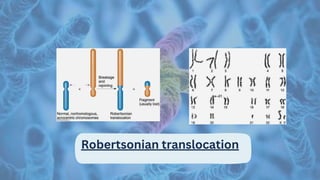
Translocation involves the exchange of parts between non-homologous chromosomes, leading to chromosomal rearrangements that can cause various disorders in humans, including mental retardation, infertility, and cancer. The document describes two main types of translocation: reciprocal, which can occur through alternate or adjacent segregation during meiosis, and Robertsonian translocation, resulting from fusions of acrocentric chromosomes. Translocations can be classified as balanced, where genetic material remains unchanged, or unbalanced, which often leads to phenotypic abnormalities.