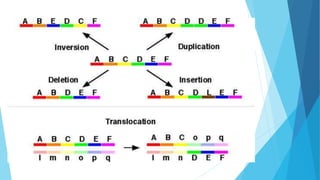
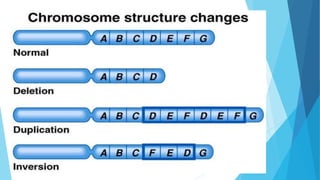
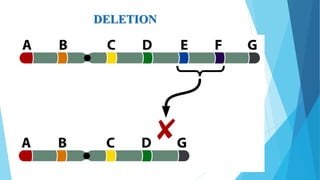
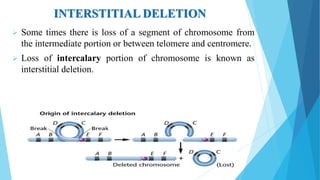
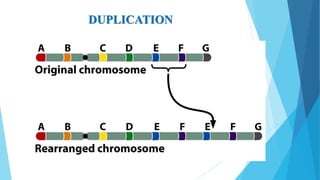
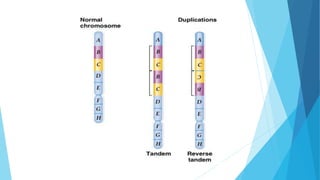
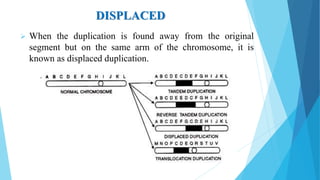
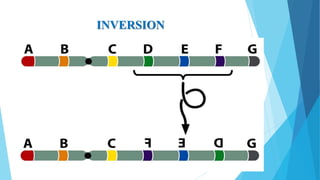
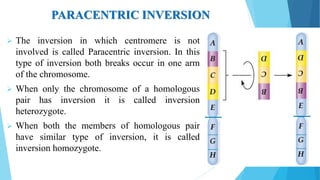
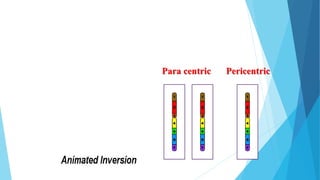
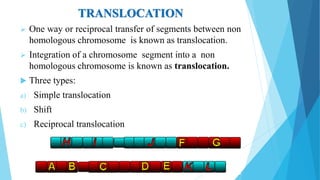
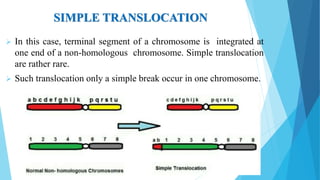
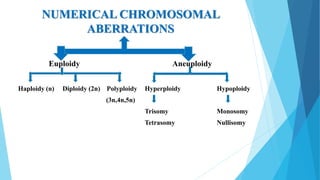
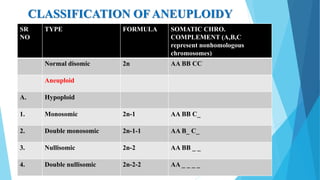
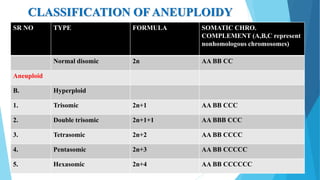
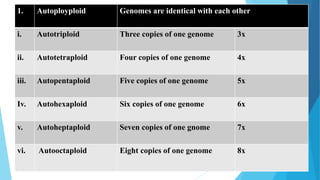
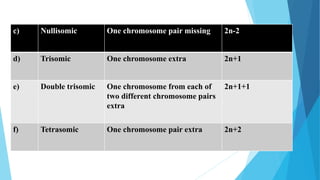
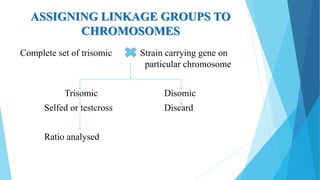
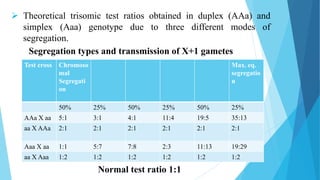
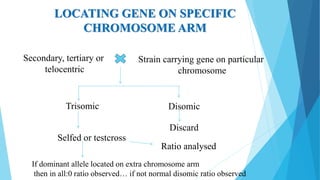
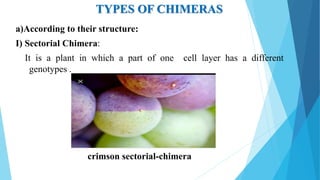
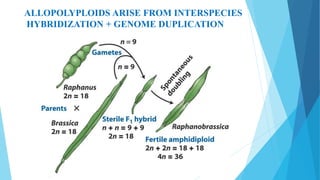
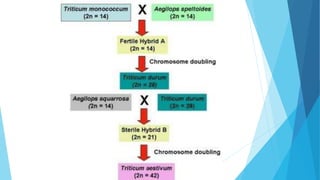
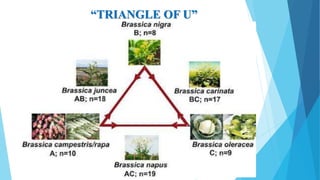
The document provides information about chromosomes and chromosomal aberrations. It begins by defining chromosomes as rod-shaped structures that carry genes and become visible during cell division. It then discusses two main types of chromosomal aberrations - structural aberrations which alter chromosome structure, and numerical aberrations which change chromosome number. Specific structural aberrations described include deletions, duplications, inversions, and translocations. Numerical aberrations include changes in ploidy levels such as polyploidy, as well as aneuploid conditions like trisomy and monosomy. The effects, origins, and uses of each type of aberration are summarized.