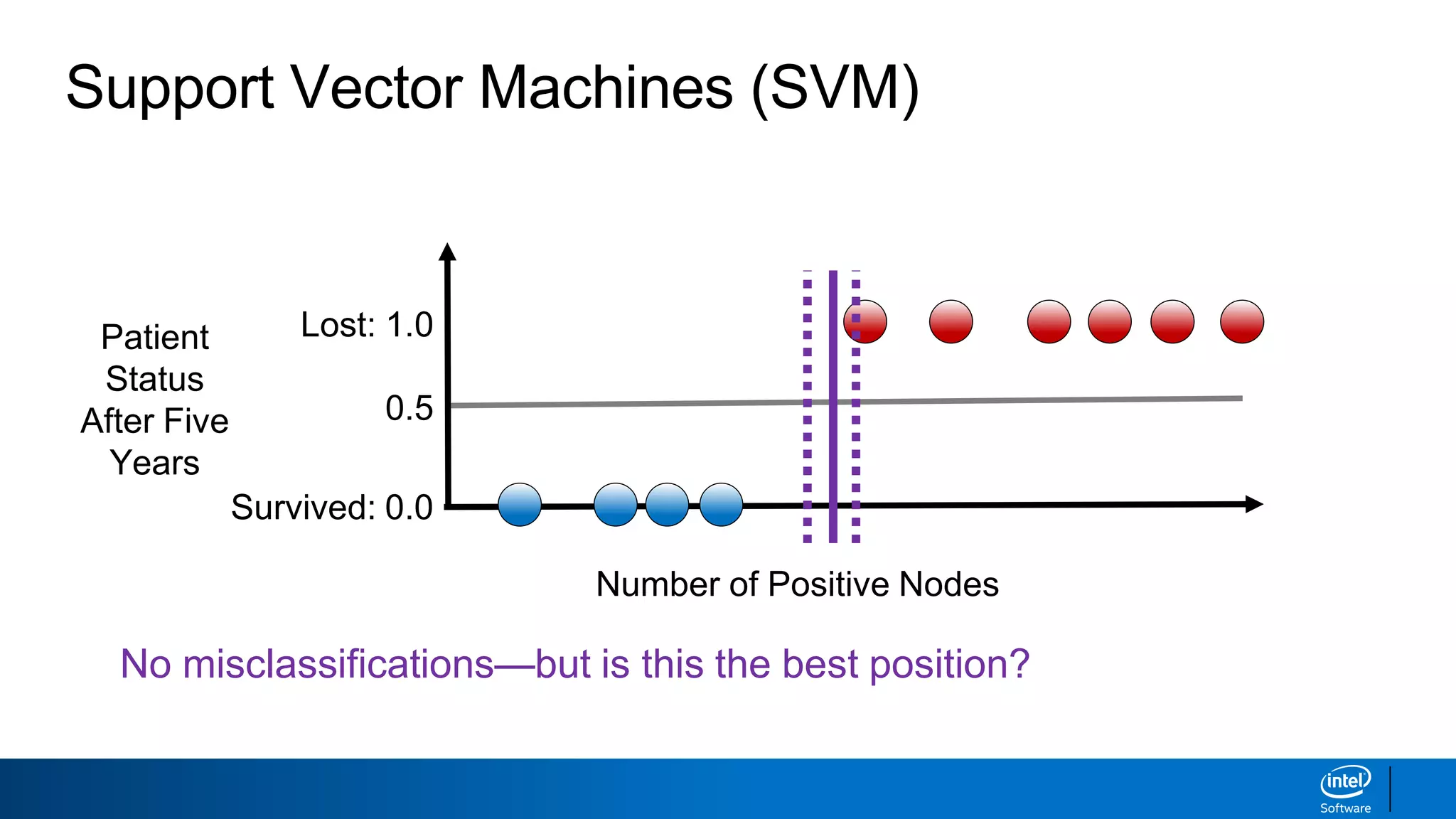
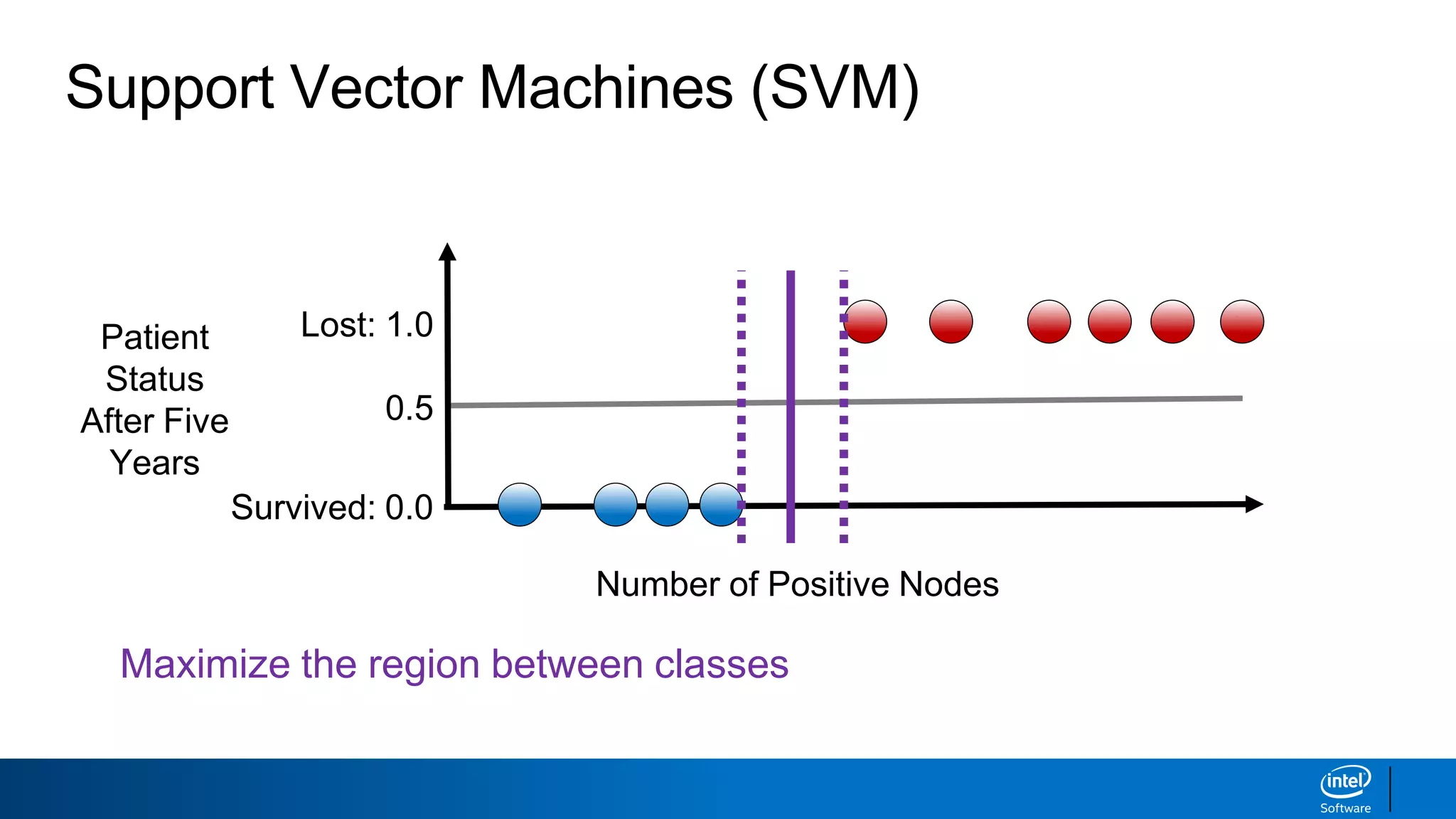
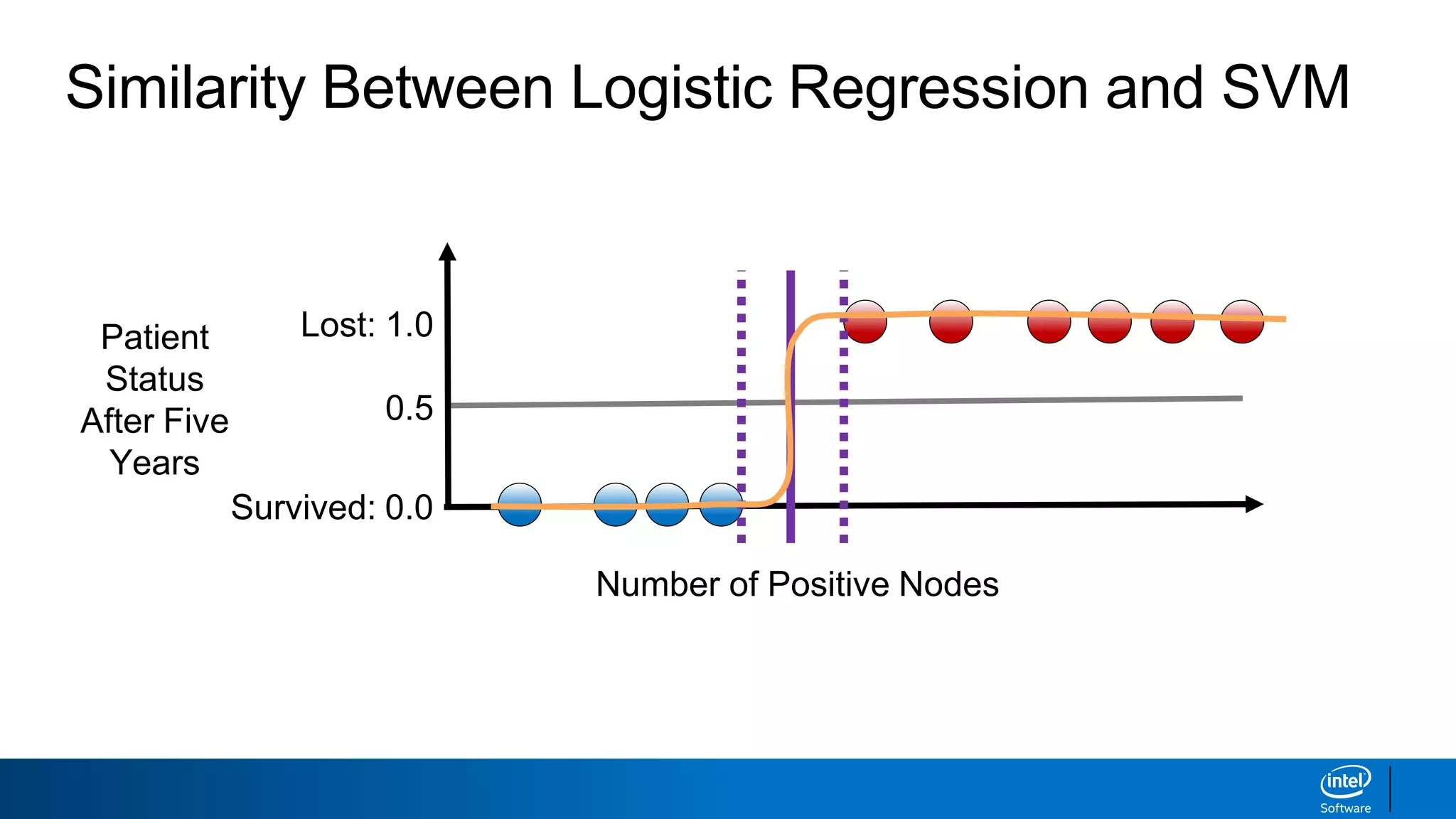
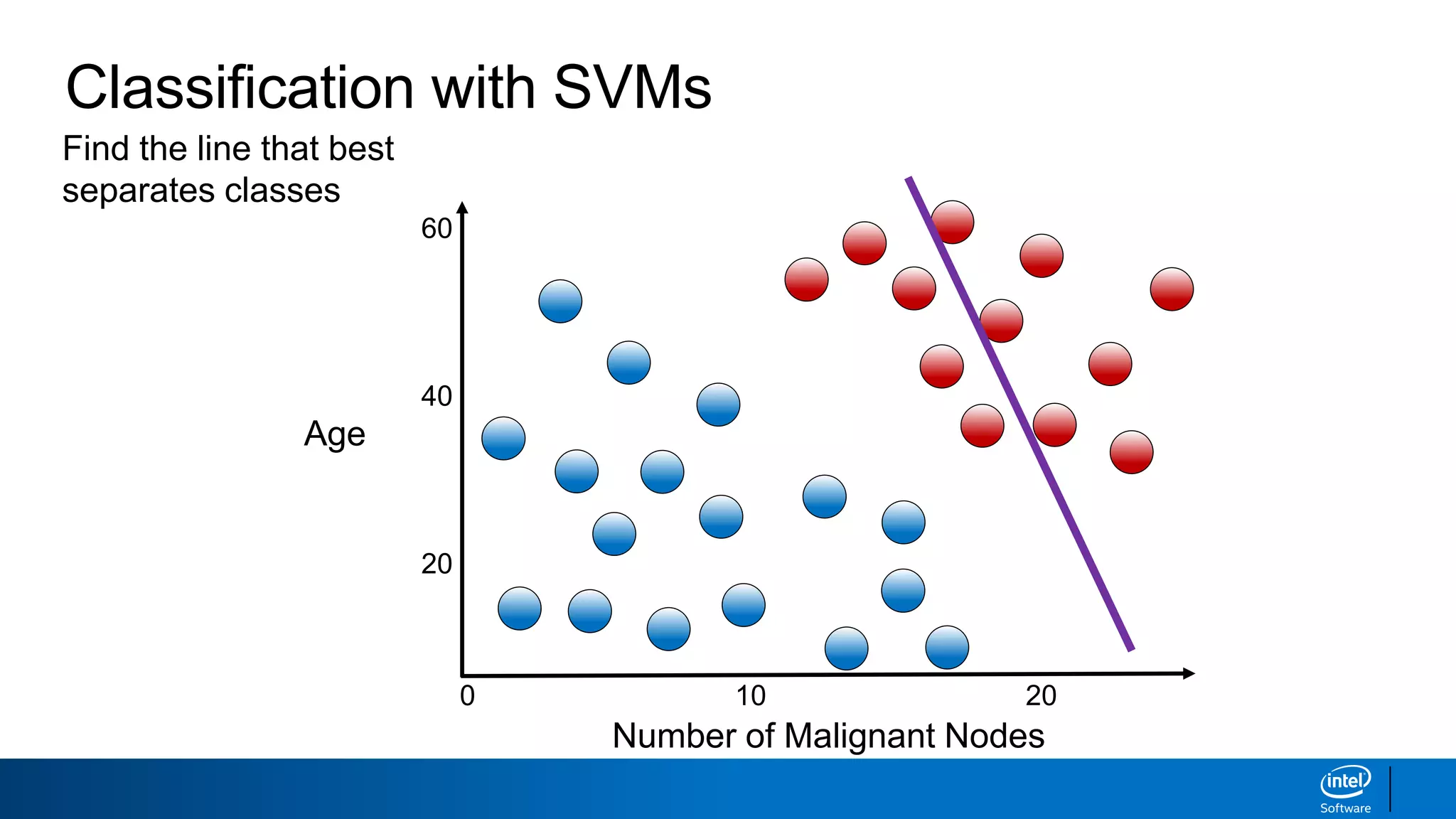
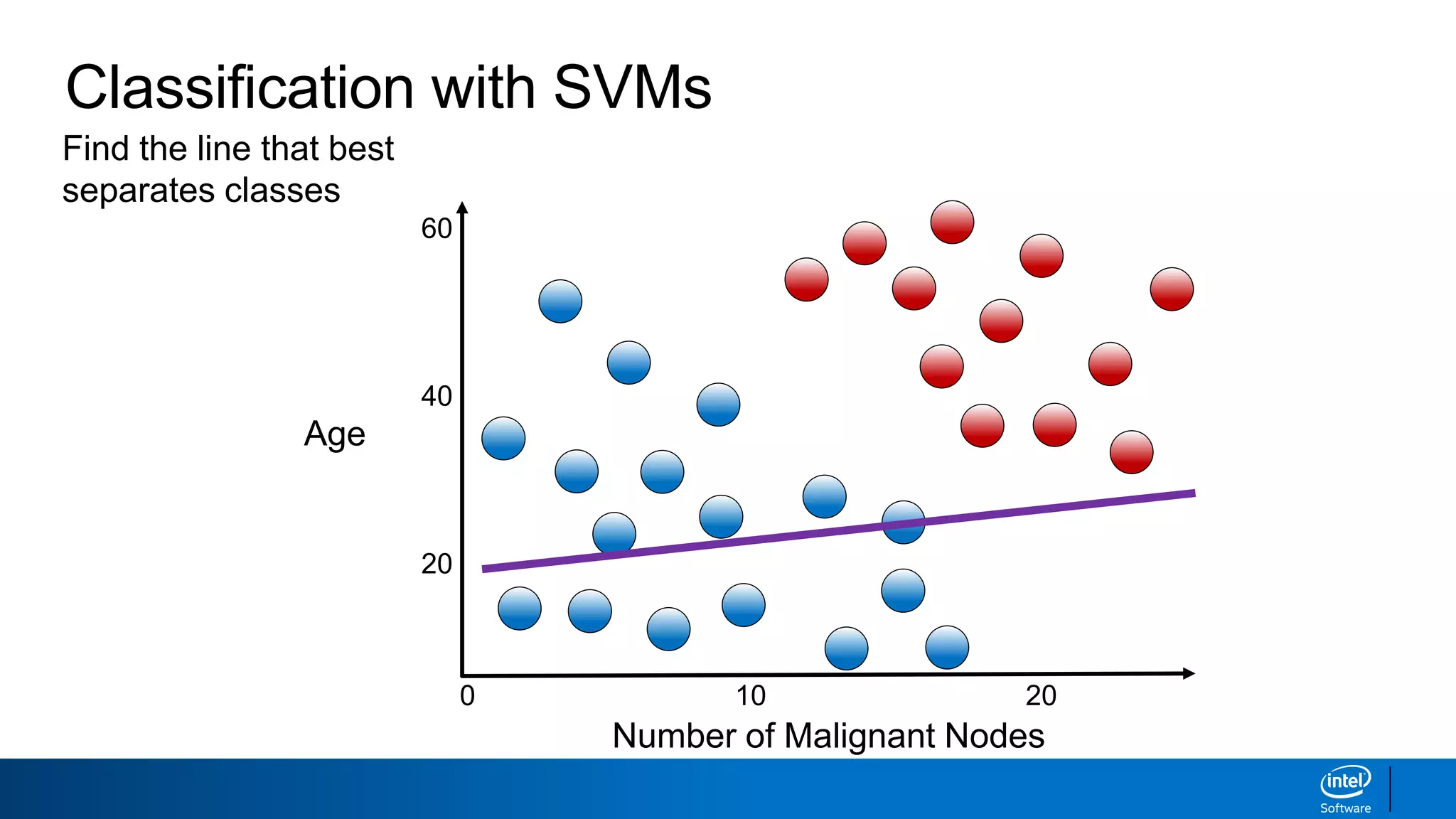
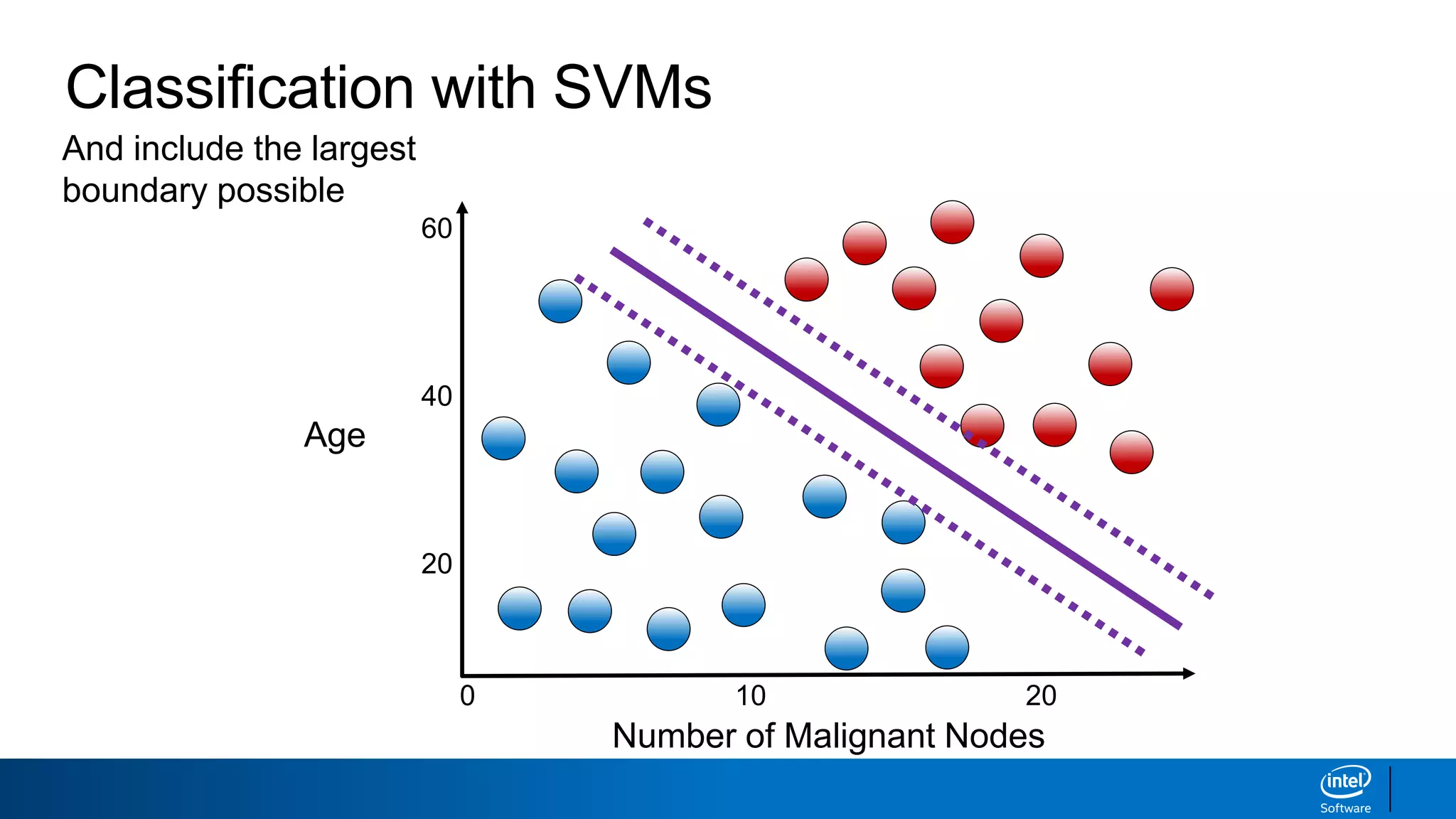
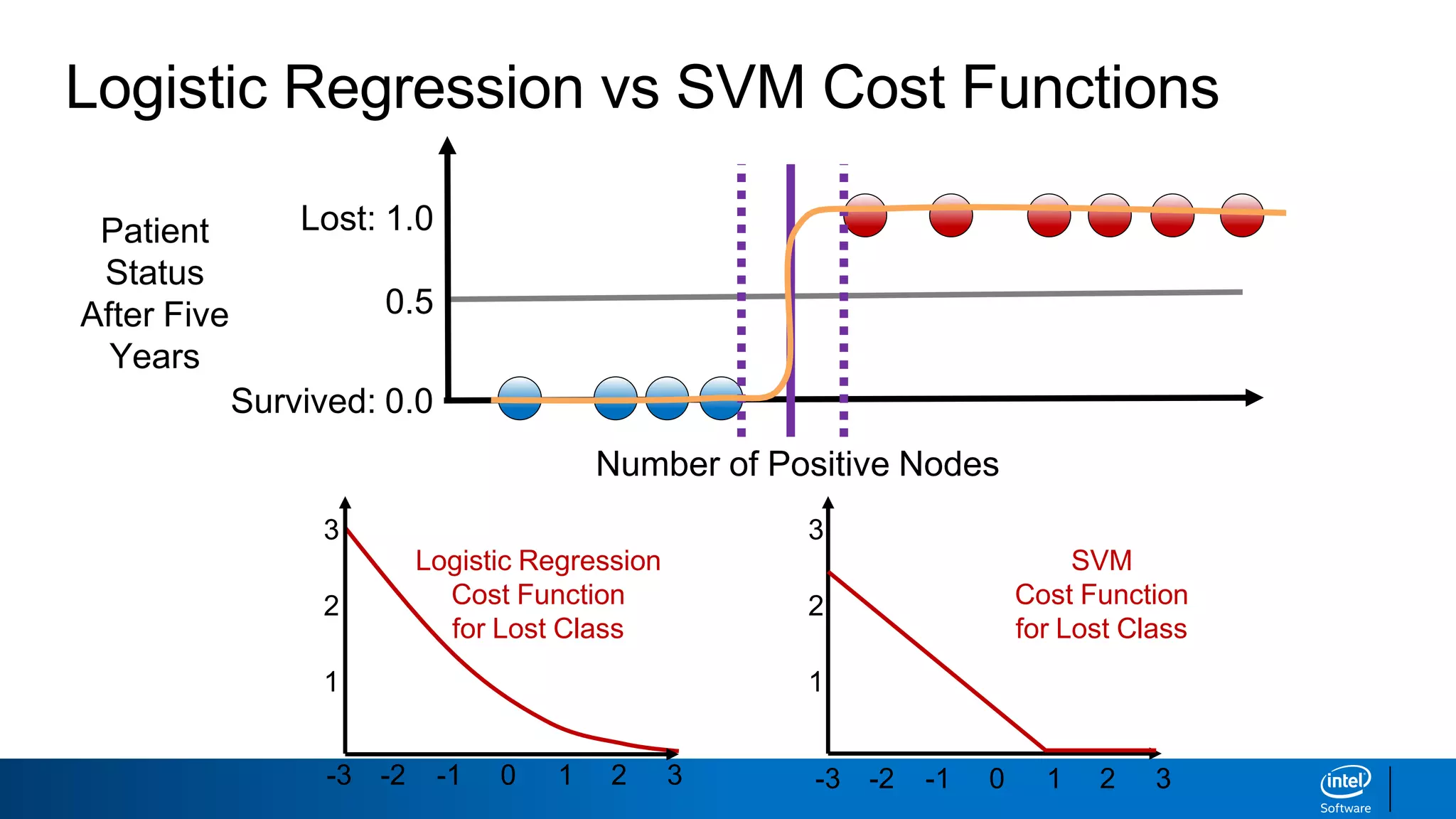
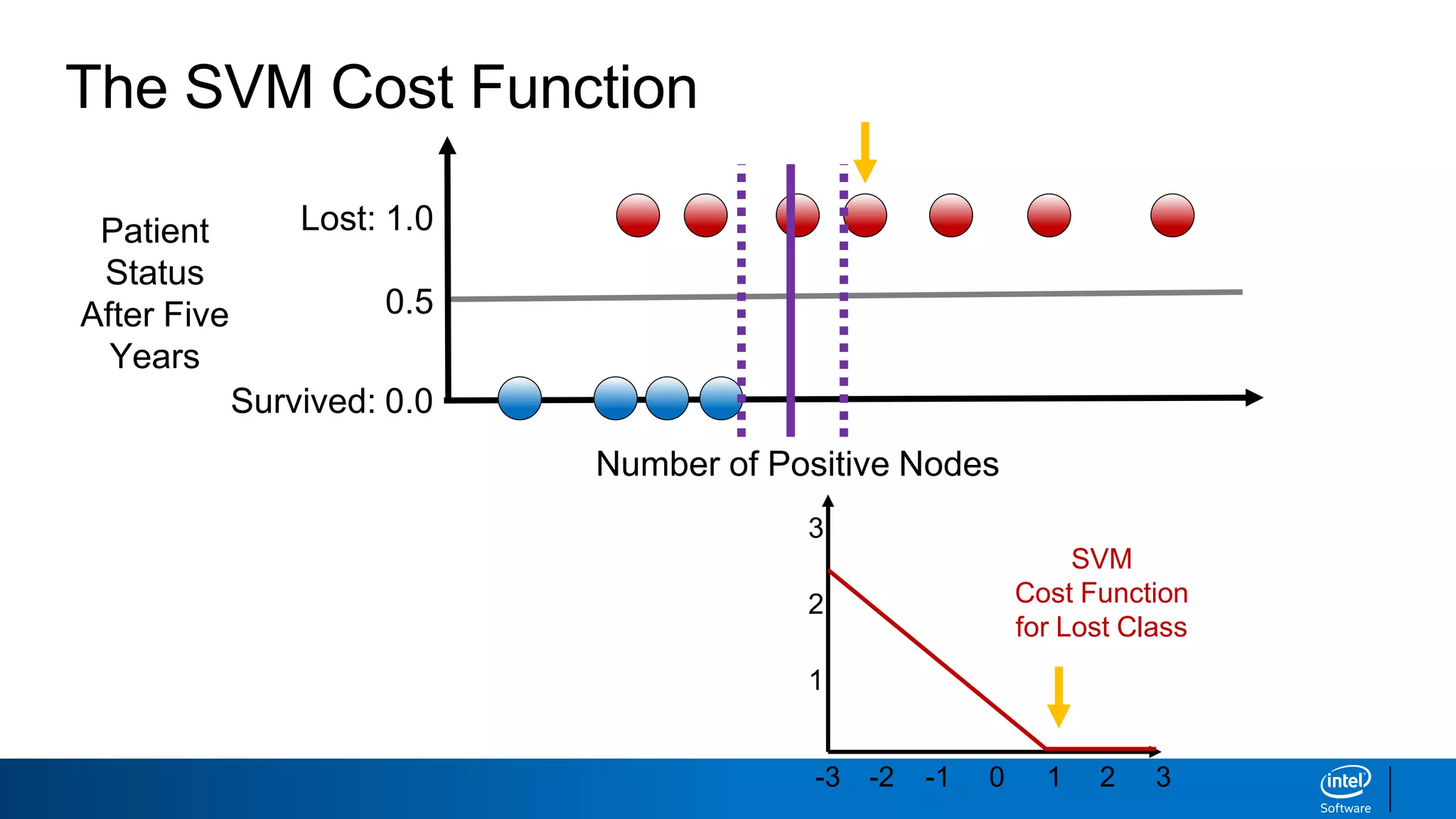
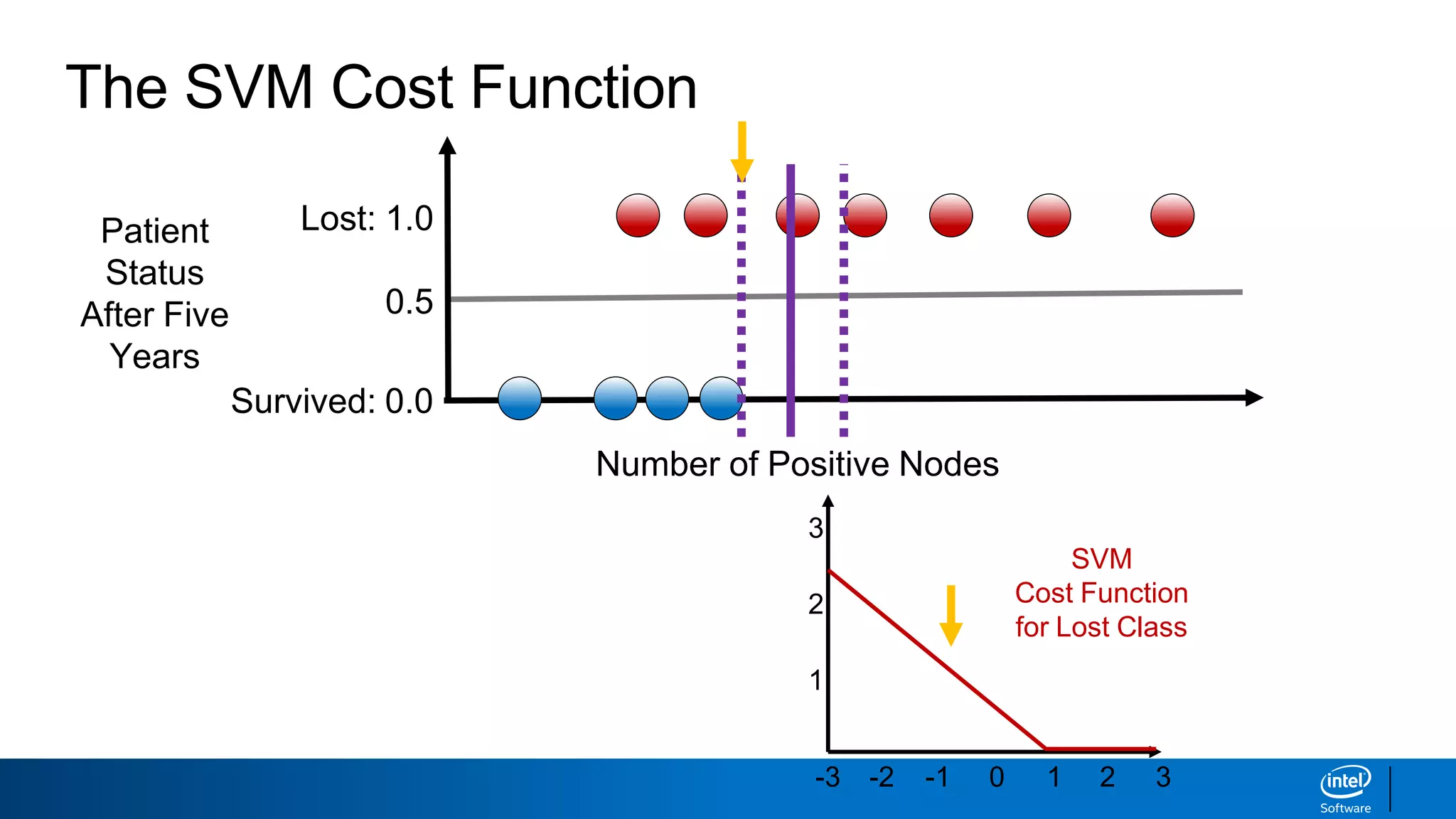
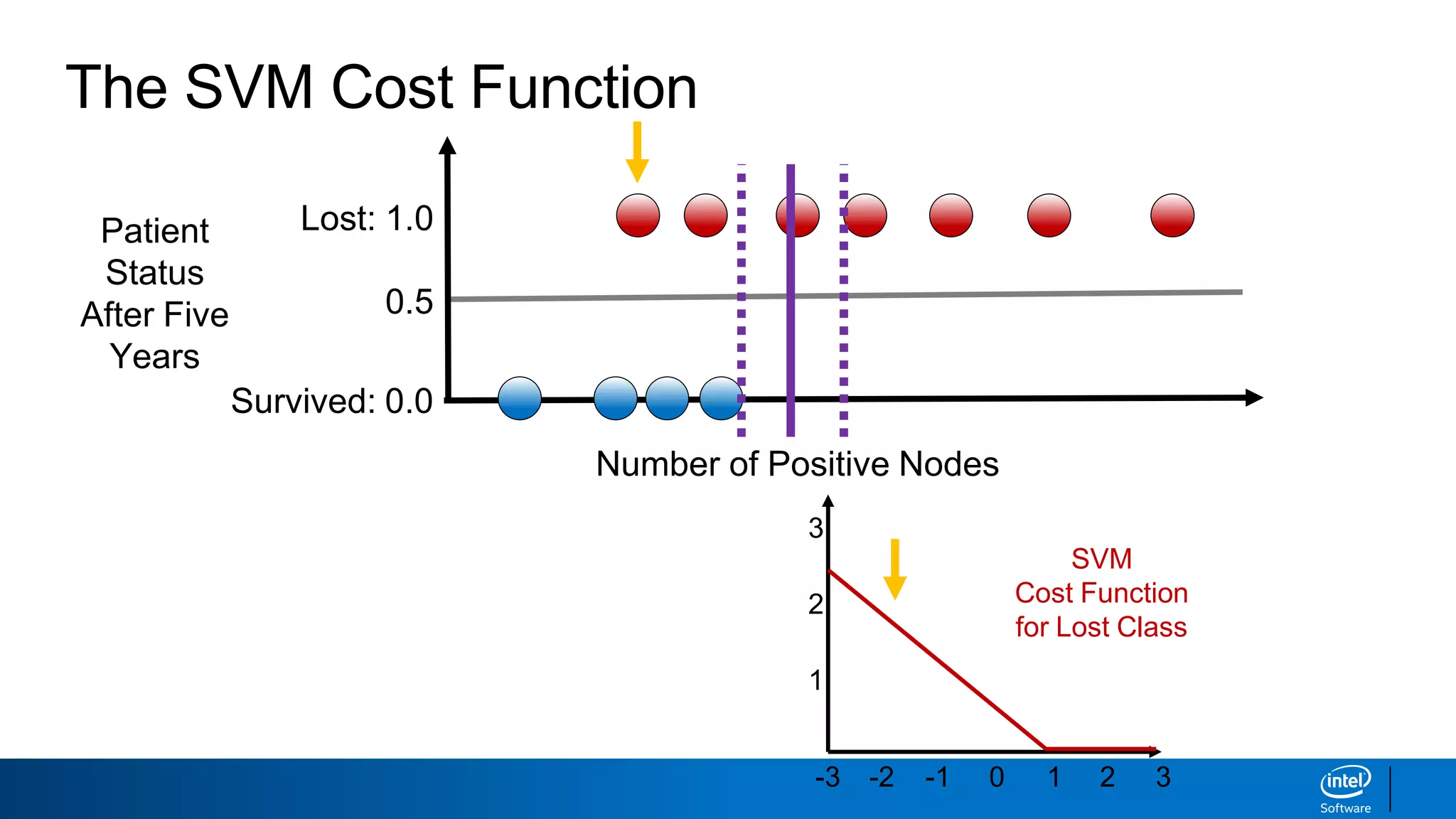
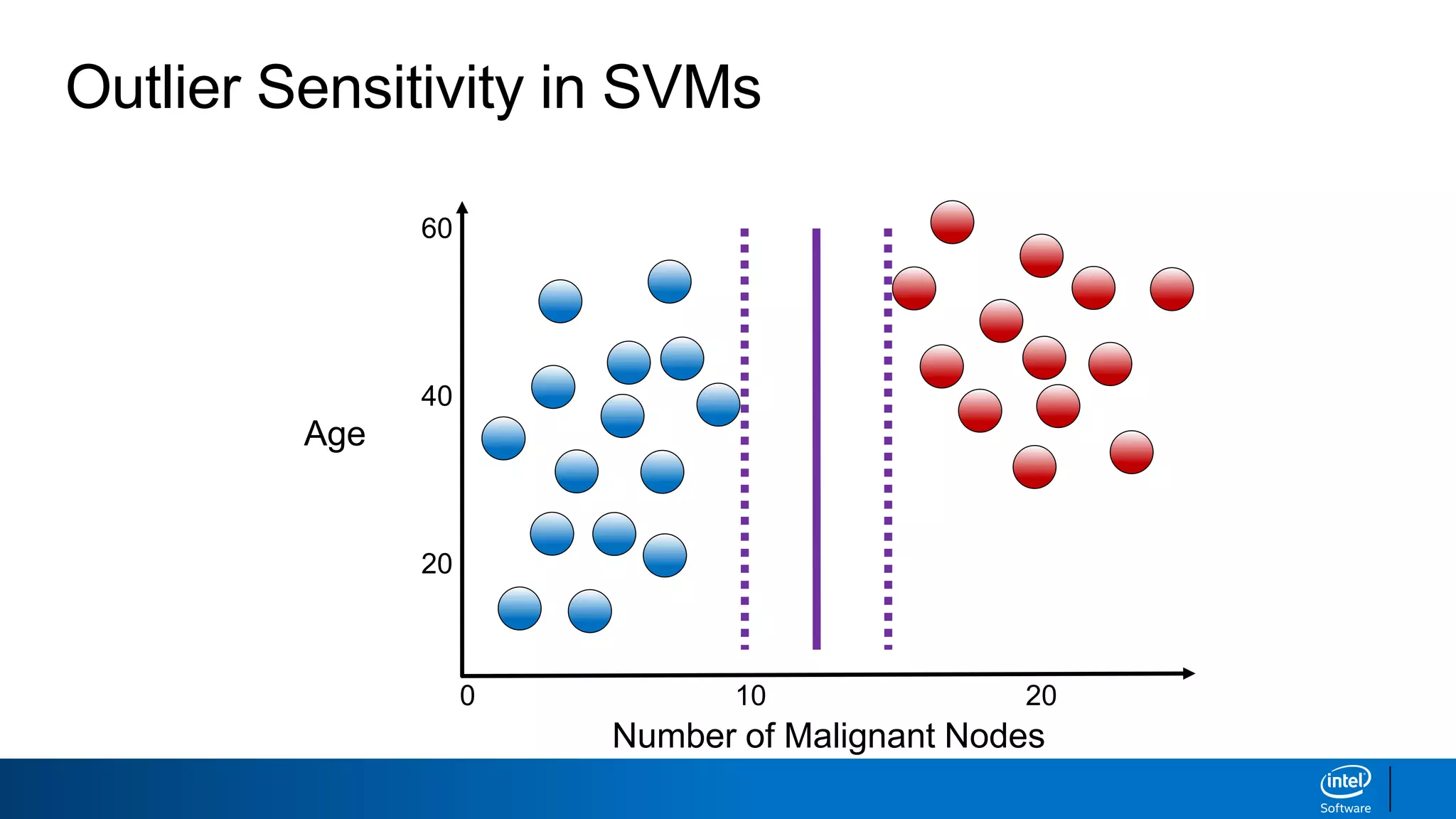
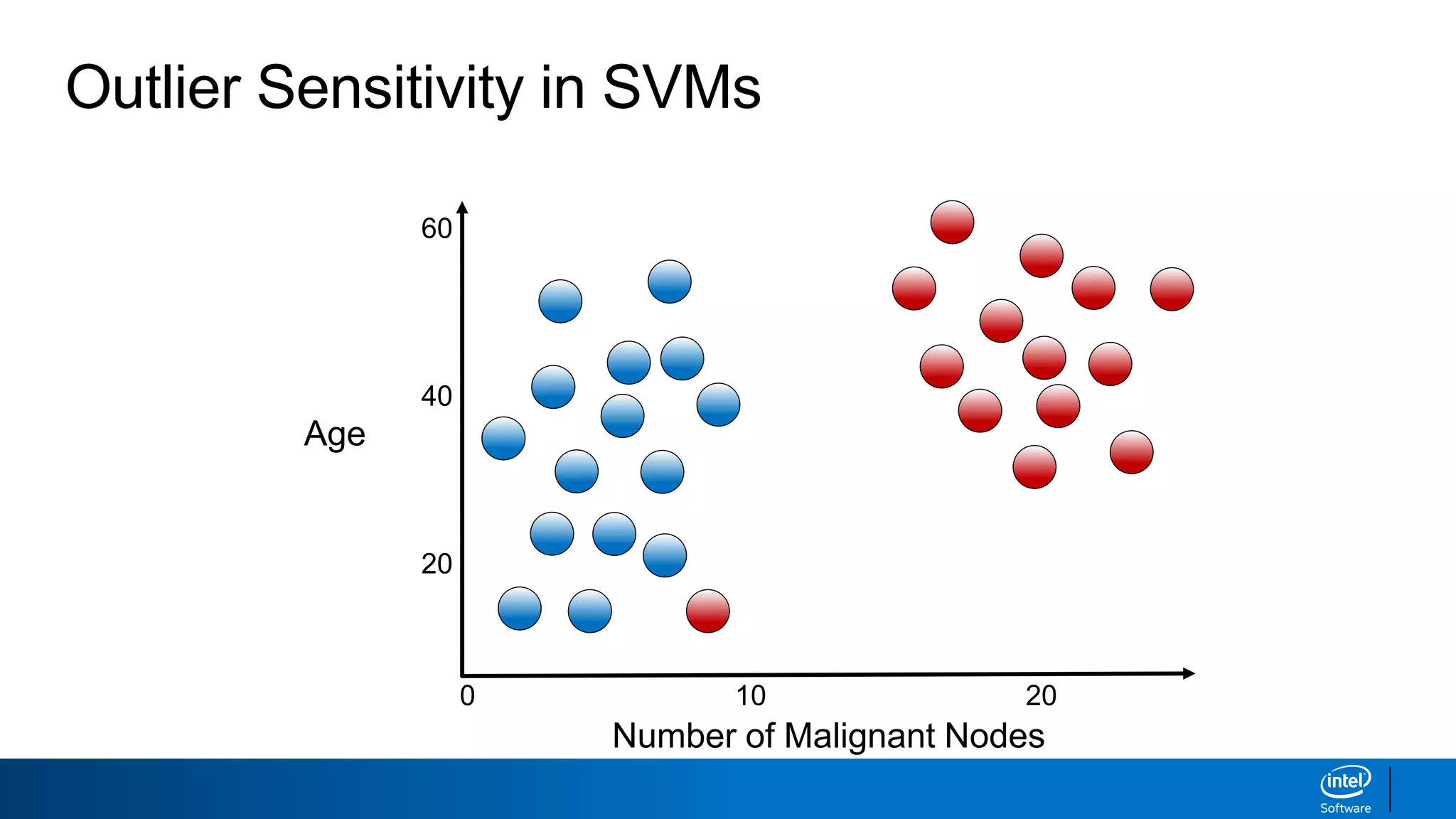
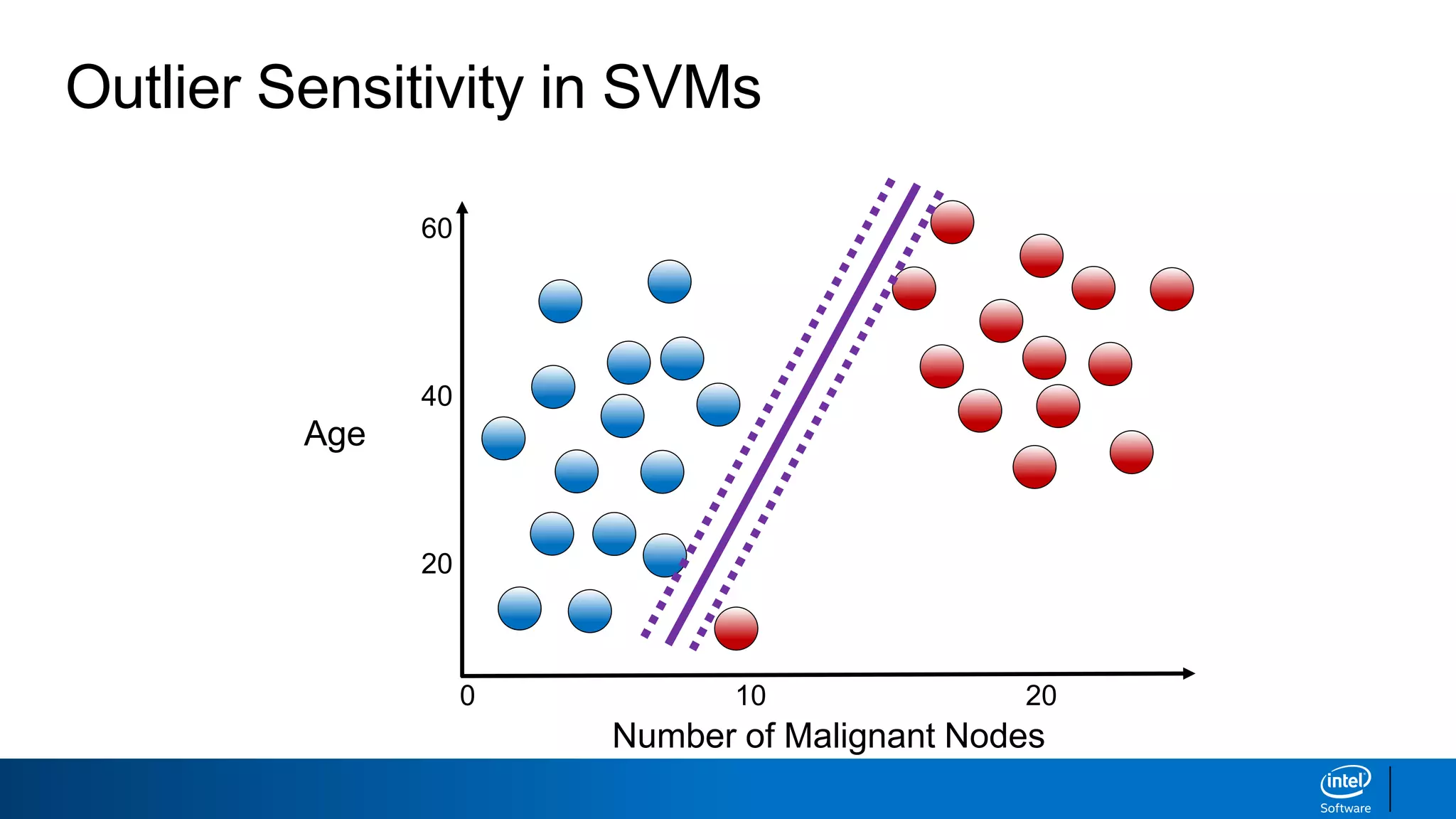
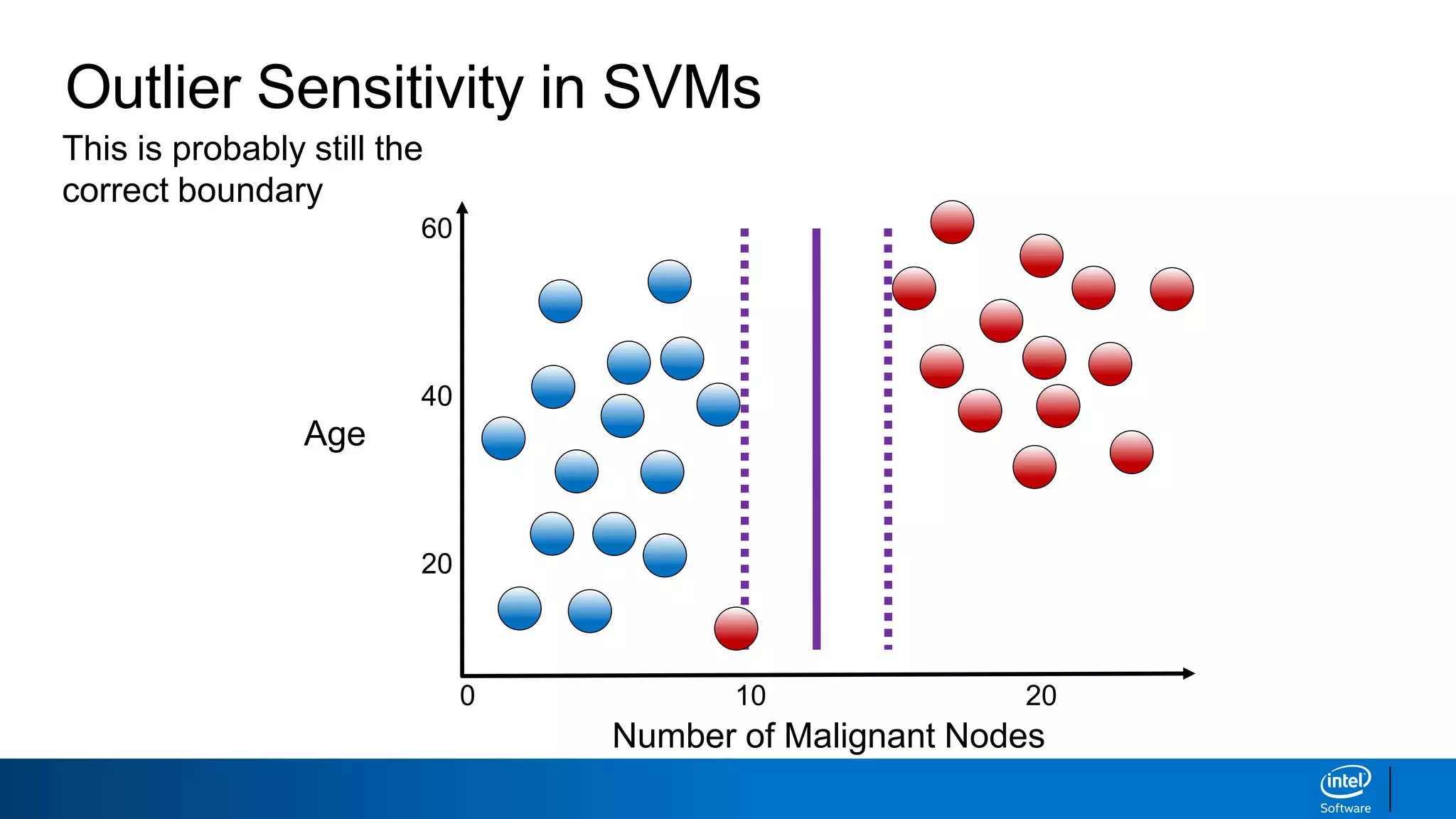
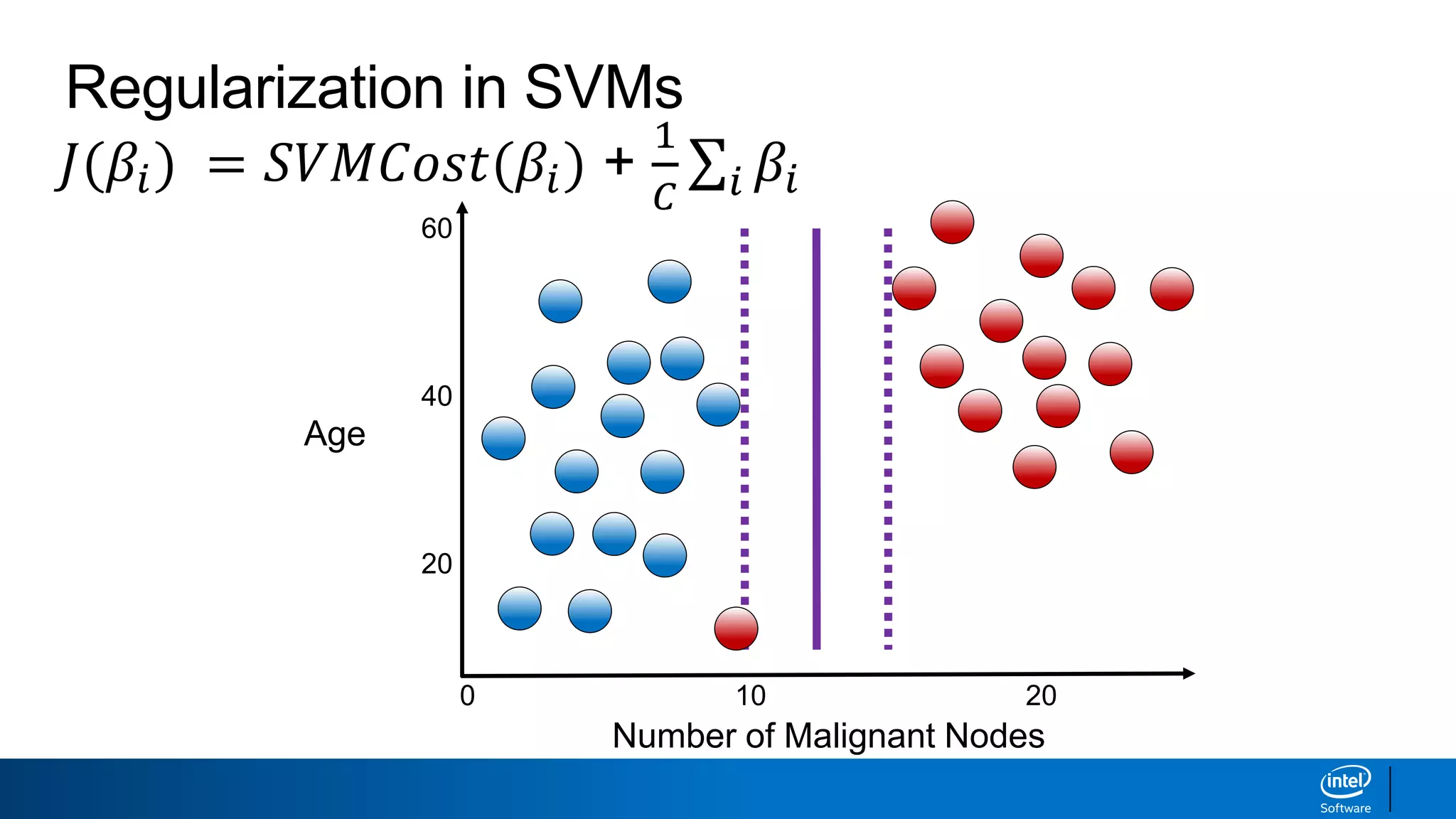
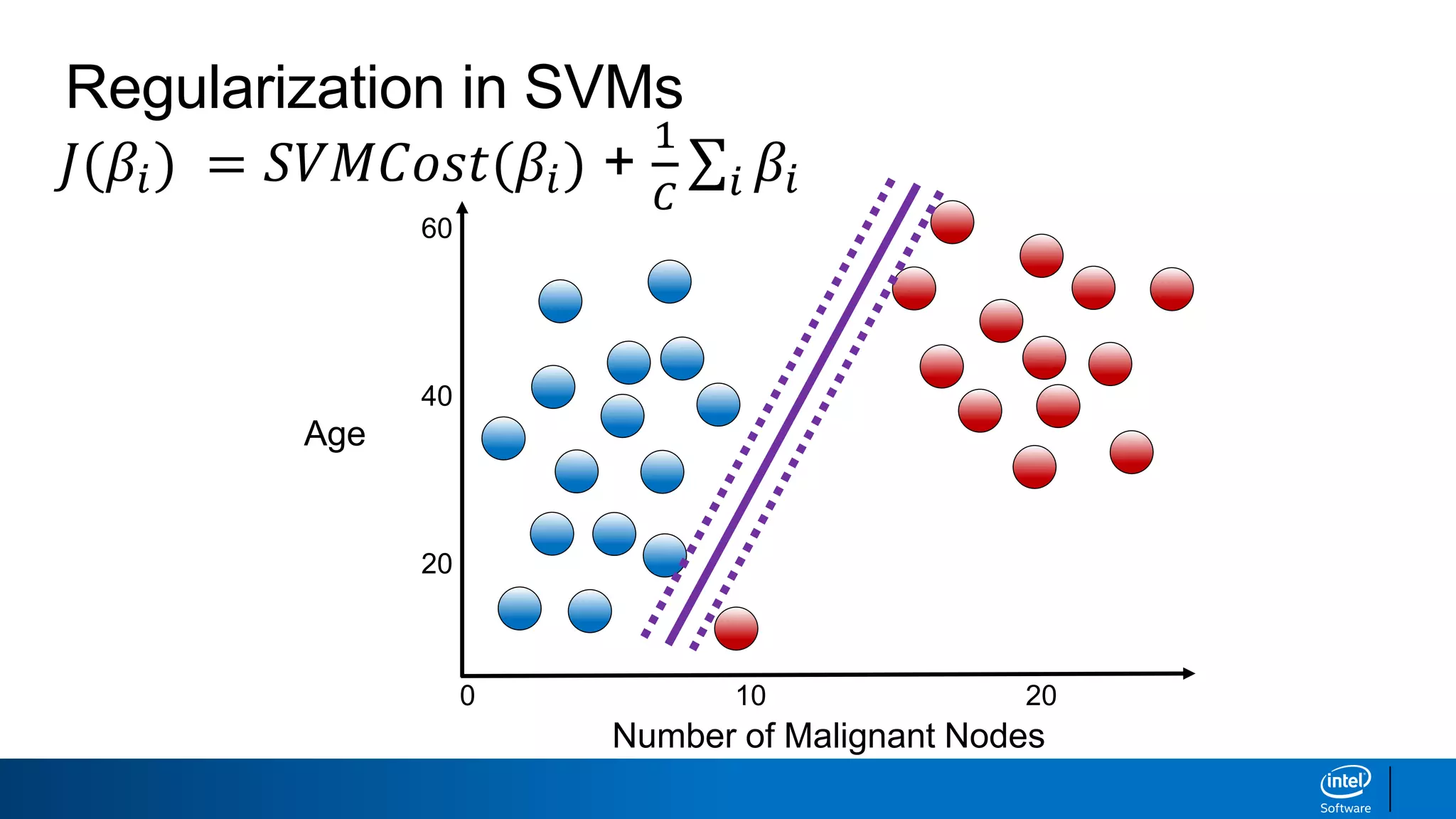
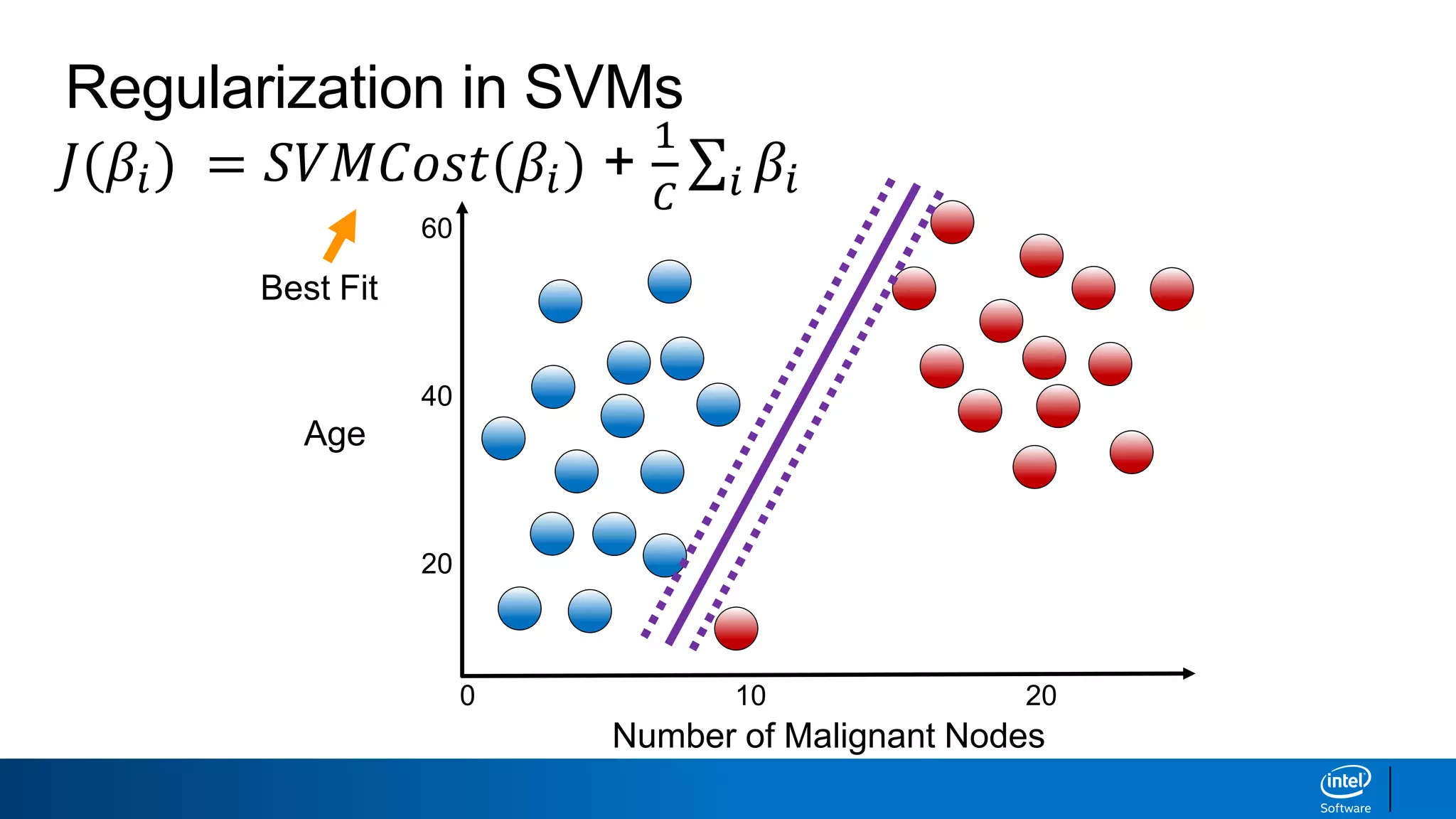
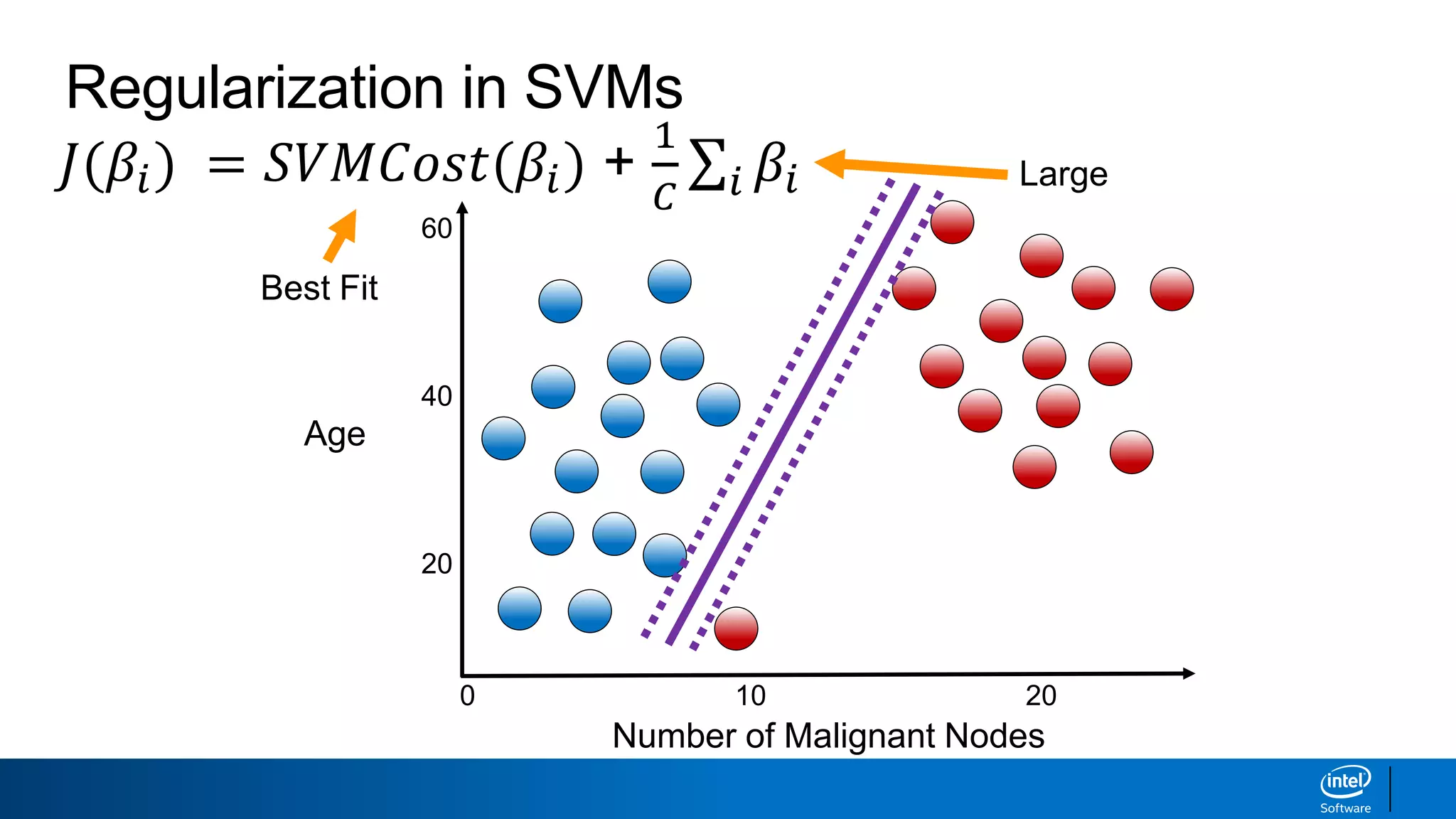
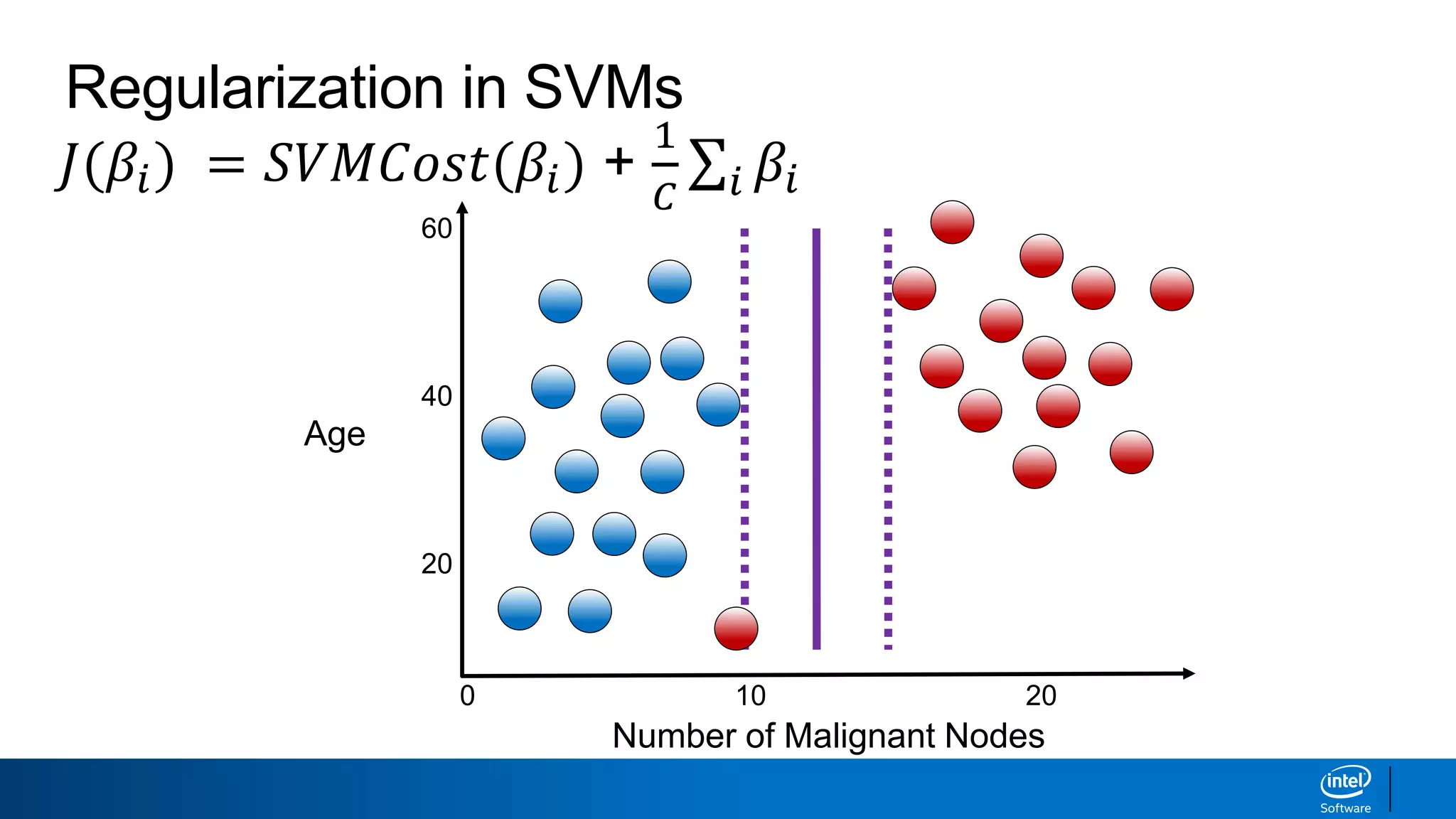
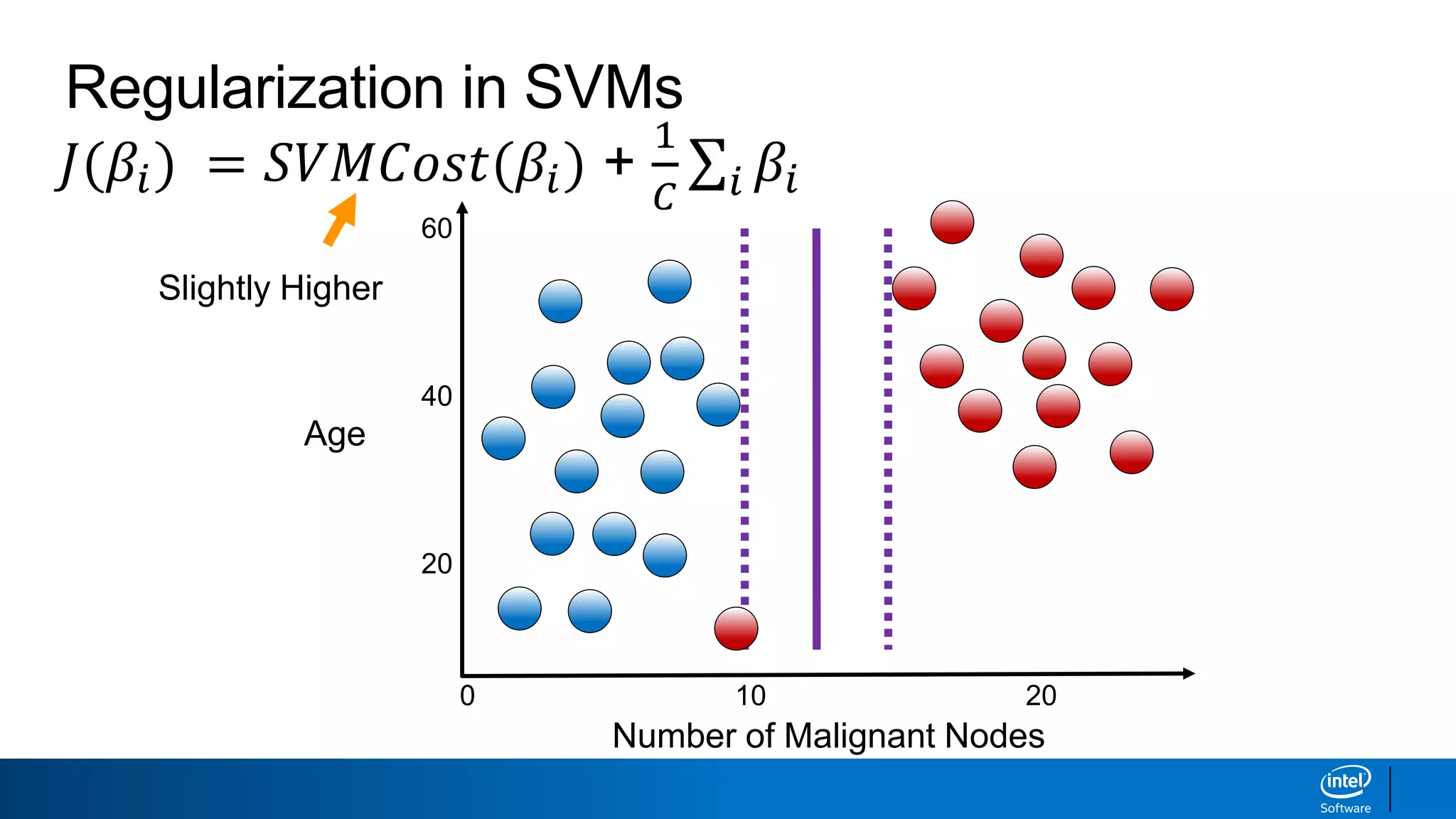
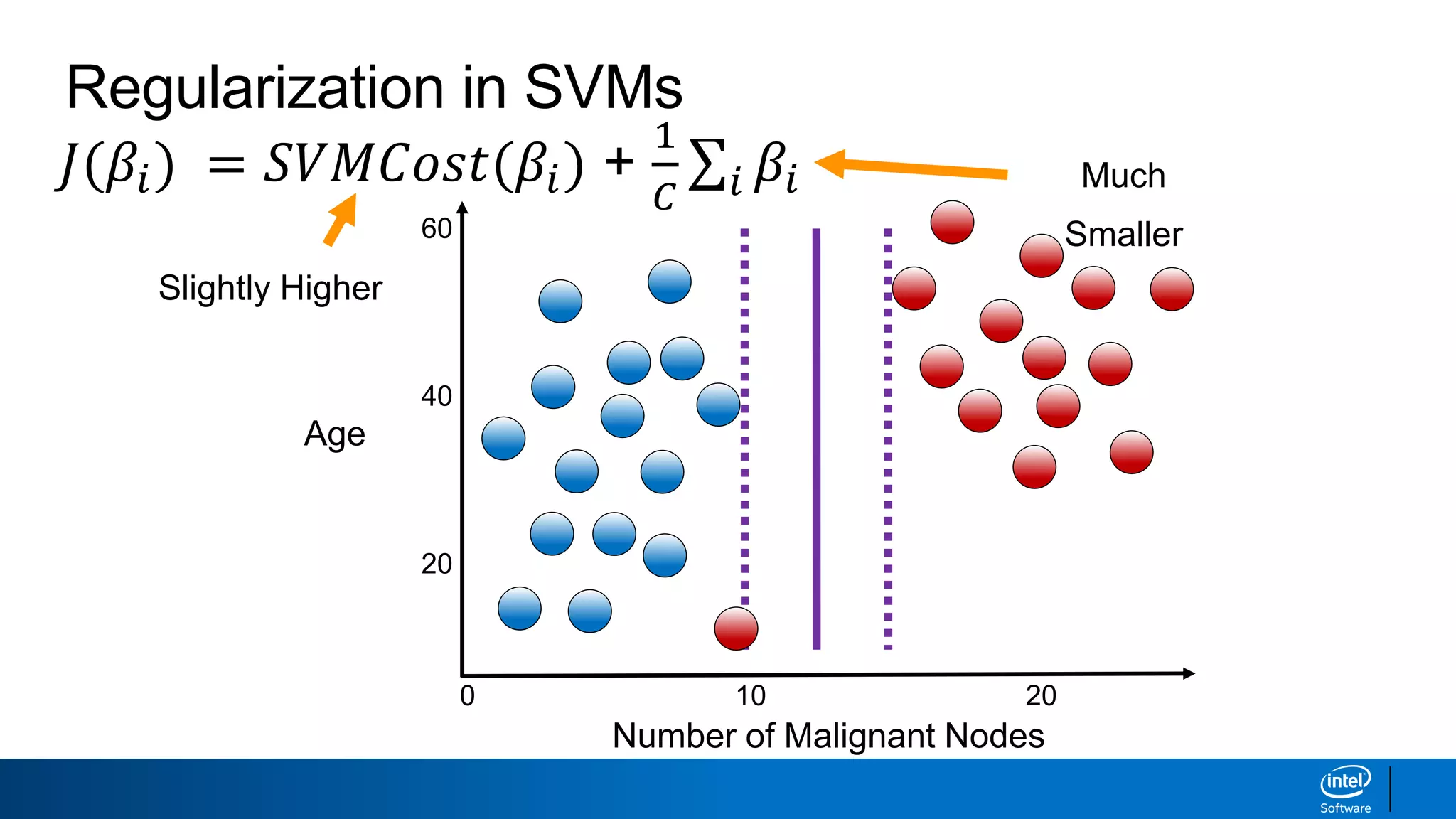
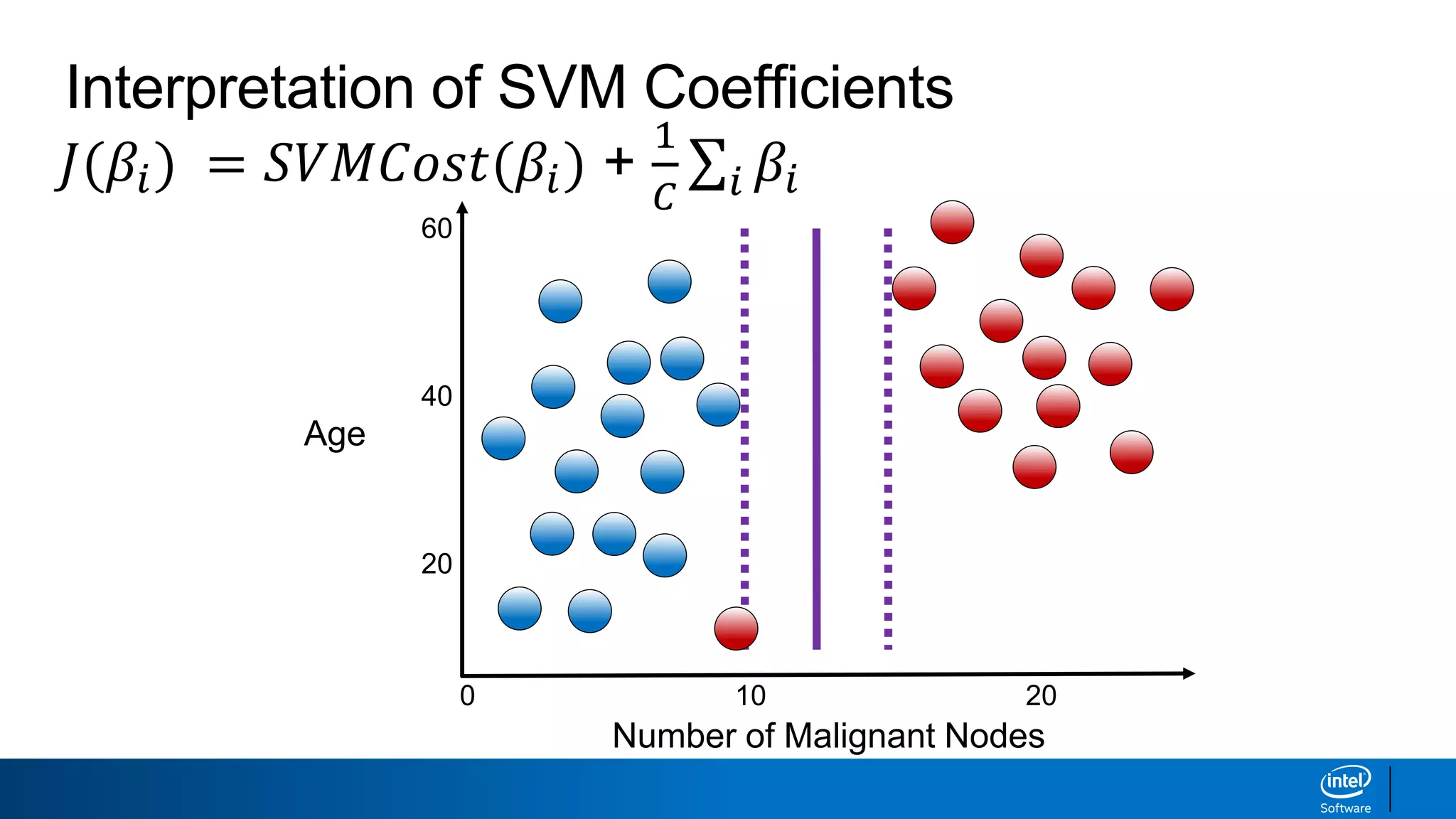
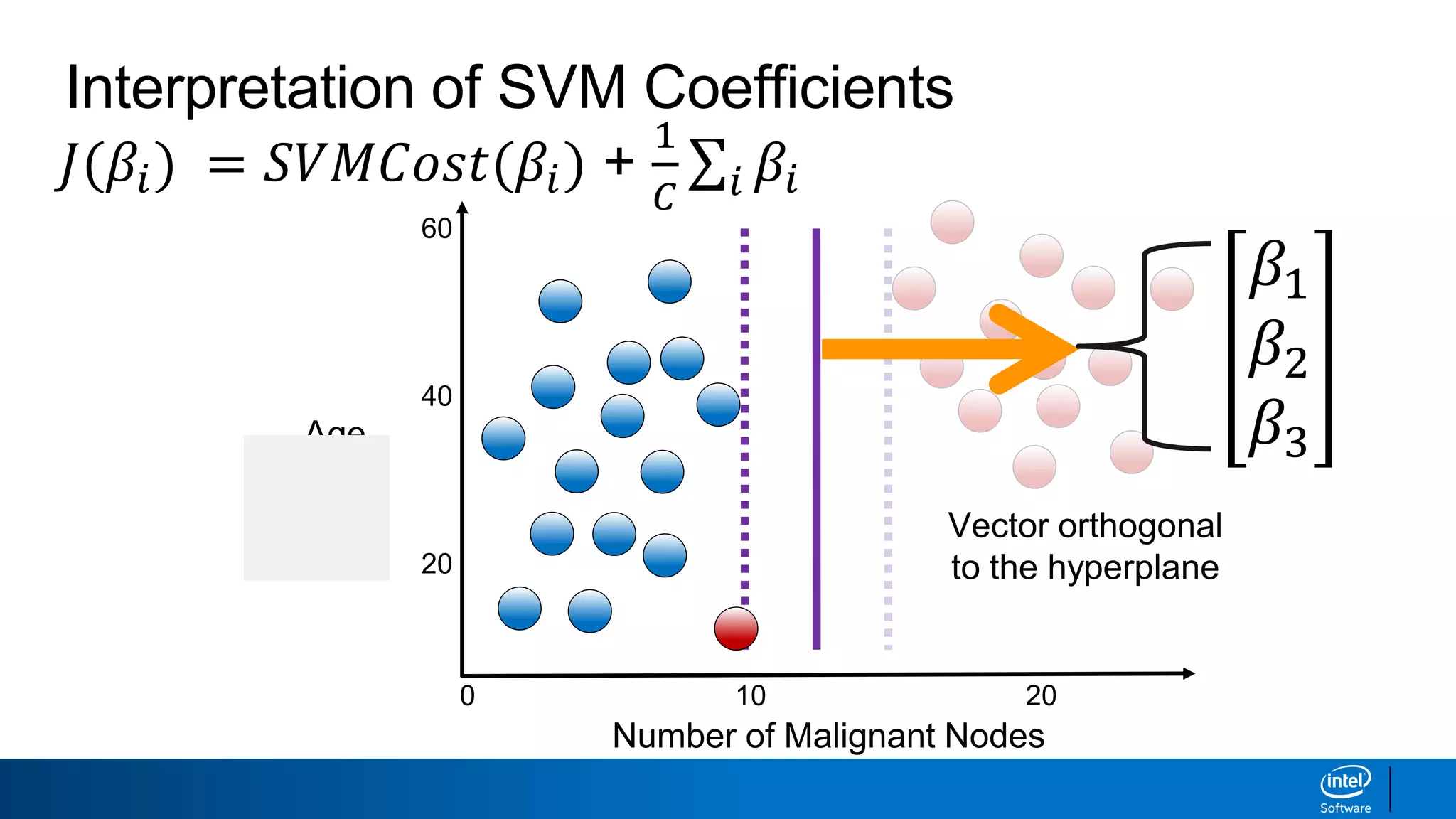
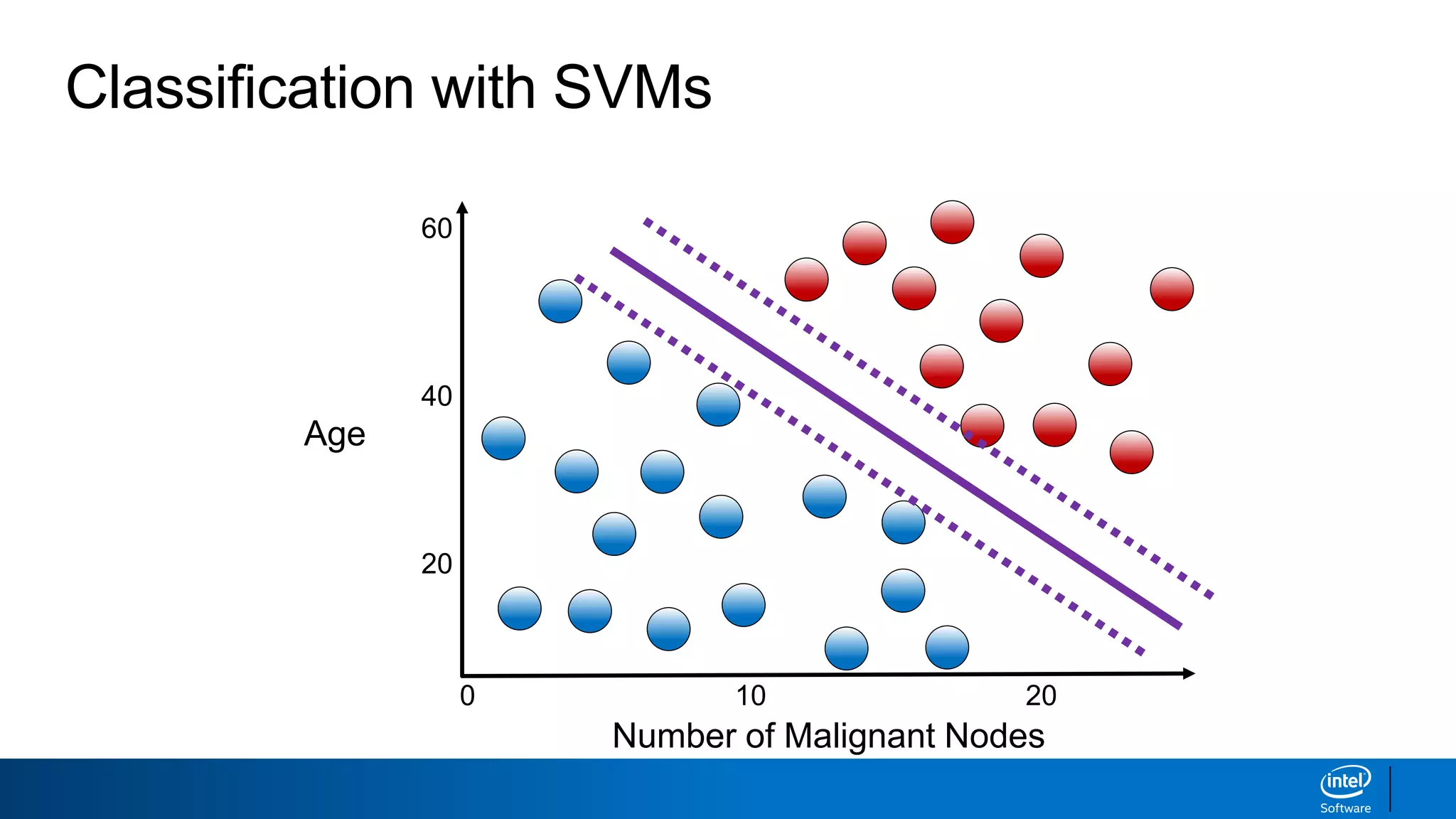
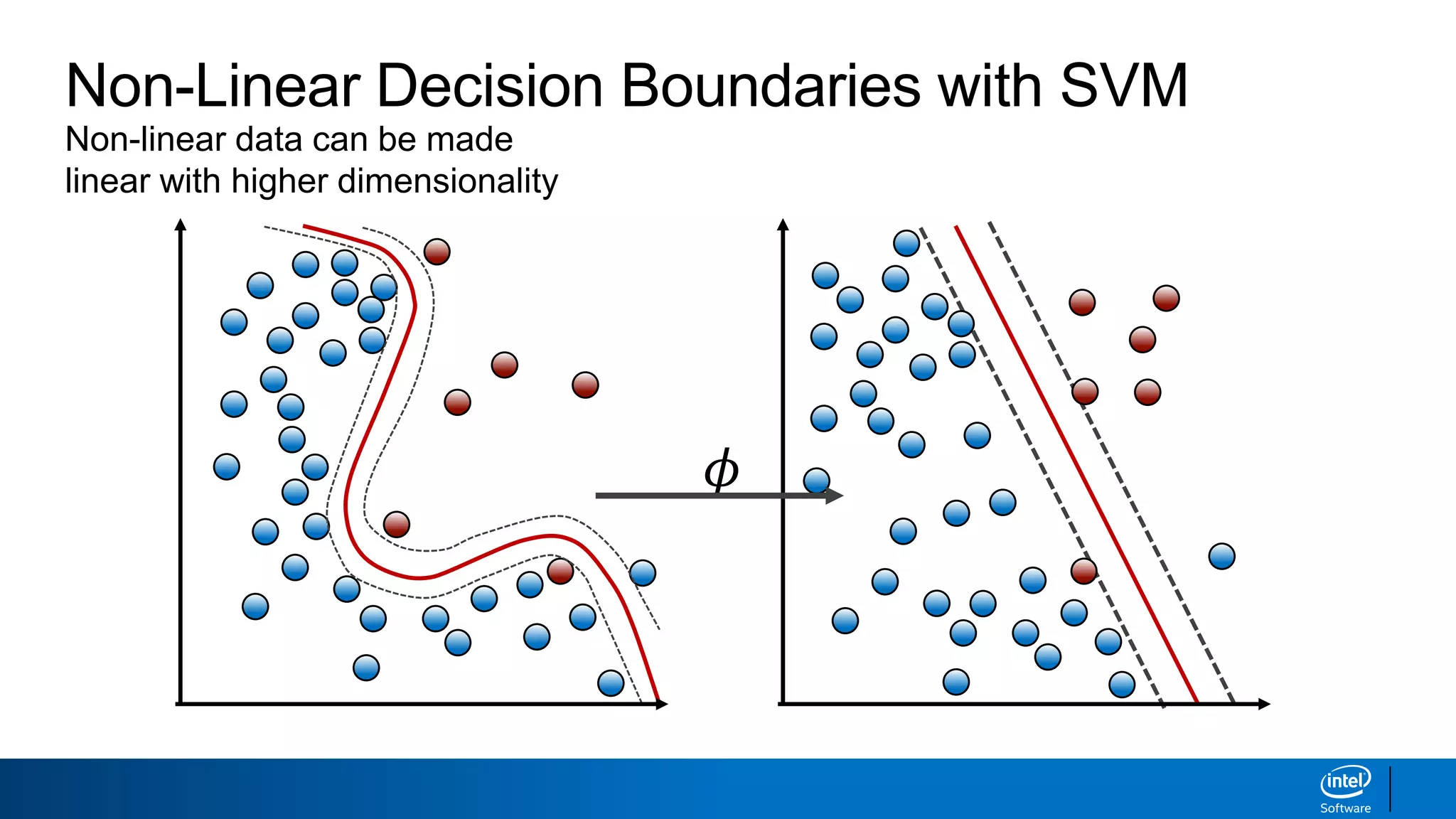
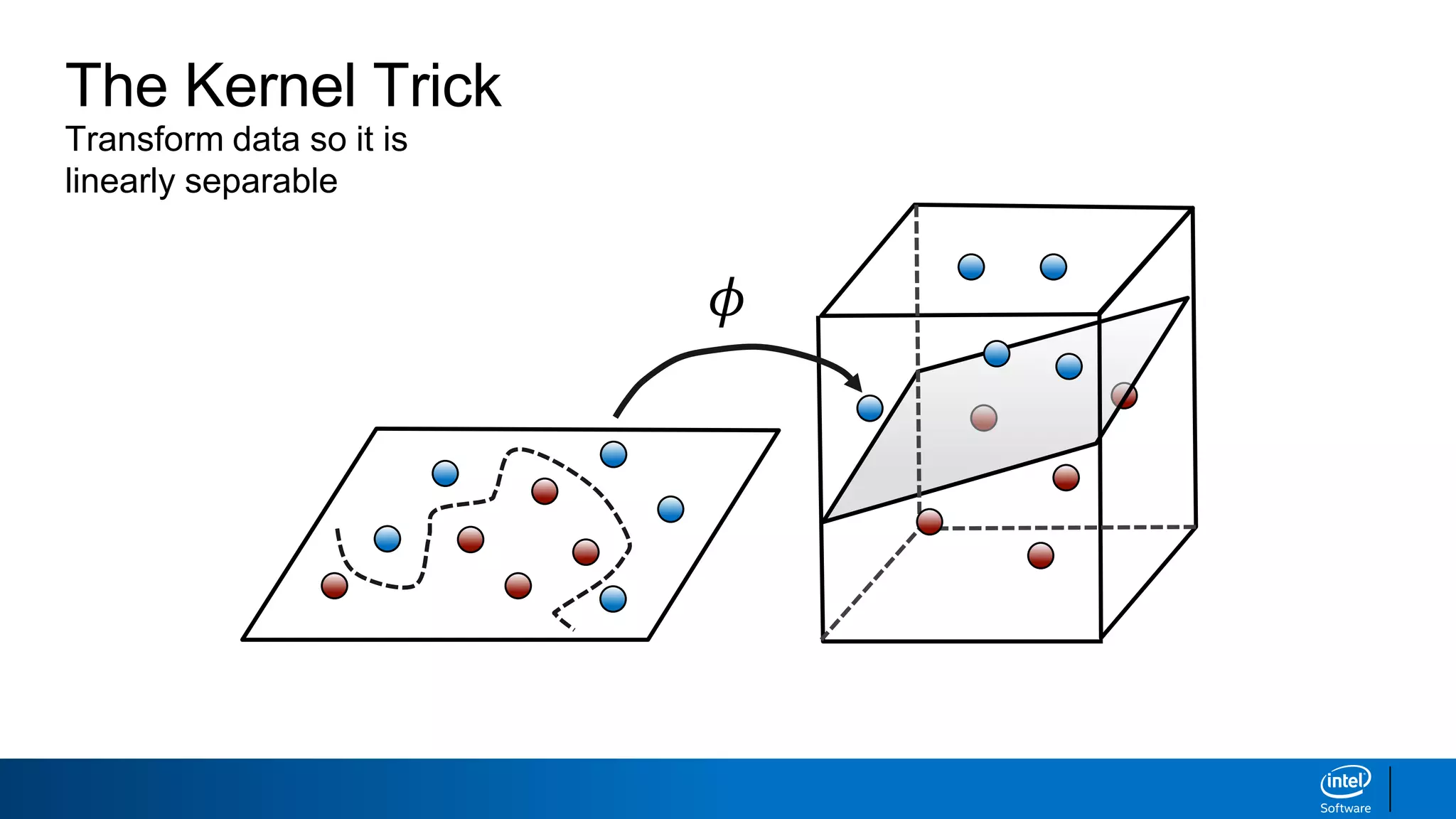
The document discusses support vector machines (SVMs). It explains that SVMs find the optimal separating hyperplane that maximizes the margin between two classes of data points. SVMs can handle both linear and non-linear classification problems by using kernels to implicitly map inputs into high-dimensional feature spaces. Common kernels include the radial basis function kernel, which transforms the data into an infinite dimensional space non-linearly.
















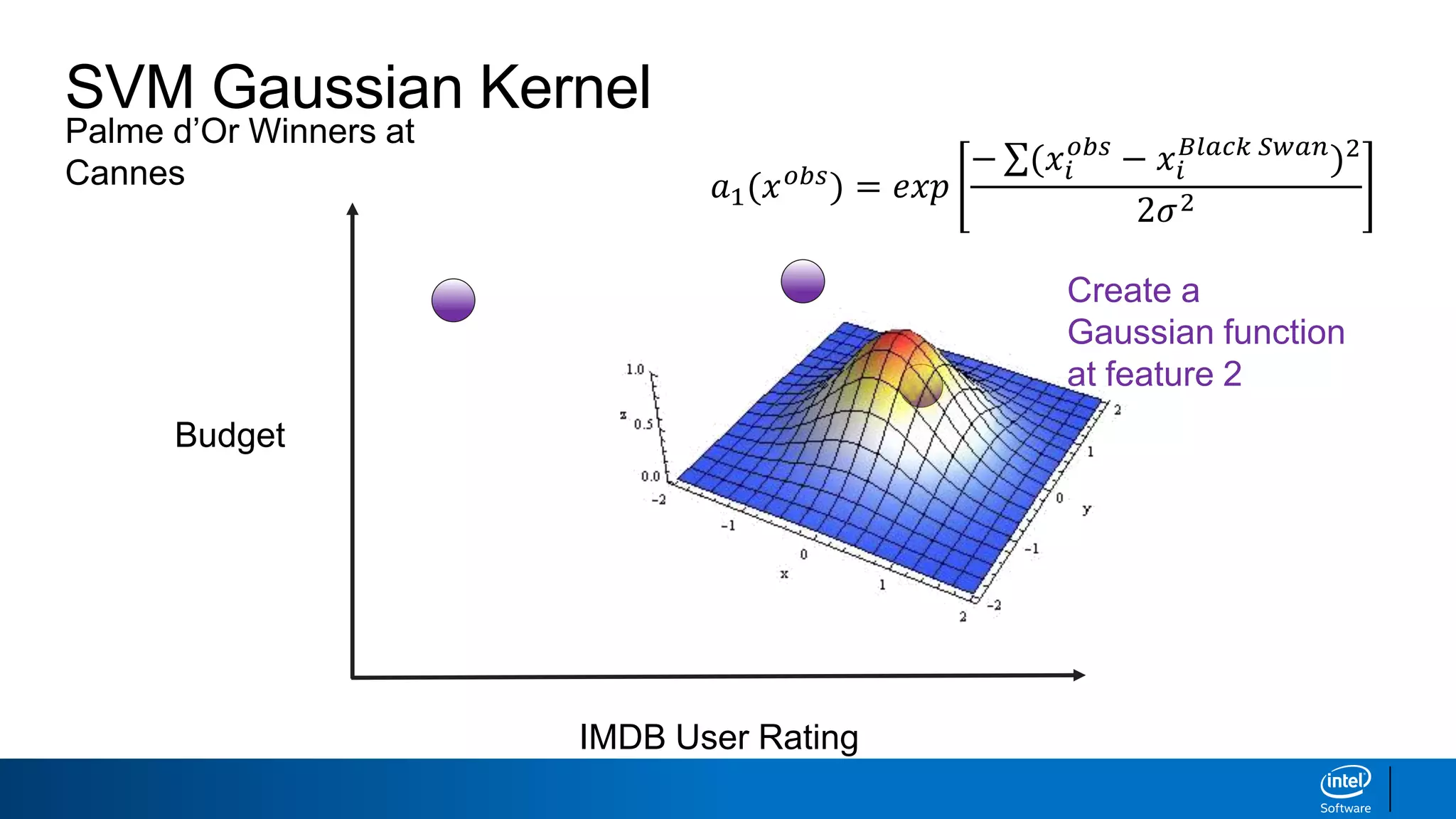
![SVM Gaussian Kernel
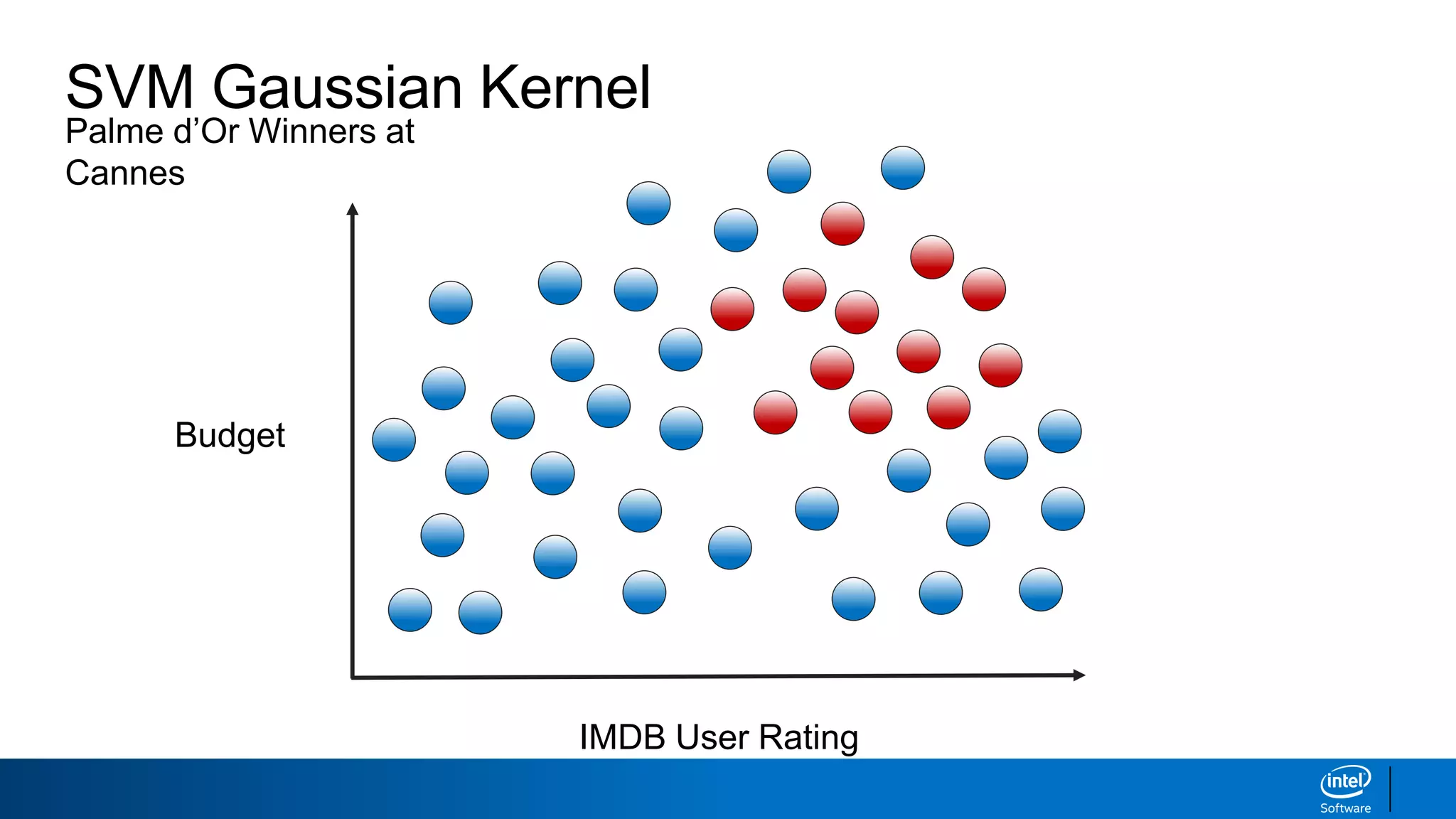
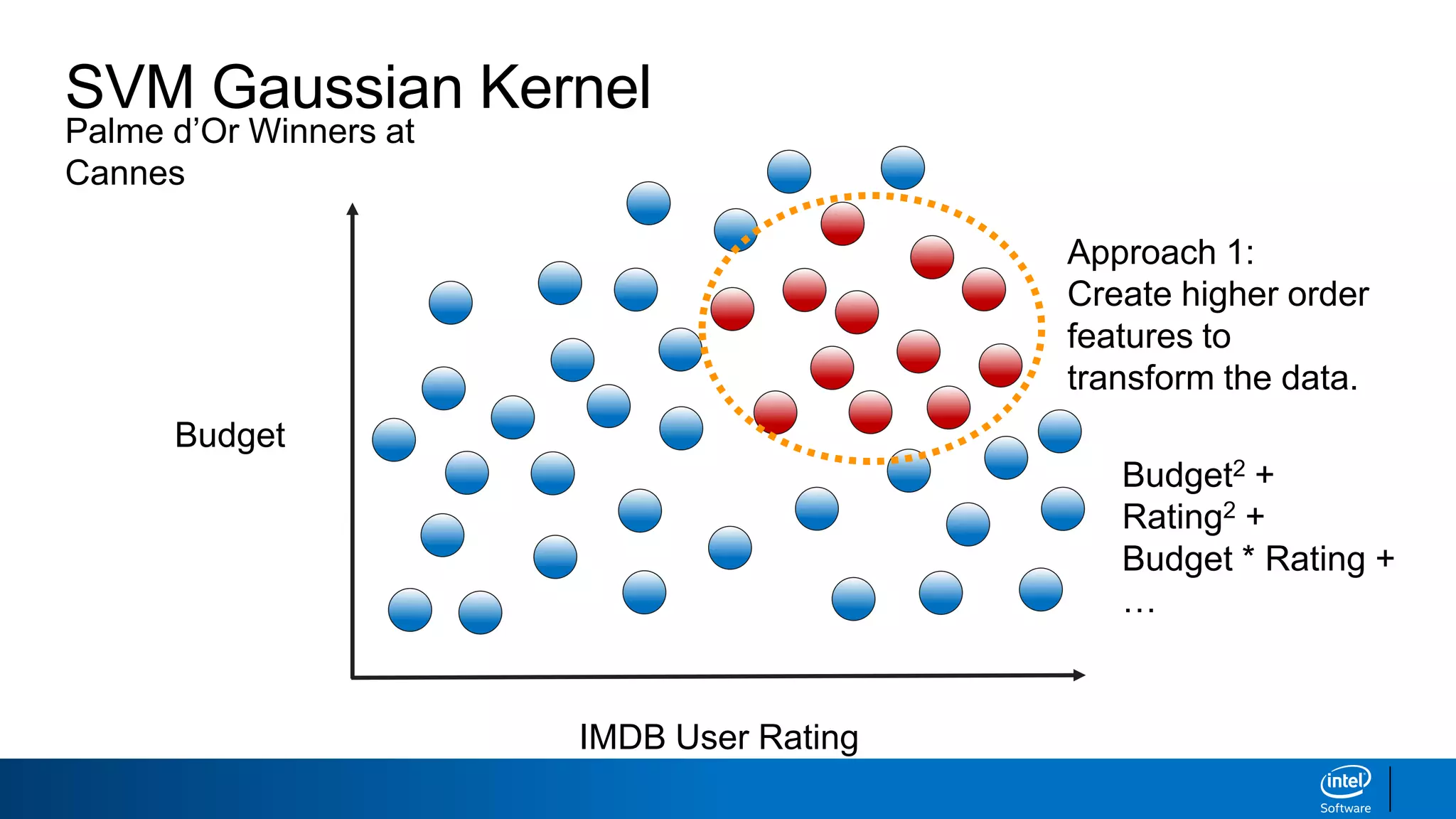
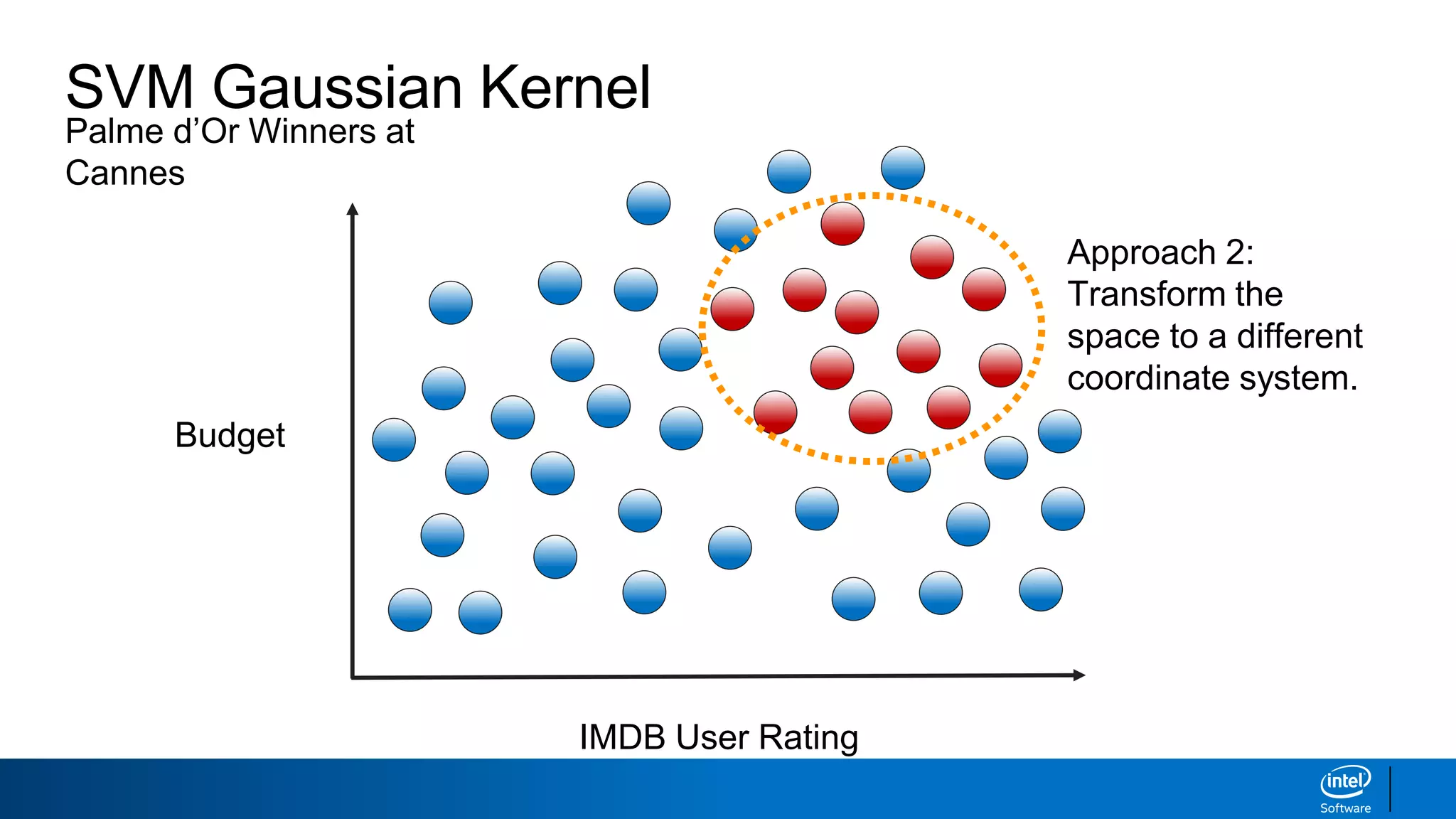
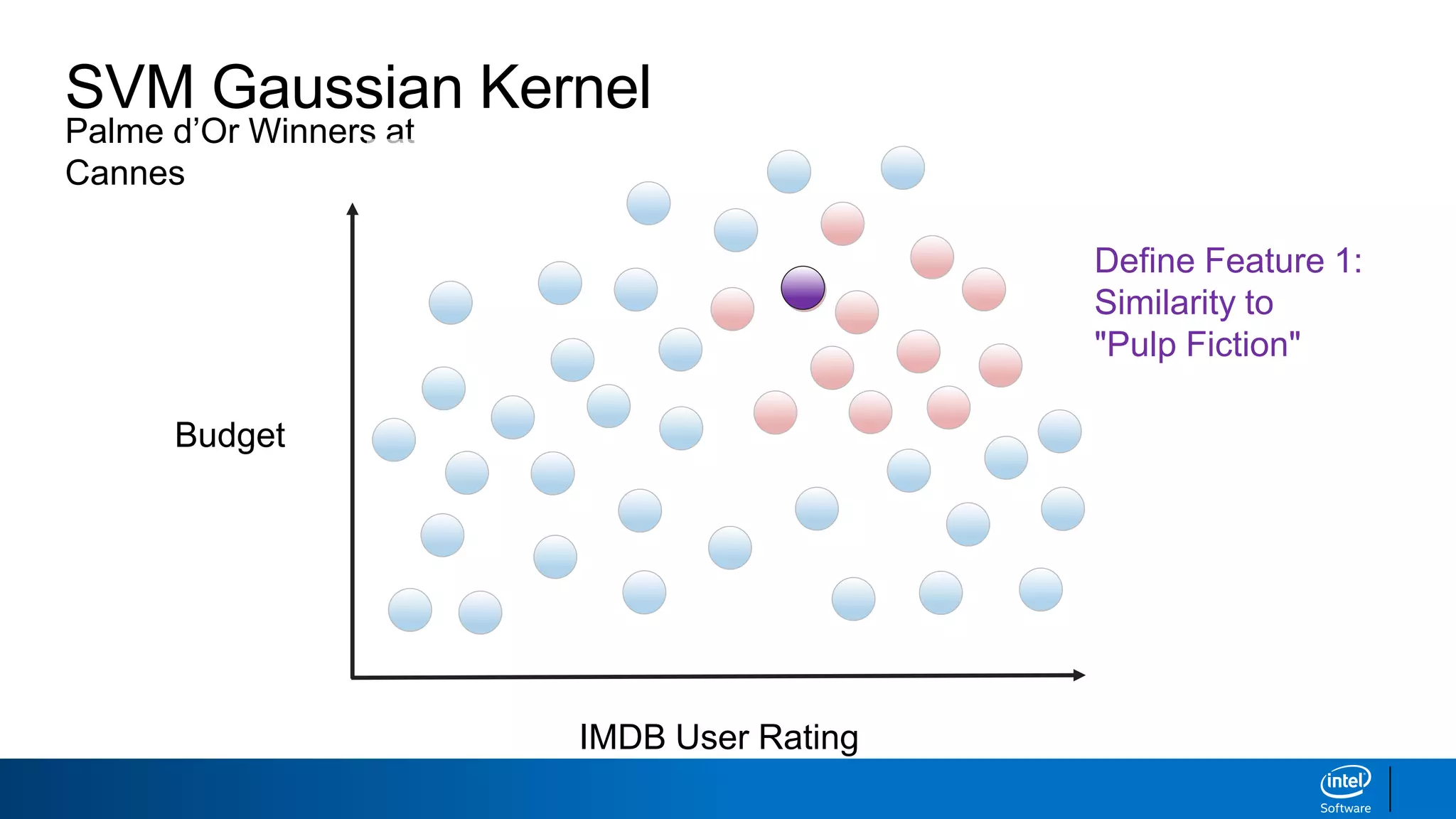
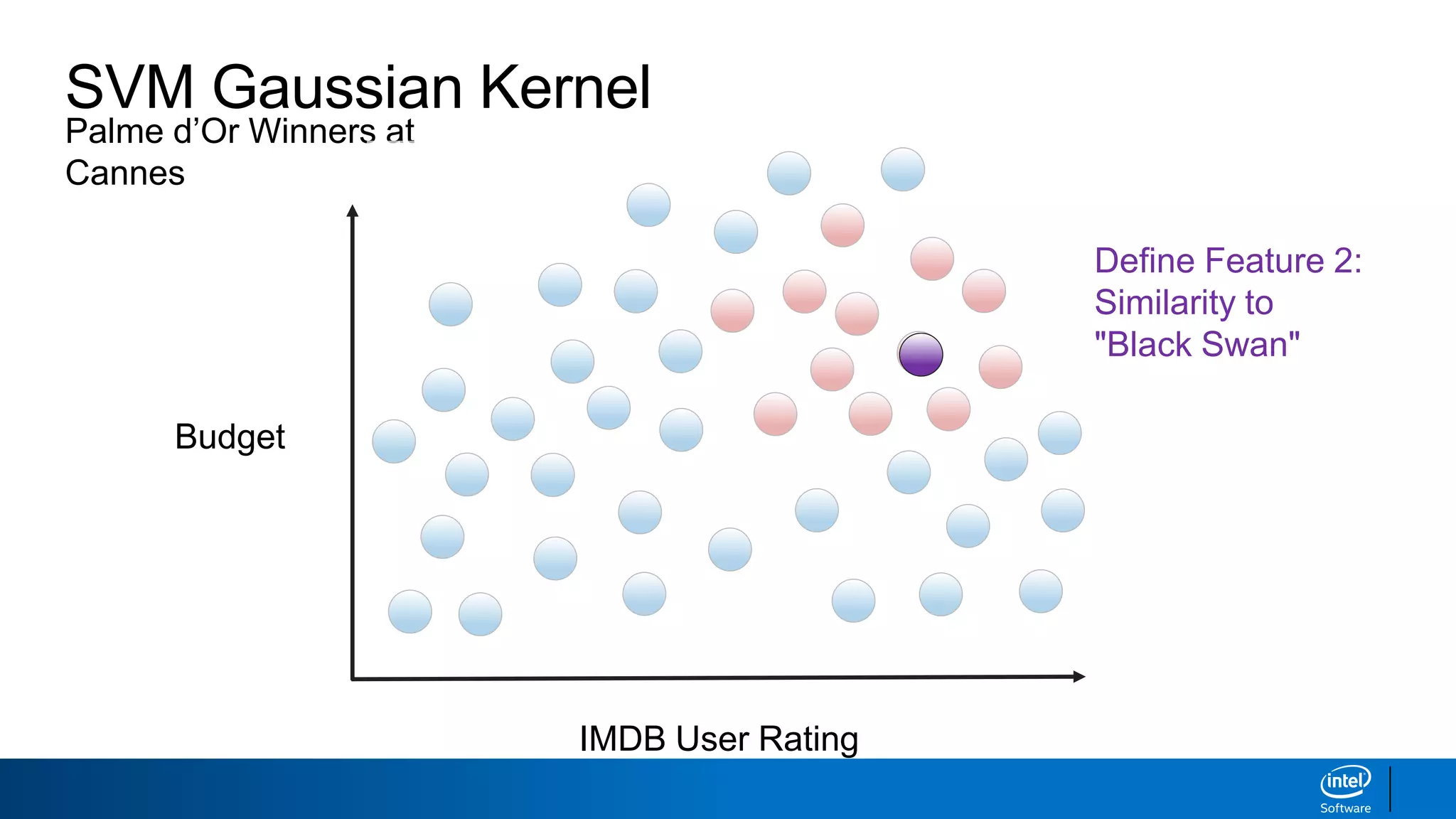
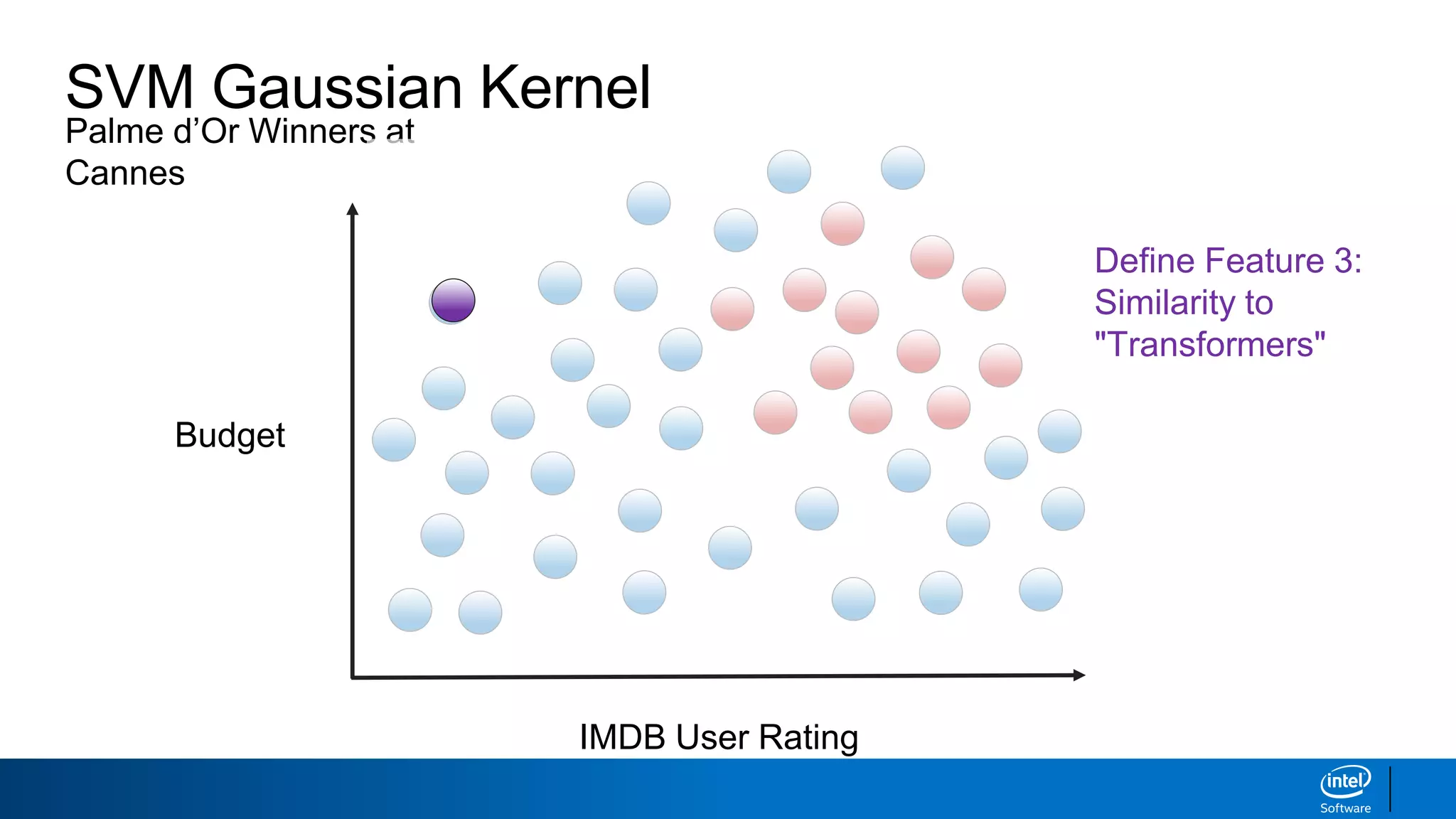
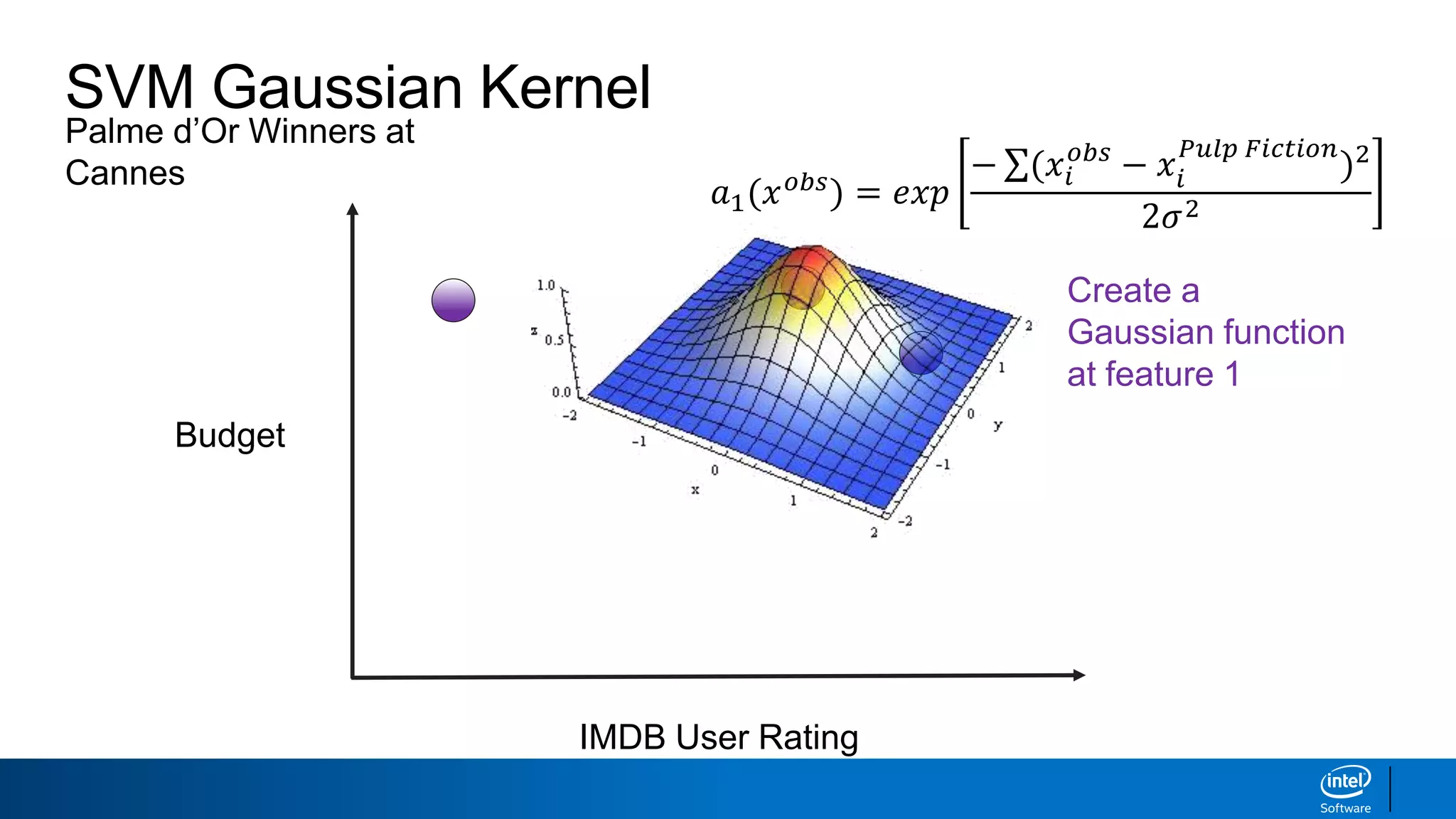
IMDB User Rating
Budget
Transformation:
[x1, x2] [0.7a1 , 0.9a2 , -0.6a3]](https://image.slidesharecdn.com/ml5-svmandkernels-190218095433/75/Ml5-svm-and-kernels-59-2048.jpg)
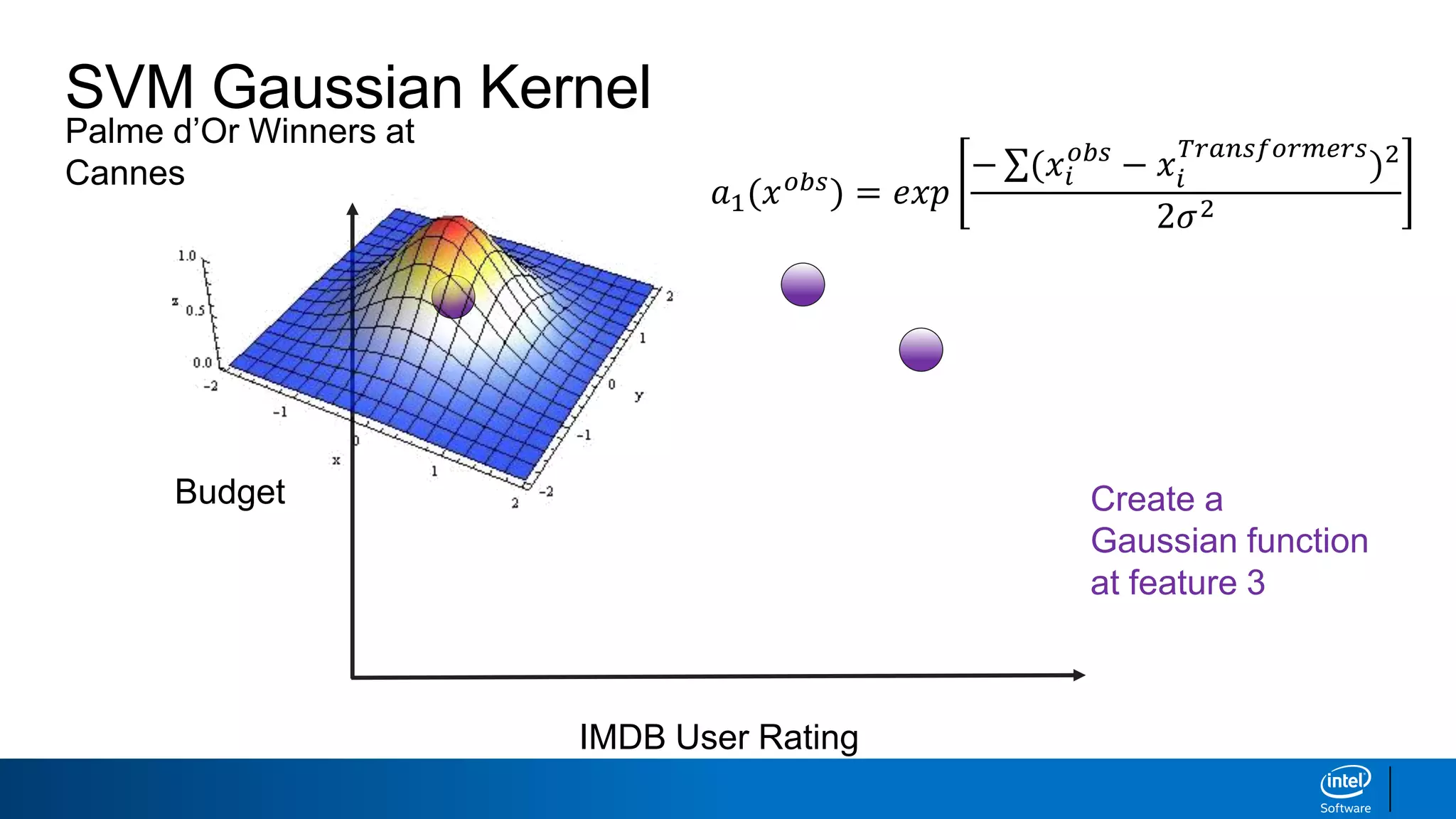
![SVM Gaussian Kernel
IMDB User Rating
Budget
a1=0.90
a2=0.92
a3=0.30
Transformation:
[x1, x2] [0.7a1 , 0.9a2 , -0.6a3]
a3
a2
a1](https://image.slidesharecdn.com/ml5-svmandkernels-190218095433/75/Ml5-svm-and-kernels-60-2048.jpg)
![SVM Gaussian Kernel
IMDB User Rating
Budget
a1=0.50
a2=0.60
a3=0.70 Transformation:
[x1, x2] [0.7a1 , 0.9a2 , -0.6a3]
a3
a1
a2](https://image.slidesharecdn.com/ml5-svmandkernels-190218095433/75/Ml5-svm-and-kernels-61-2048.jpg)
![SVM Gaussian Kernel
x2 (IMDB Rating)
x1(Budget)
a1 (Pulp Fiction)
a3 (Transformers)
a2 (Black Swan)
Transformation:
[x1, x2] [0.7a1 , 0.9a2 , -0.6a3]](https://image.slidesharecdn.com/ml5-svmandkernels-190218095433/75/Ml5-svm-and-kernels-62-2048.jpg)
![Classification in the New Space
a1 (Pulp Fiction)
a3 (Transformers)
a2 (Black Swan)
Transformation:
[x1, x2] [0.7a1 , 0.9a2 , -0.6a3]](https://image.slidesharecdn.com/ml5-svmandkernels-190218095433/75/Ml5-svm-and-kernels-63-2048.jpg)