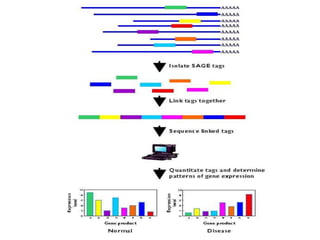
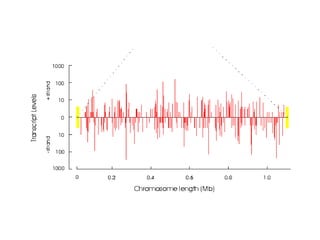
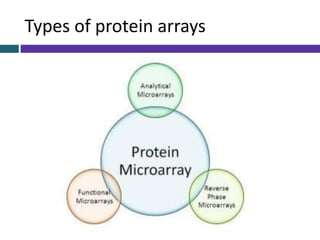
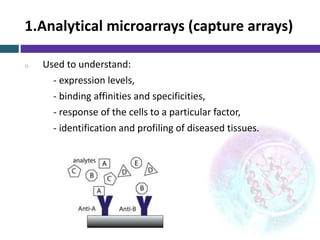
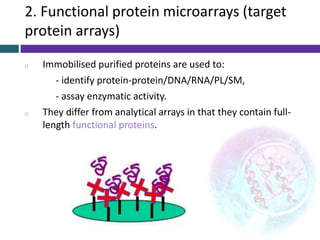
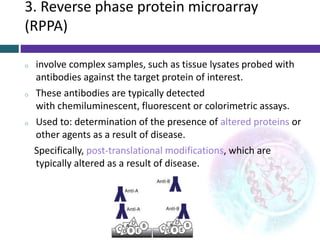
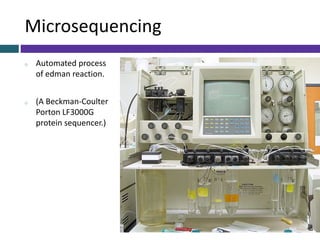
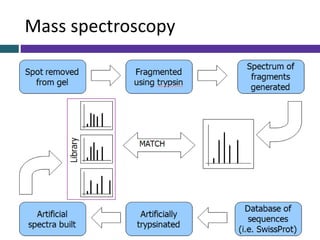
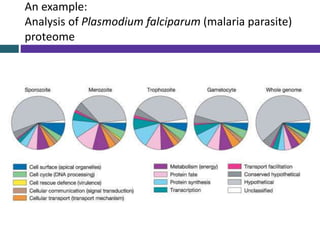
This document provides an overview of common proteomics techniques. It describes proteomics as the study of proteins including their roles, structures, localization, interactions and other factors. The key techniques discussed include molecular techniques like DNA microarrays and yeast two-hybrid analysis, separation techniques like gel electrophoresis and chromatography, protein identification methods like mass spectroscopy and Edman sequencing, and protein structure determination methods like NMR, X-ray crystallography and computational prediction. The document provides examples and details of several of these techniques.