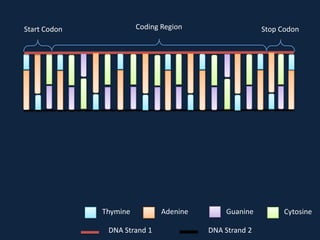
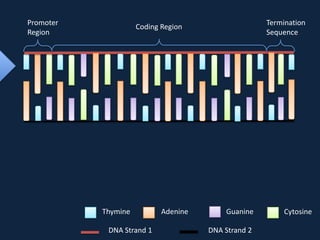
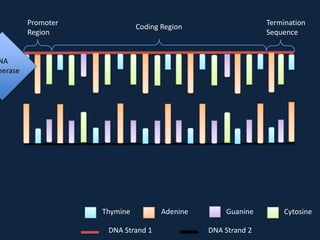
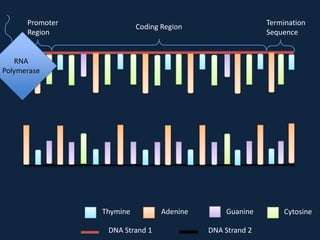
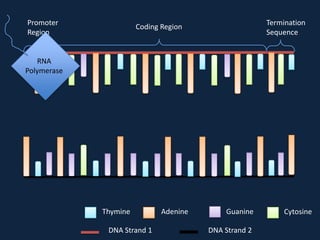
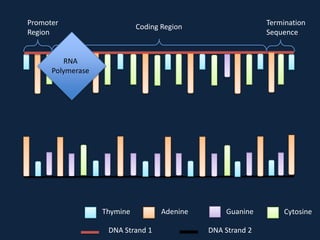
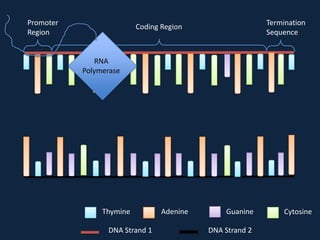
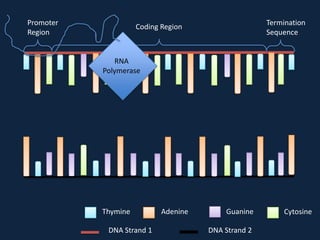
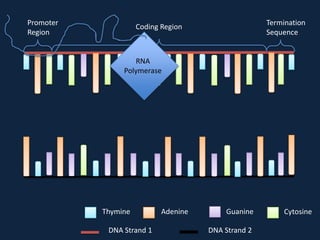
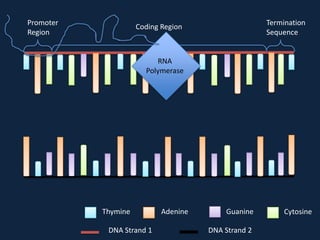
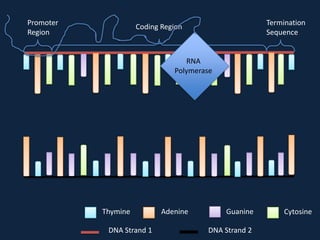
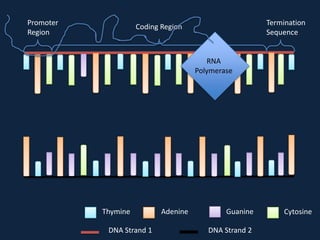
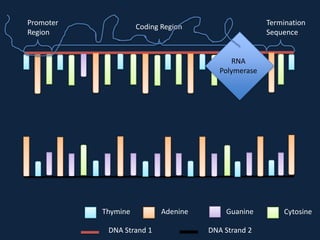
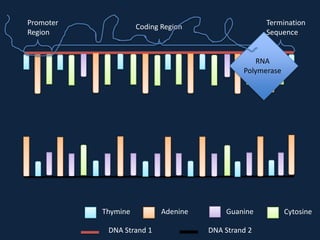
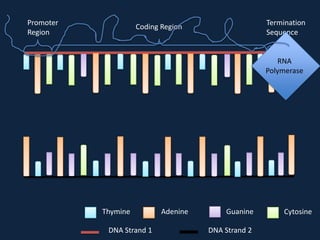
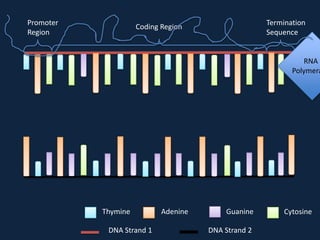
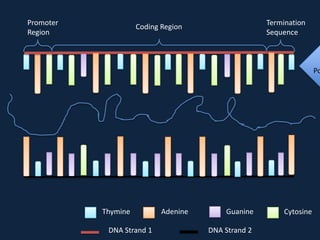
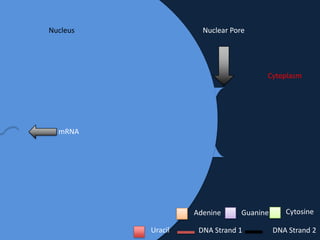
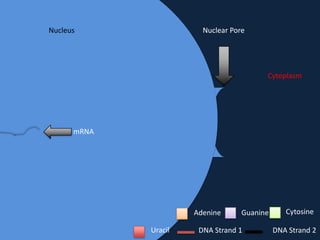
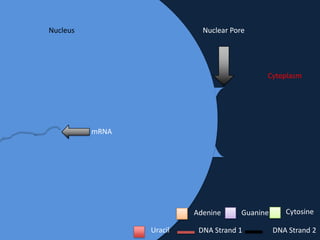
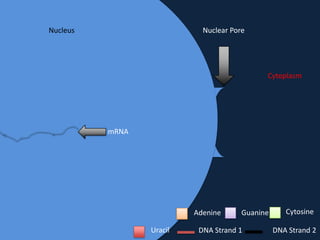
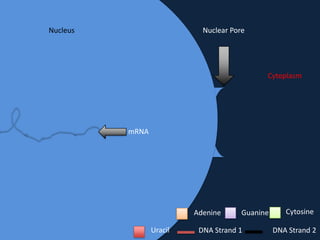
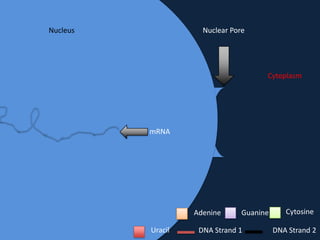
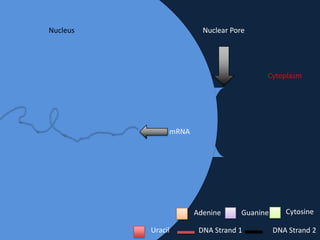
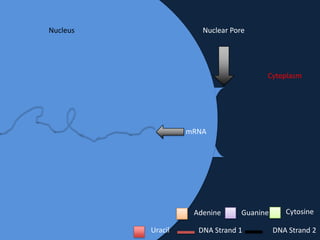
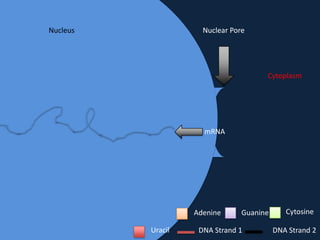
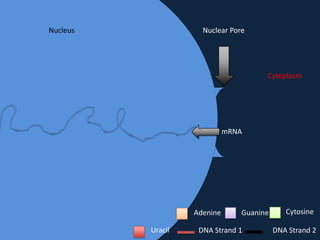
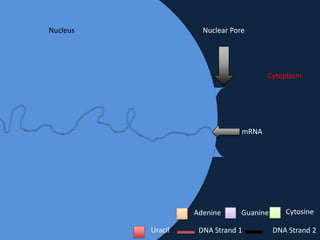
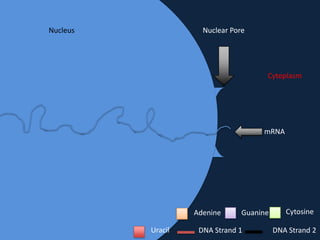
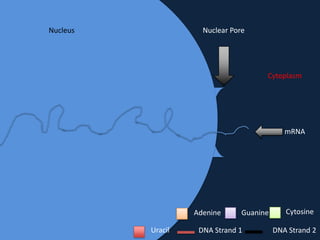
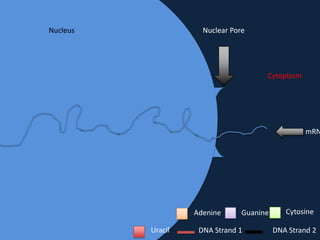
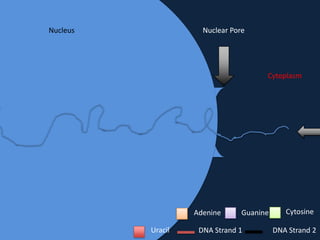
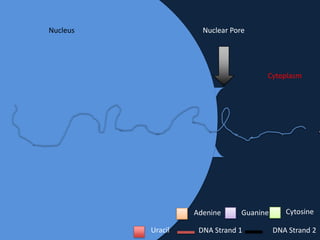
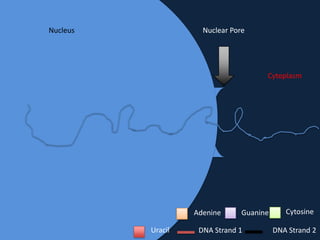
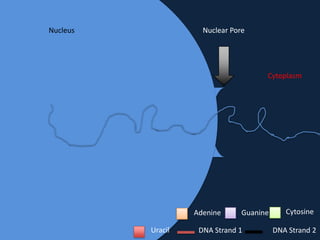
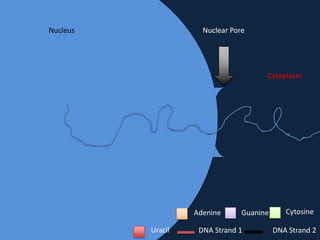
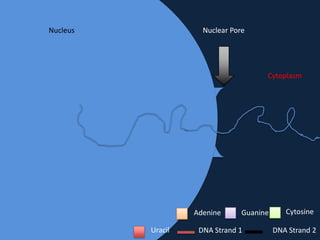
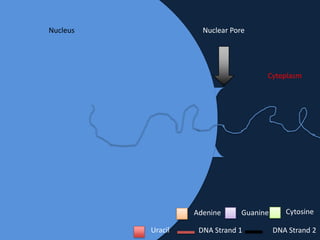
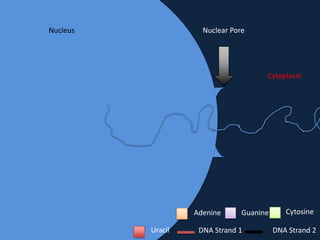
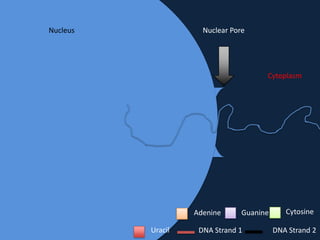
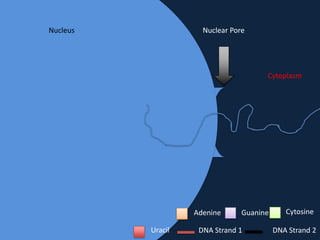
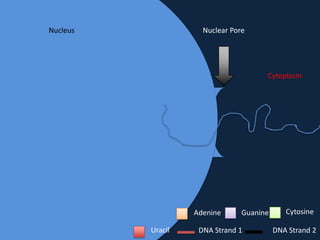
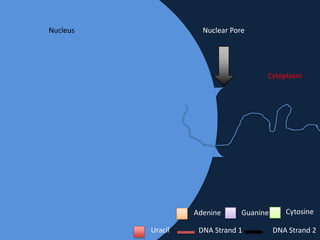
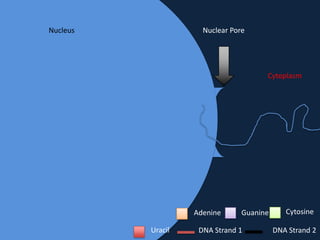
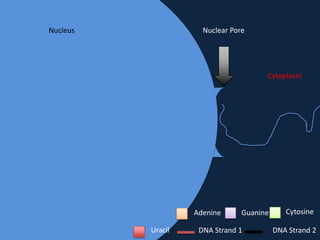
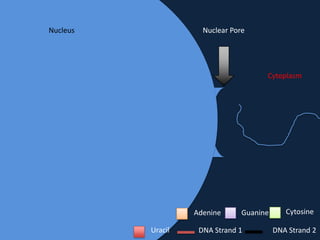
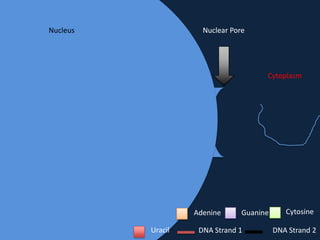
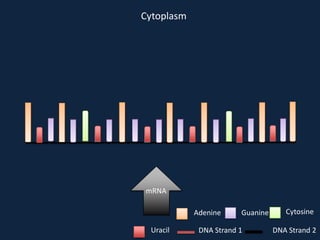
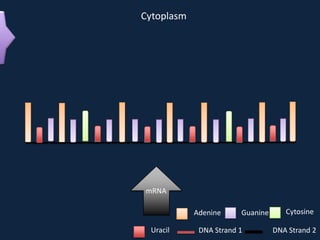
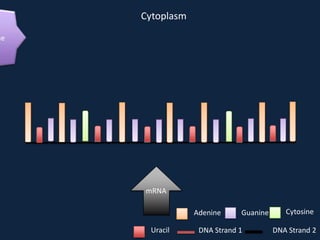
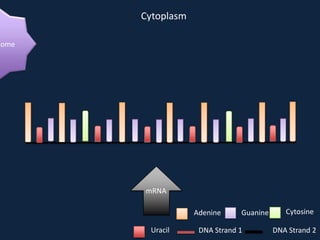
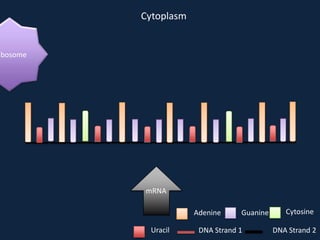
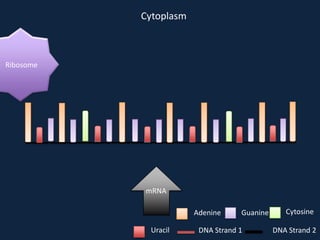
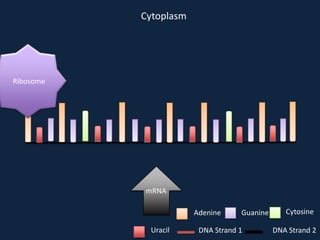
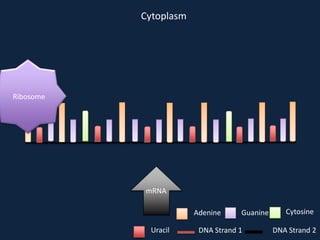
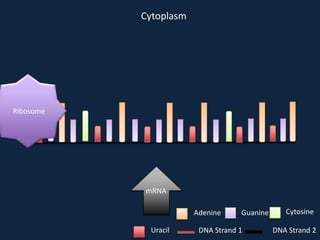
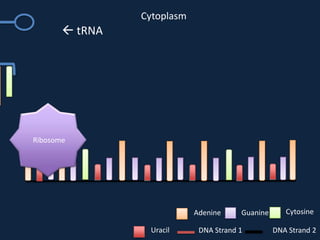
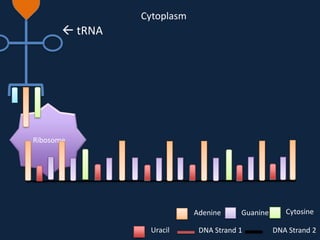
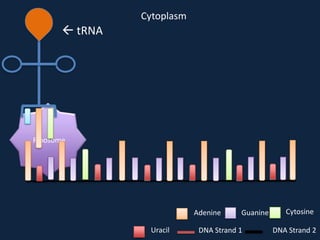
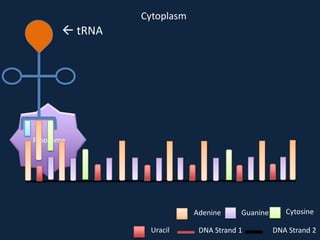
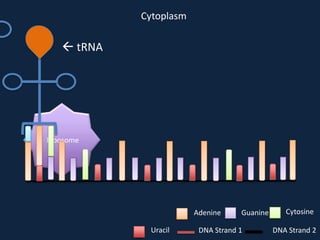
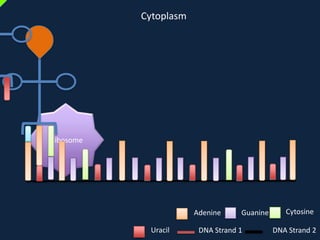
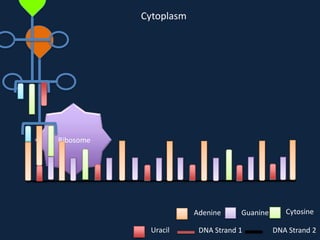
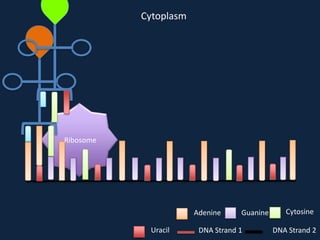
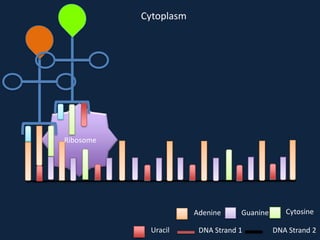
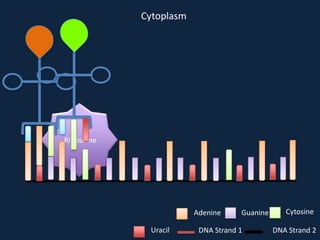
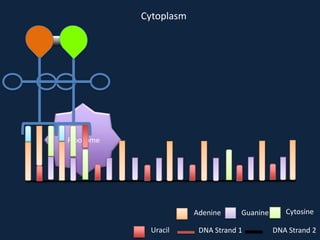
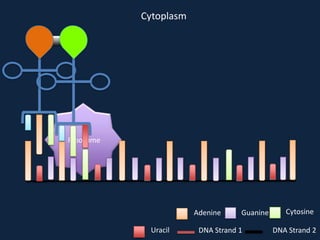
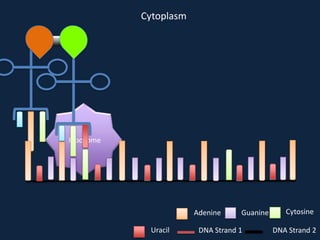
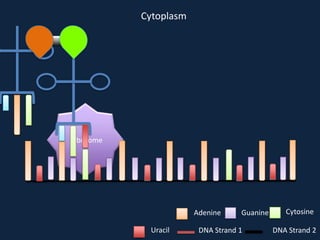
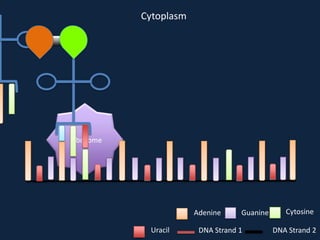
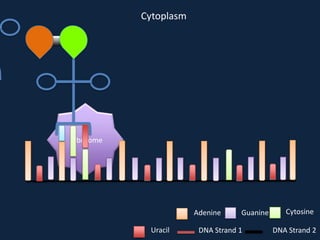
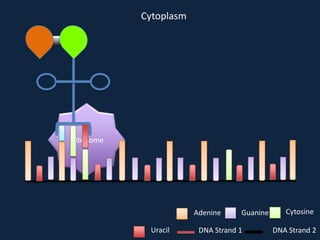
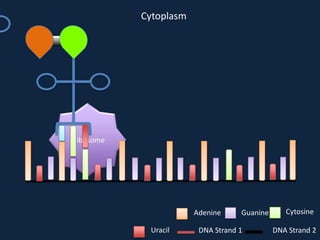
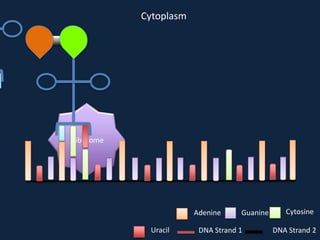
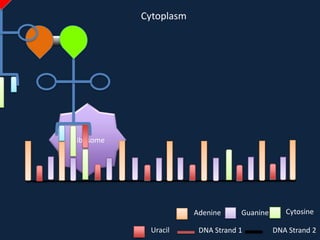
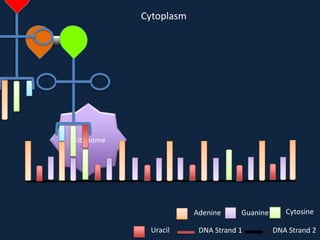
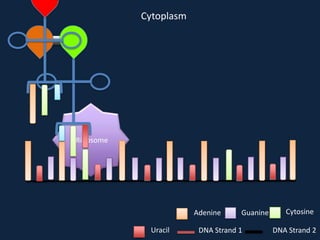
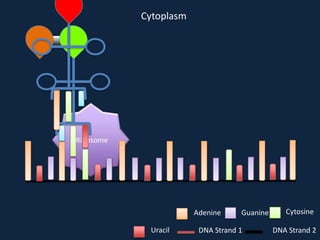
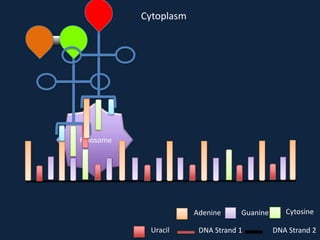
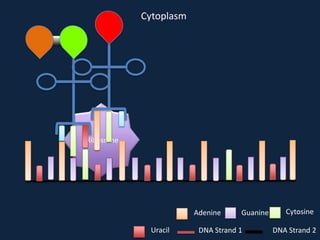
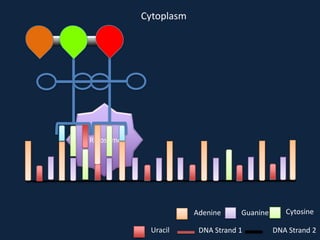
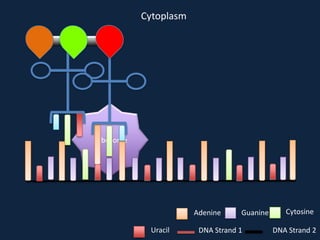
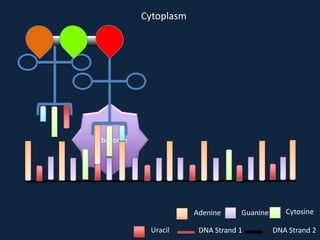
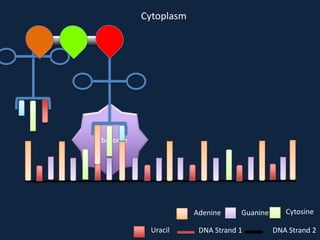
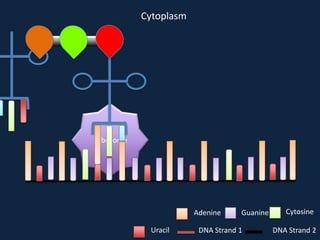
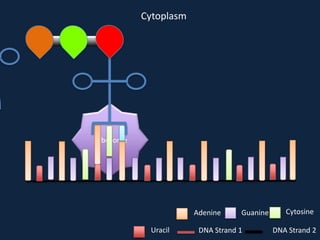
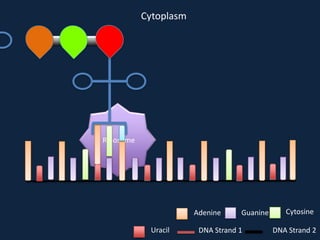
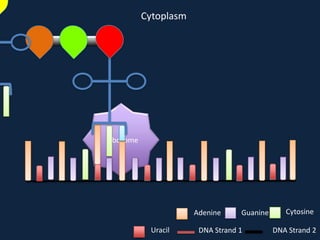
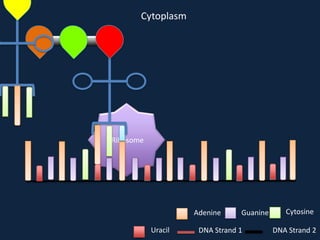
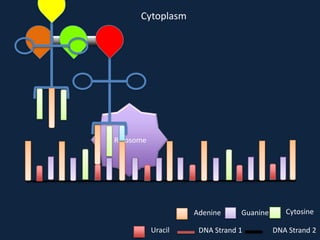
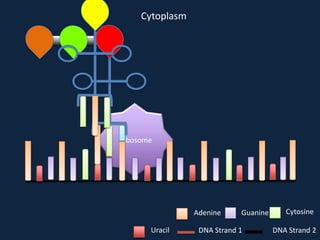
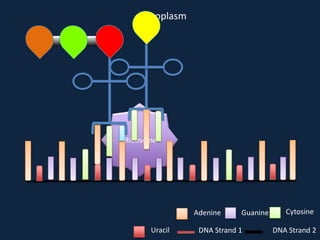
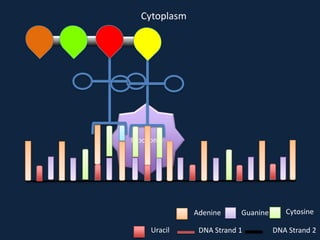
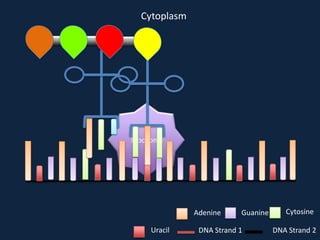
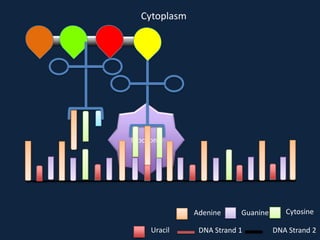
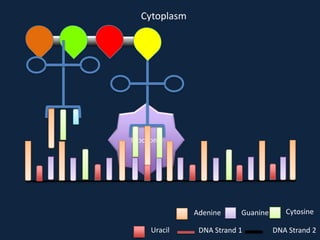
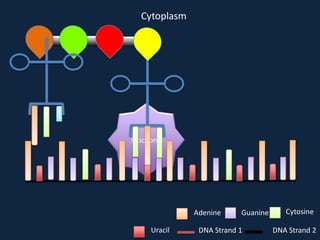
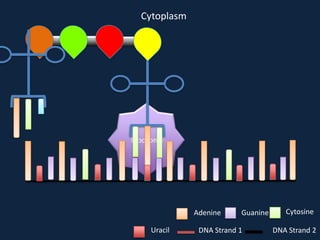
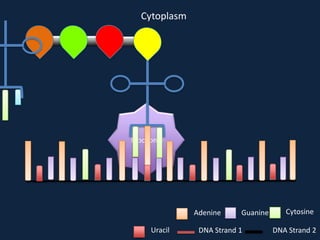
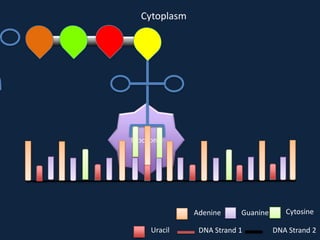
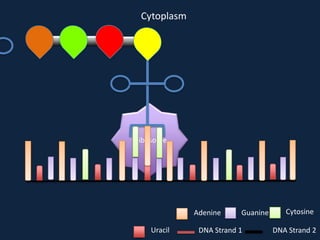
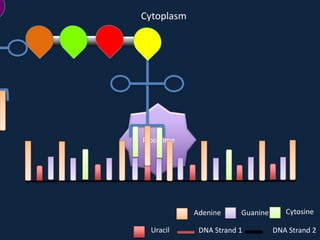
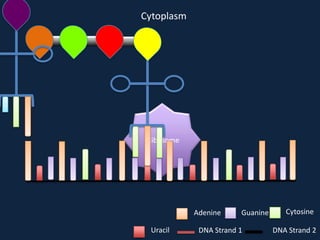
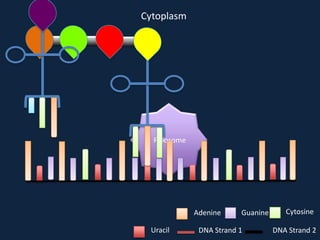
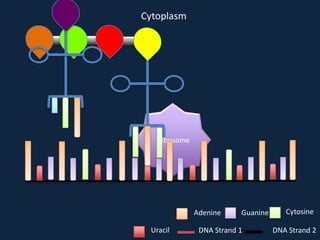
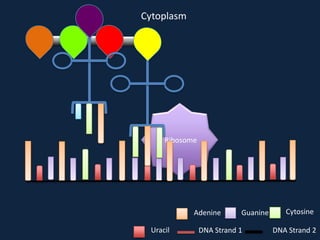
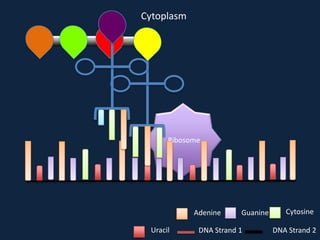
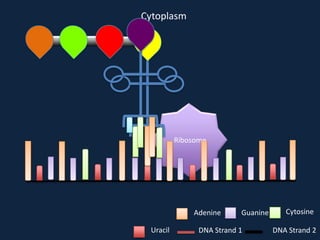
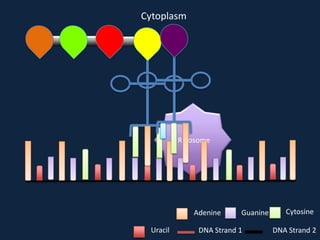
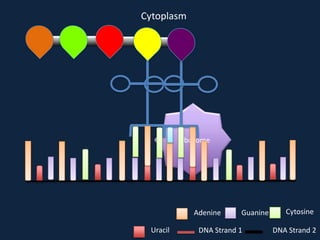
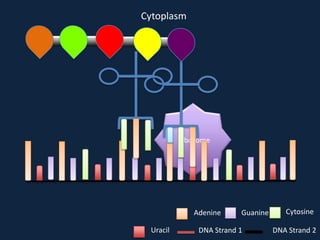
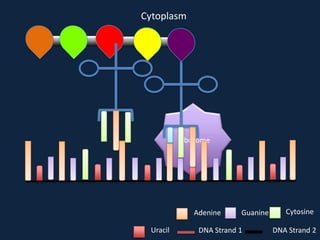
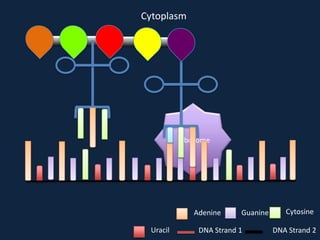
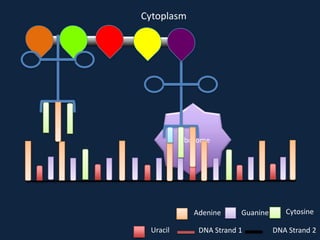
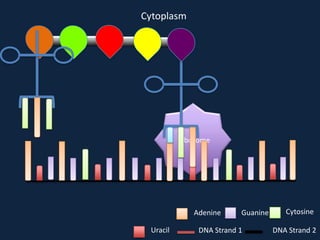
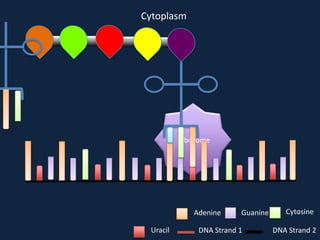
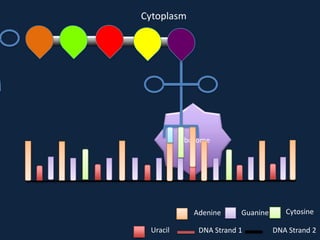
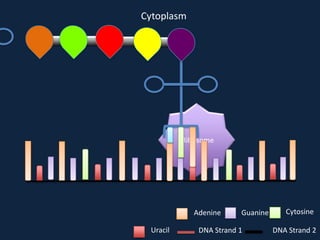
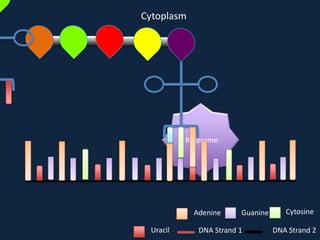
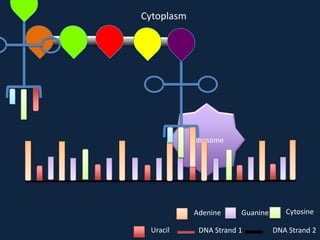
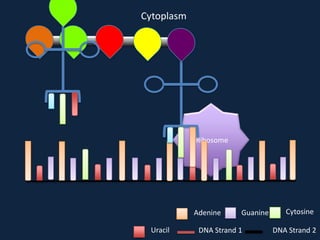
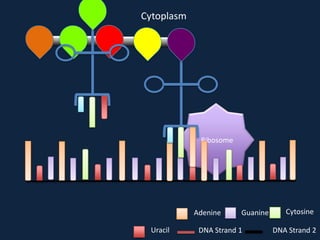
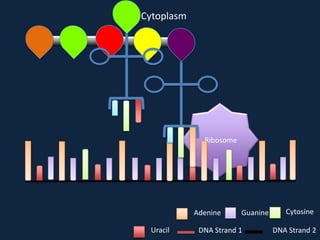
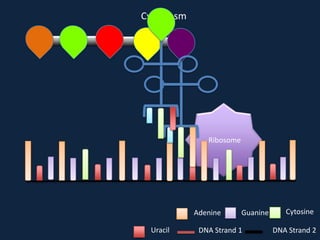
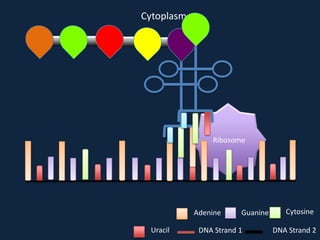
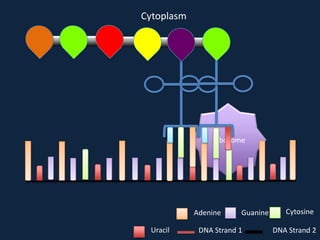
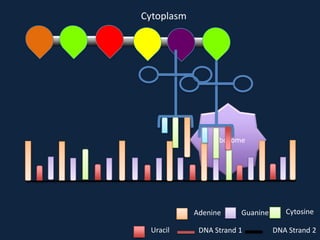
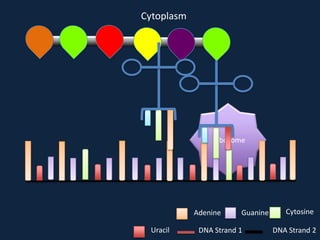
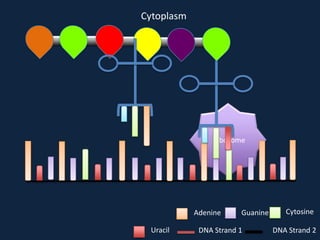
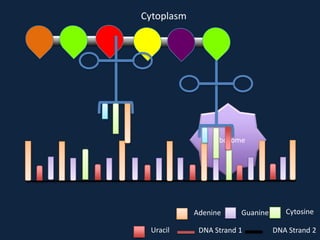
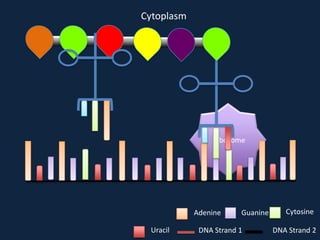
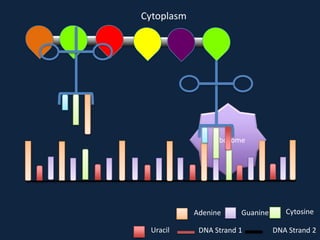
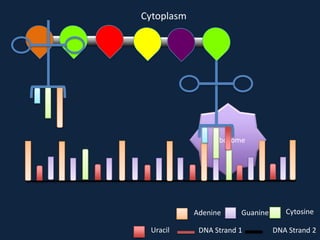
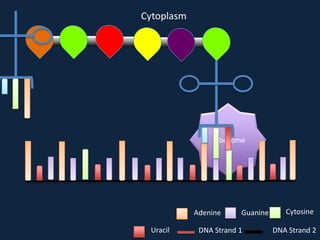
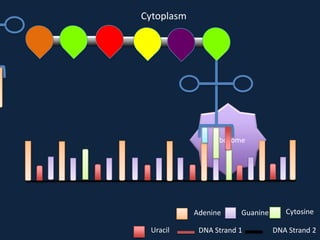
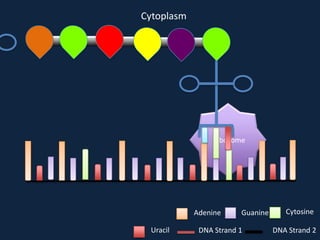
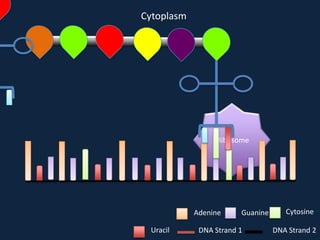
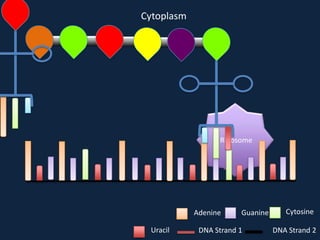
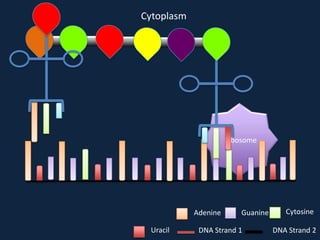
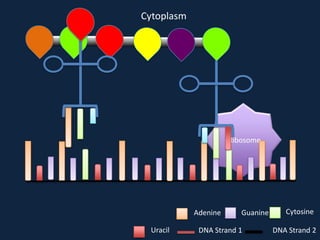
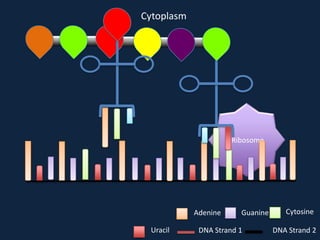
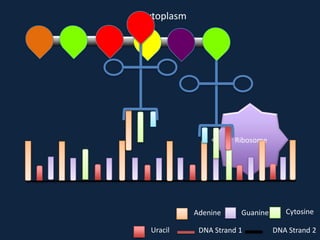
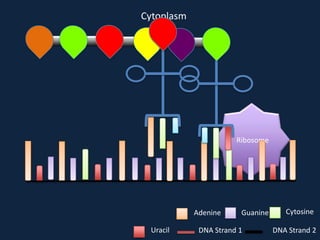
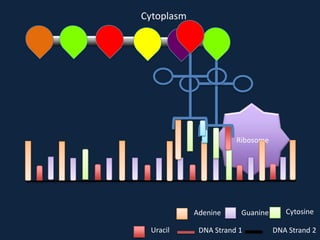
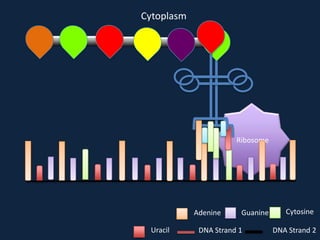
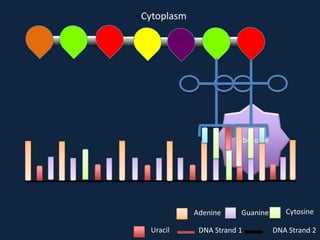
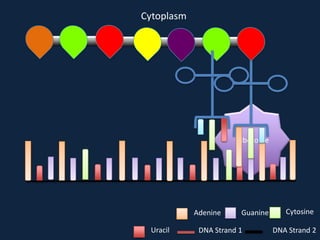
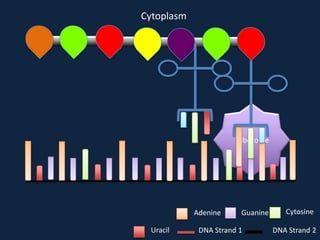
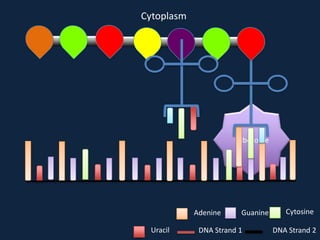
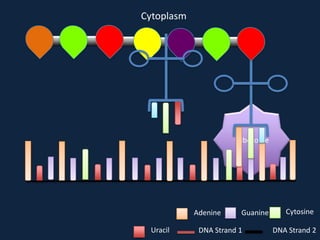
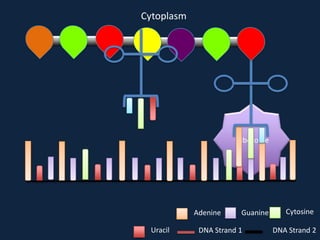
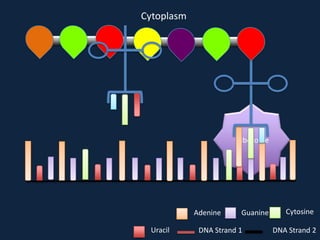
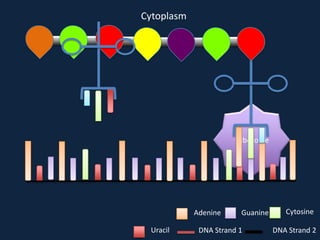
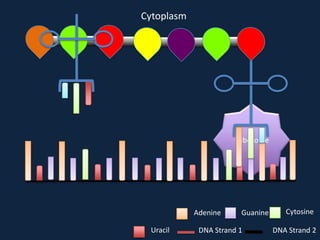
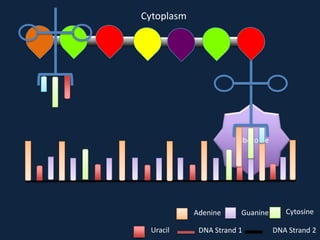
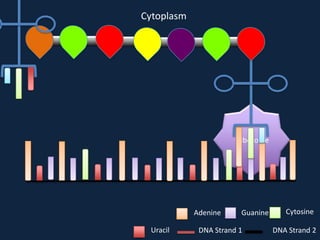
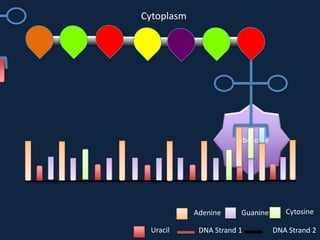
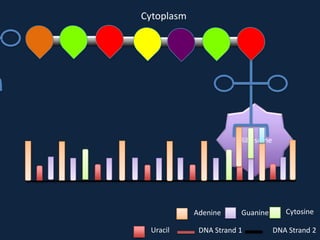
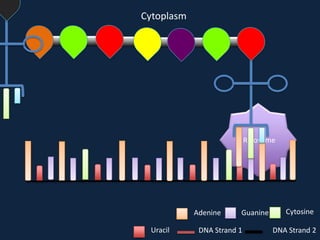
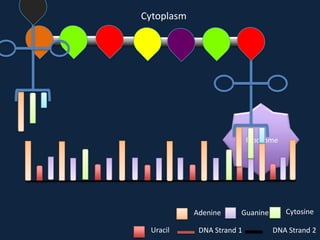
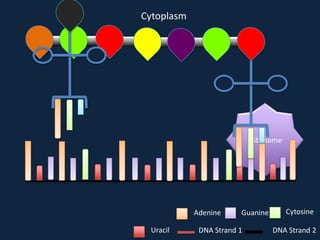
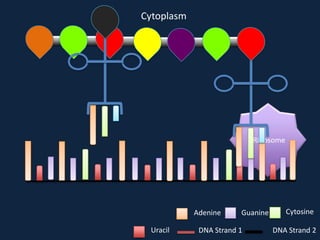
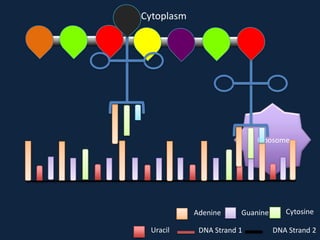
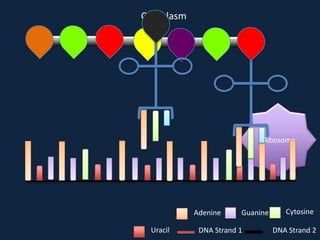
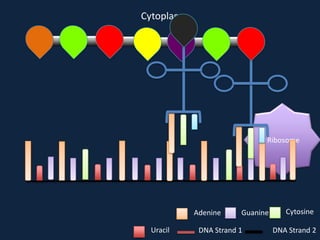
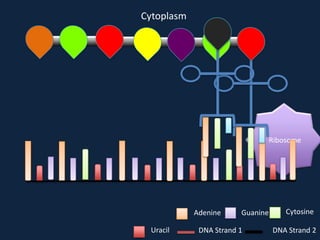
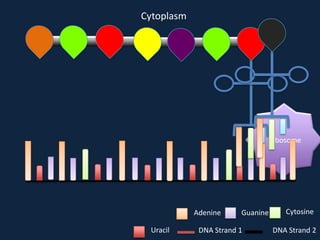
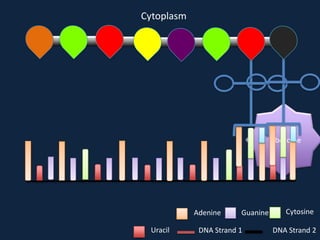
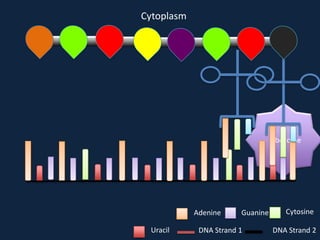
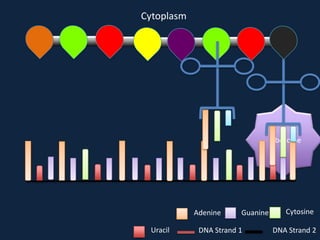
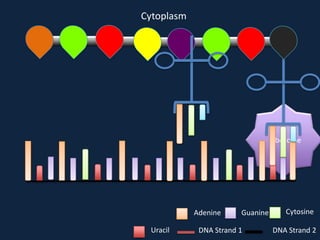
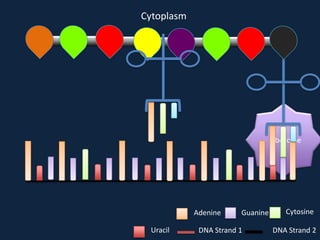
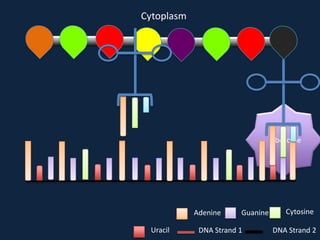
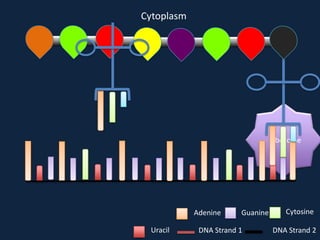
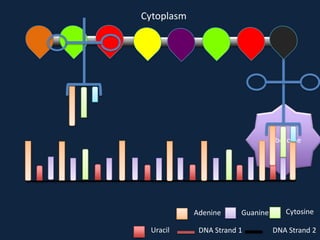
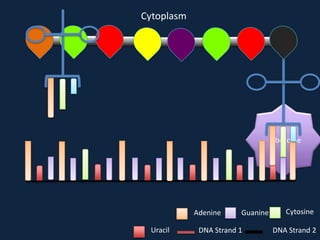
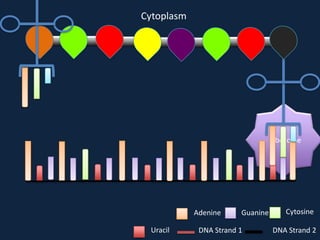
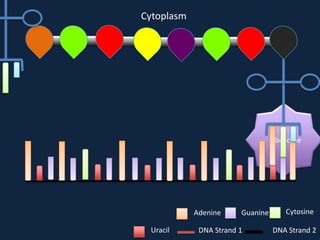
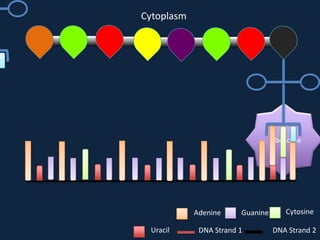
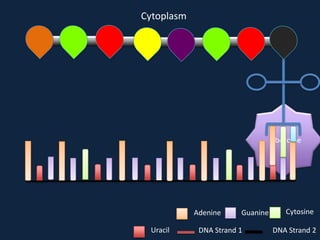
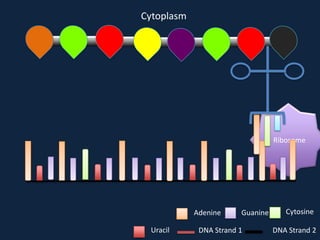
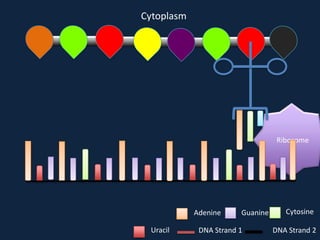
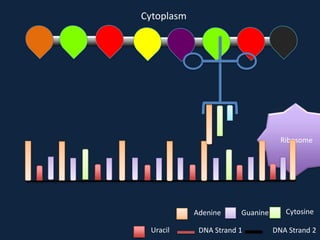
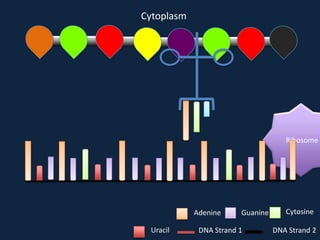
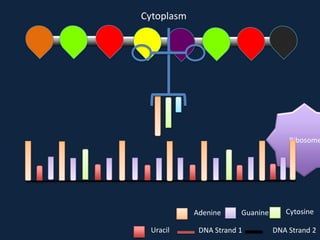
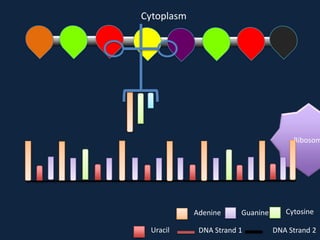
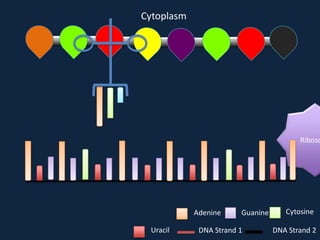
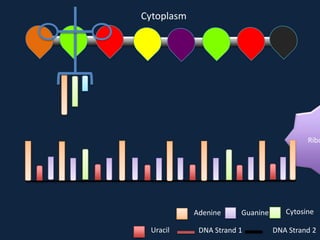
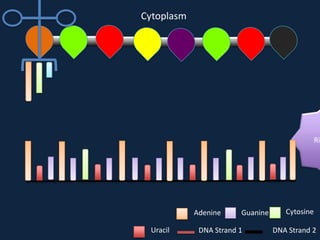
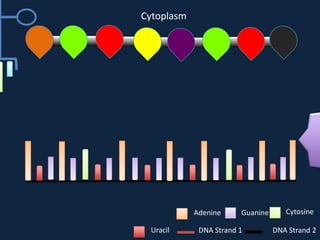
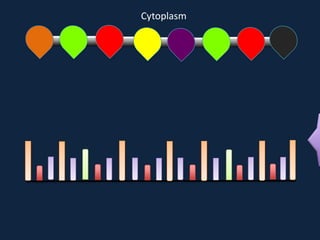
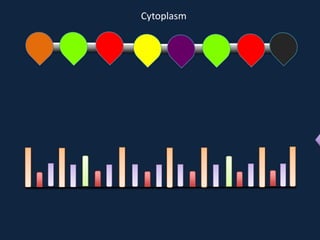
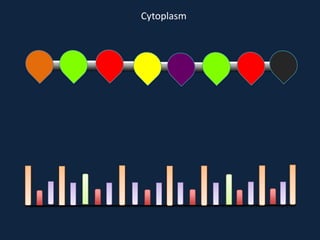
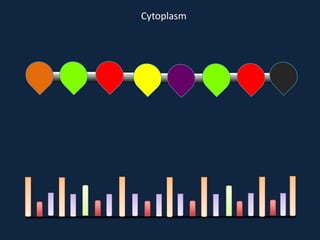
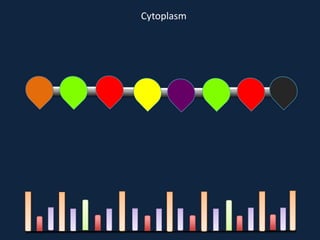
Transcription occurs in the nucleus, where RNA polymerase copies DNA into mRNA. The mRNA then exits the nucleus through the nuclear pore and enters the cytoplasm. In the cytoplasm, ribosomes use the mRNA as a template to assemble amino acids specified by the mRNA into a polypeptide chain through translation.