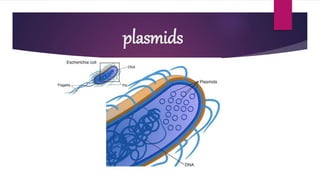
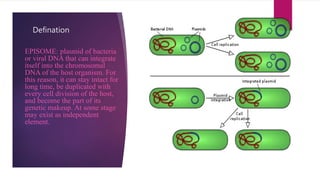
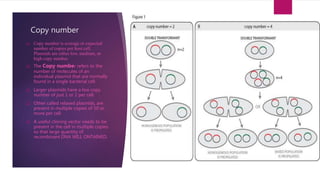
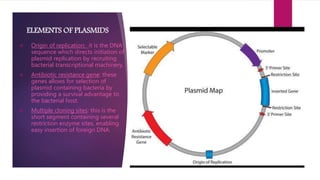
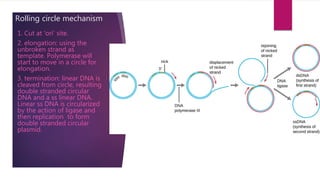
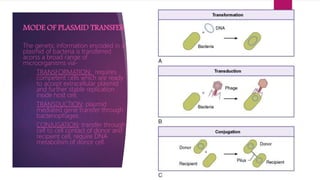
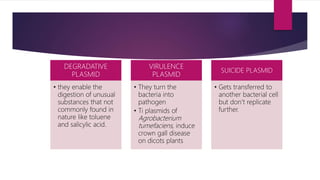
Plasmids are small, circular pieces of extrachromosomal DNA found in bacteria and other organisms. They can replicate independently of the chromosomal DNA and often contain genes that provide advantages to the host cell like antibiotic resistance. Key features of plasmids include an origin of replication, antibiotic resistance genes, and mechanisms for horizontal transfer between organisms like conjugation. Plasmids are important tools in biotechnology as they can be used to clone genes and amplify DNA fragments.