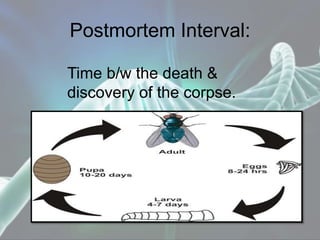
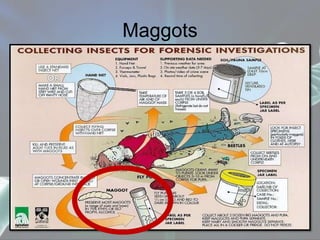
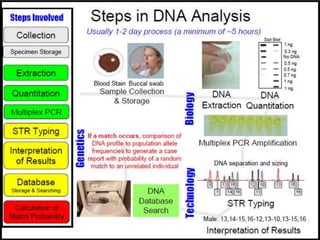
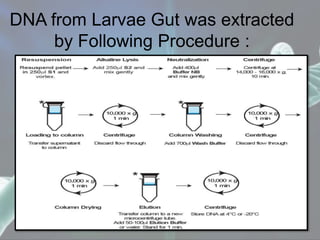
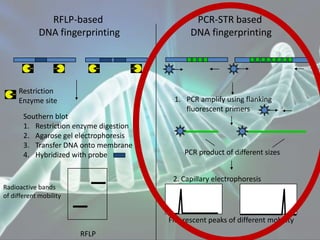
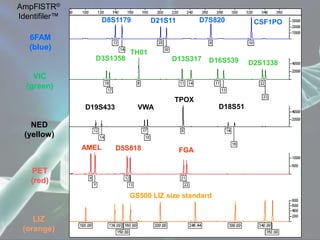
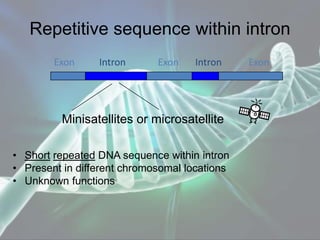
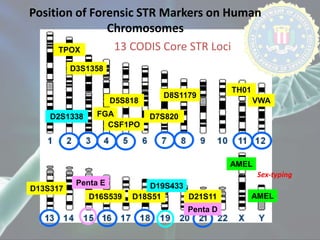
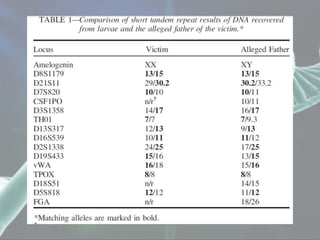
The document presents a forensic case report involving a badly burned female corpse, with key elements including the use of forensic entomology and DNA analysis for victim identification. The methodology described includes the extraction of DNA from maggots and buccal samples from an alleged father, alongside various DNA fingerprinting techniques such as PCR and capillary electrophoresis. The findings suggest a high probability of paternity testing and highlight the significance of entomological evidence in forensic investigations.