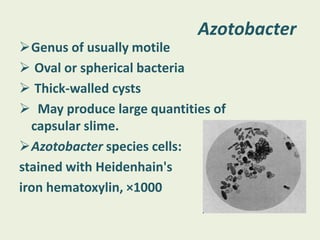
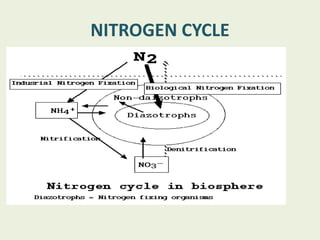
Azotobacter is a genus of free-living, motile bacteria known for forming cysts and their ability to fix atmospheric nitrogen into ammonia. They are predominantly found in neutral to alkaline soils and have significant metabolic capabilities, making them valuable for agricultural nitrogen fixation. Their unique nitrogenase enzymes and high metabolic rate contribute to their role in nutrient competition and growth promotion in various environments.