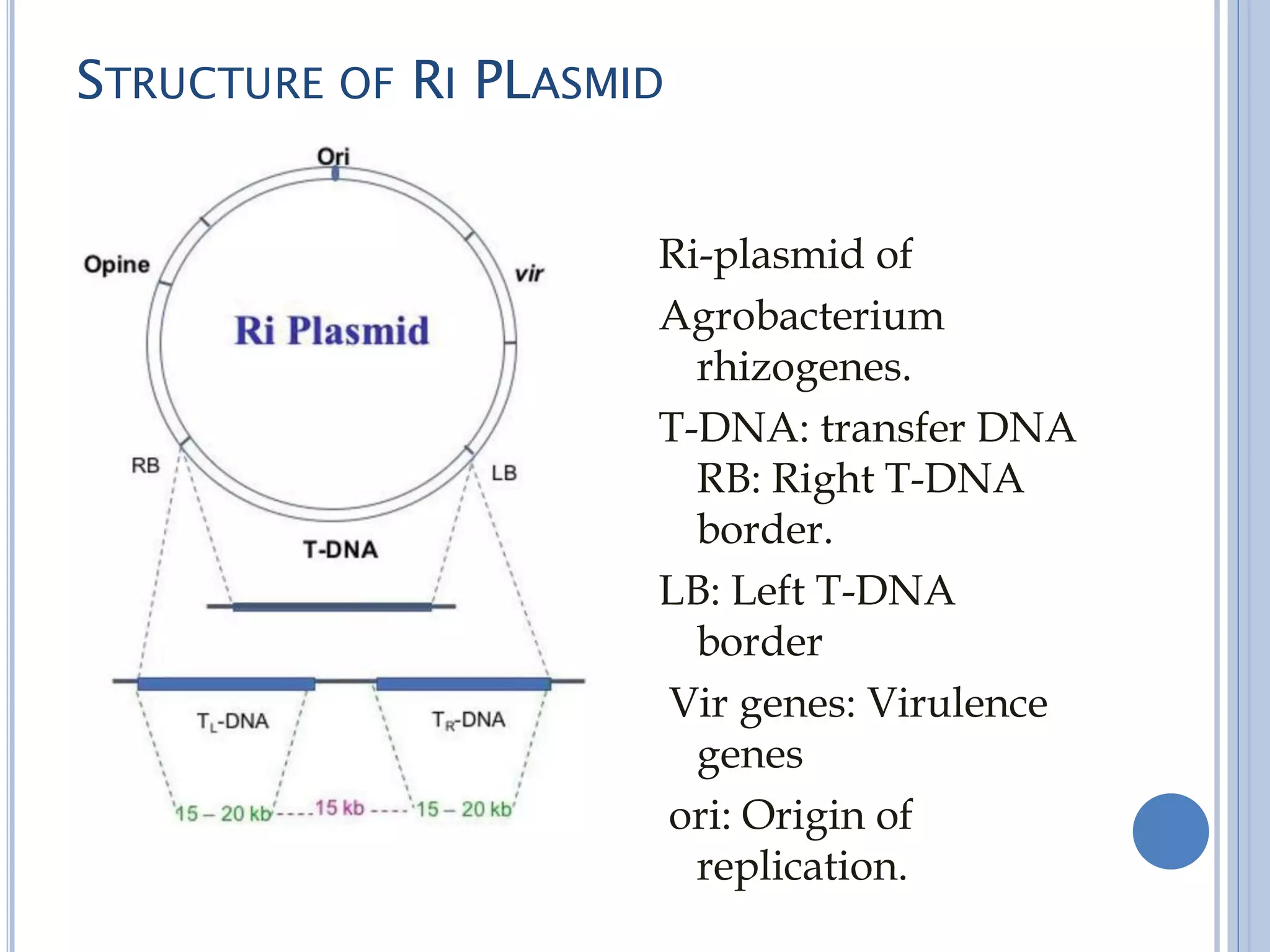
The Ri plasmid is a plasmid found in Agrobacterium rhizogenes bacteria that causes hairy root disease in plants. The plasmid contains genes that allow it to integrate portions of its DNA (T-DNA regions) into the plant genome. These integrated genes cause uncontrolled root growth and the formation of hairy roots. The Ri plasmid shares similarities with the Ti plasmid found in Agrobacterium tumefaciens, including virulence genes that mediate the transfer of T-DNA into plant cells and opine synthesis genes. Integration of the T-DNA from the Ri plasmid alters plant hormone production and induces hairy root formation.