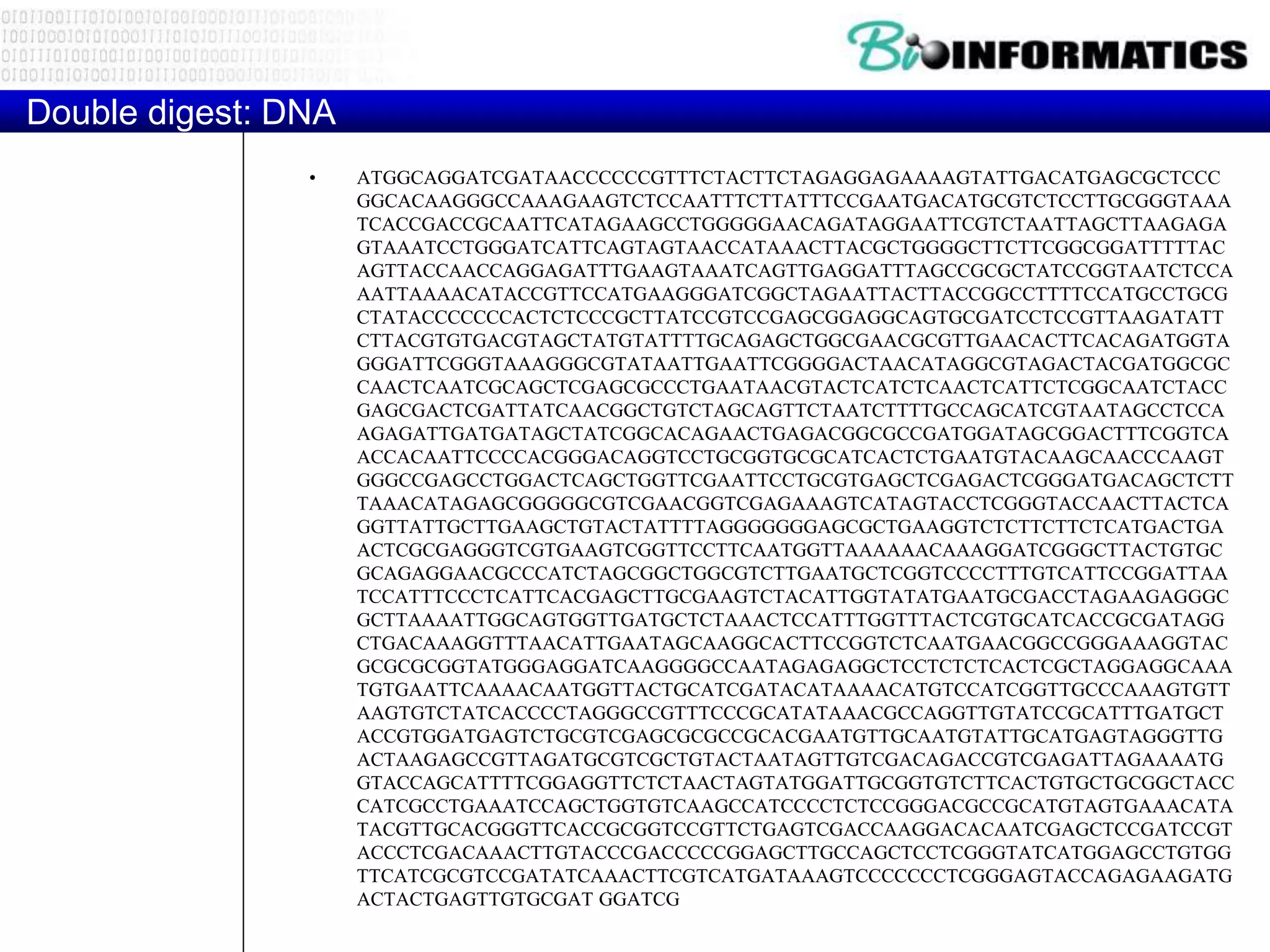
The document discusses regular expressions and provides examples of common regex patterns used for tasks like searching DNA sequences or scanning for protein domains. It then provides a sample of DNA/protein sequences and suggests using translated regex patterns to scan the sequences for specific consensus patterns representing protein domains.


![Recap
if condition:
statements
[elif condition:
statements] ...
else:
statements
while condition:
statements
for var in sequence:
statements
break
continue
Strings](https://image.slidesharecdn.com/p132017pythonexercises-171023184352/75/P1-3-2017_python_exercises-4-2048.jpg)
![Lists
• Flexible arrays, not Lisp-like linked
lists
• a = [99, "bottles of beer", ["on", "the",
"wall"]]
• Same operators as for strings
• a+b, a*3, a[0], a[-1], a[1:], len(a)
• Item and slice assignment
• a[0] = 98
• a[1:2] = ["bottles", "of", "beer"]
-> [98, "bottles", "of", "beer", ["on", "the", "wall"]]
• del a[-1] # -> [98, "bottles", "of", "beer"]](https://image.slidesharecdn.com/p132017pythonexercises-171023184352/75/P1-3-2017_python_exercises-5-2048.jpg)
![Dictionaries
• Hash tables, "associative arrays"
• d = {"duck": "eend", "water": "water"}
• Lookup:
• d["duck"] -> "eend"
• d["back"] # raises KeyError exception
• Delete, insert, overwrite:
• del d["water"] # {"duck": "eend", "back": "rug"}
• d["back"] = "rug" # {"duck": "eend", "back":
"rug"}
• d["duck"] = "duik" # {"duck": "duik", "back":
"rug"}](https://image.slidesharecdn.com/p132017pythonexercises-171023184352/75/P1-3-2017_python_exercises-6-2048.jpg)
![if condition:
statements
[elif condition:
statements] ...
else:
statements
while condition:
statements
for var in sequence:
statements
break
continue
Strings
REGULAR EXPRESSIONS](https://image.slidesharecdn.com/p132017pythonexercises-171023184352/75/P1-3-2017_python_exercises-7-2048.jpg)
![Regular Expression Quick Guide
^ Matches the beginning of a line
$ Matches the end of the line
. Matches any character
s Matches whitespace
S Matches any non-whitespace character
* Repeats a character zero or more times
*? Repeats a character zero or more times (non-greedy)
+ Repeats a chracter one or more times
+? Repeats a character one or more times (non-greedy)
[aeiou] Matches a single character in the listed set
[^XYZ] Matches a single character not in the listed set
[a-z0-9] The set of characters can include a range
( Indicates where string extraction is to start
) Indicates where string extraction is to end](https://image.slidesharecdn.com/p132017pythonexercises-171023184352/75/P1-3-2017_python_exercises-8-2048.jpg)


![Towards a protein prosite scanner
N-{P}-[ST]-{P}.
[RK](2)-x-[ST].
[ST]-x-[RK].
[ST]-x(2)-[DE].
[RK]-x(2,3)-[DE]-x(2,3)-Y.
G-{EDRKHPFYW}-x(2)-[STAGCN]-{P}.
x-G-[RK]-[RK].
C-x-[DN]-x(4)-[FY]-x-C-x-C.
E-x(2)-[ERK]-E-x-C-x(6)-[EDR]-x(10,11)-[FYA]-[YW].
[DEQGSTALMKRH]-[LIVMFYSTAC]-[GNQ]-[LIVMFYAG]-[DNEKHS]-S-[LIVMST]-{PCFY}-[STAGCPQLIVMF]-[LIVMATN]-[DENQGTAKRHLM]-[LIVMWSTA]-
[LIVGSTACR]-{LPIY}-{VY}-[LIVMFA].
[KRHQSA]-[DENQ]-E-L>.
R-G-D.
[AG]-x(4)-G-K-[ST].
D-{W}-[DNS]-{ILVFYW}-[DENSTG]-[DNQGHRK]-{GP}-[LIVMC]-[DENQSTAGC]-x(2)-[DE]-[LIVMFYW].
[EQ]-{LNYH}-x-[ATV]-[FY]-{LDAM}-{T}-W-{PG}-N.
[LIVM]-x-[SGNL]-[LIVMN]-[DAGHENRS]-[SAGPNVT]-x-[DNEAG]-[LIVM]-x-[DEAGQ]-x(4)-[LIVM]-x-[LM]-[SAG]-[LIVM]-[LIVMT]-[WS]-x(0,1)-[LIVM](2).
[FY]-C-[RH]-[NS]-x(7,8)-[WY]-C.
C-x-C-x(2)-{V}-x(2)-G-{C}-x-C.
C-x(2)-P-F-x-[FYWIV]-x(7)-C-x(8,10)-W-C-x(4)-[DNSR]-[FYW]-x(3,5)-[FYW]-x-[FYWI]-C.
[LIFAT]-{IL}-x(2)-W-x(2,3)-[PE]-x-{VF}-[LIVMFY]-[DENQS]-[STA]-[AV]-[LIVMFY].
[KRH]-x(2)-C-x-[FYPSTV]-x(3,4)-[ST]-x(3)-C-x(4)-C-C-[FYWH].
C-x(4,5)-C-C-S-x(2)-G-x-C-G-x(3,4)-[FYW]-C.
[LIVMFYG]-[ASLVR]-x(2)-[LIVMSTACN]-x-[LIVM]-{Y}-x(2)-{L}-[LIV]-[RKNQESTAIY]-[LIVFSTNKH]-W-[FYVC]-x-[NDQTAH]-x(5)-[RKNAIMW].
C-x(2,4)-C-x(3)-[LIVMFYWC]-x(8)-H-x(3,5)-H.
L-x(6)-L-x(6)-L-x(6)-L.
C-x(2)-C-x(1,2)-[DENAVSPHKQT]-x(5,6)-[HNY]-[FY]-x(4)-C-x(2)-C-x(2)-F(2)-x-R.
[LIVMFE]-[FY]-P-W-M-[KRQTA].
L-M-A-[EQ]-G-L-Y-N.
IRED_1R-P-C-x(11)-C-V-S.
[RKQ]-R-[LIM]-x-[LF]-G-[LIVMFY]-x-Q-x-[DNQ]-V-G.
[KR]-x(1,3)-[RKSAQ]-N-{VL}-x-[SAQ](2)-{L}-[RKTAENQ]-x-R-{S}-[RK].
[LIVMF](2)-D-E-A-D-[RKEN]-x-[LIVMFYGSTN].
[KRQ]-[LIVMA]-x(2)-[GSTALIV]-{FYWPGDN}-x(2)-[LIVMSA]-x(4,9)-[LIVMF]-x-{PLH}-[LIVMSTA]-[GSTACIL]-{GPK}-{F}-x-[GANQRF]-[LIVMFY]-x(4,5)-[LFY]-x(3)-
[FYIVA]-{FYWHCM}-{PGVI}-x(2)-[GSADENQKR]-x-[NSTAPKL]-[PARL].
Scan for the following prosite patterns in your 4 sequences
Hint: translate the patters to regexes and then scan](https://image.slidesharecdn.com/p132017pythonexercises-171023184352/75/P1-3-2017_python_exercises-12-2048.jpg)
![reuse galacto.py in github
Consensus_pattern="G-R-x-N-[LIV]-I-G-[DE]-H-x-D-Y"
pattern=Consensus_pattern.replace("-","")
pattern=pattern.replace("x","[A-Z]")
#print(pattern)
count=0
for s in sequences:
count=count+1
print ("searching seq",count)
s=s.replace(" ","")
matches = re.finditer(pattern,s)
for match in matches:
print (match.group(0),"from: ",match.start(),"to: ",match.end())](https://image.slidesharecdn.com/p132017pythonexercises-171023184352/75/P1-3-2017_python_exercises-13-2048.jpg)
