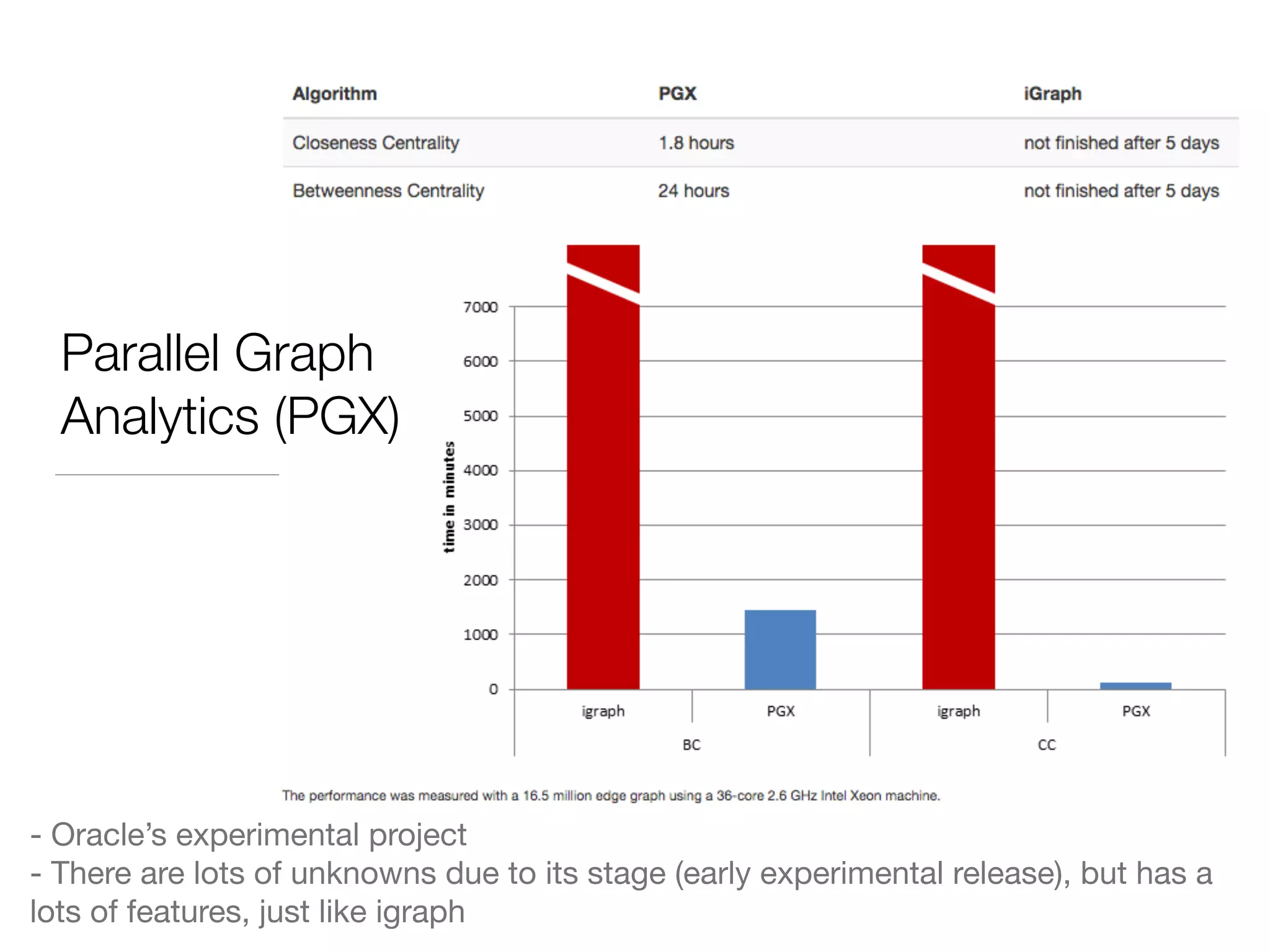
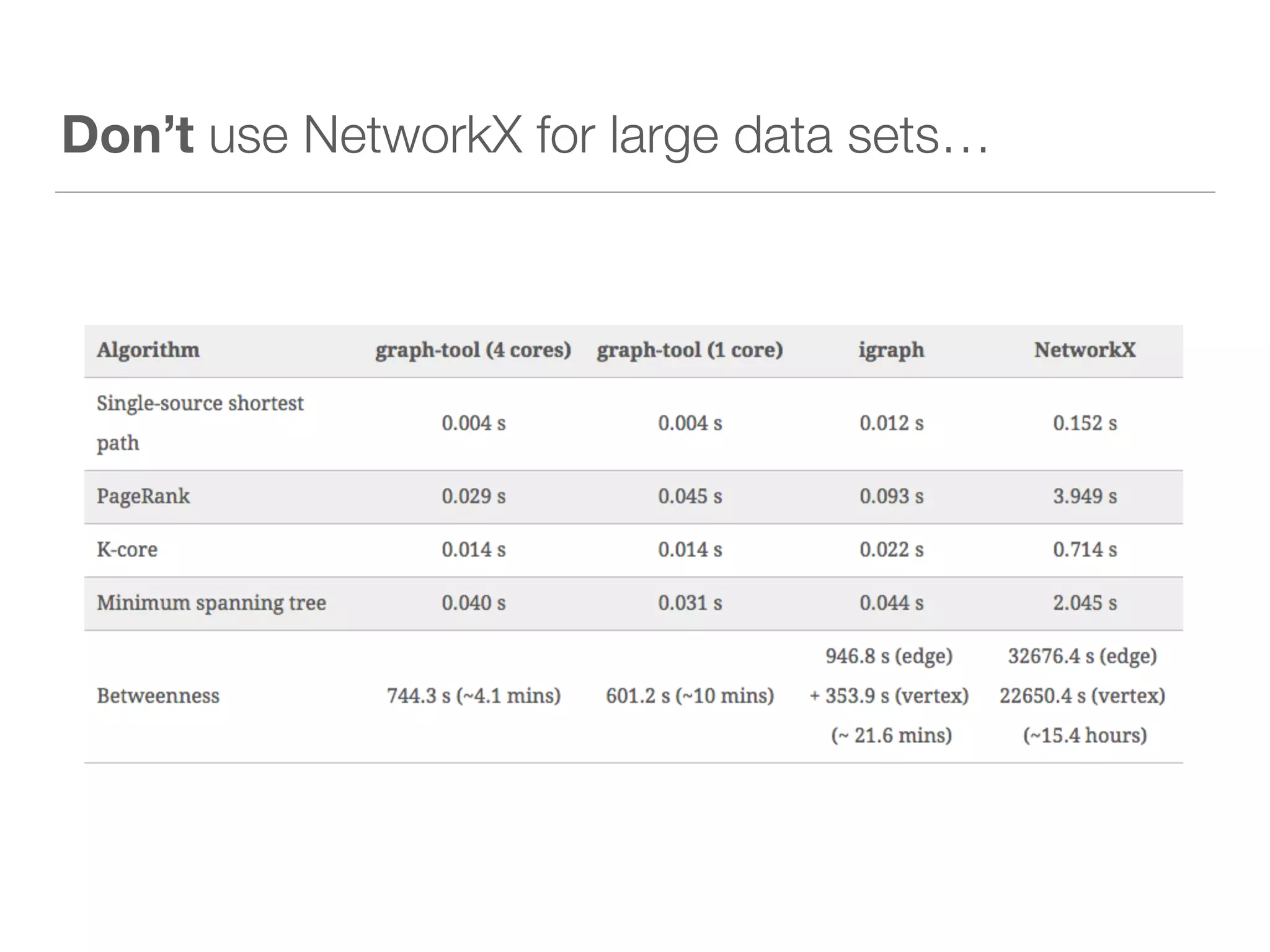
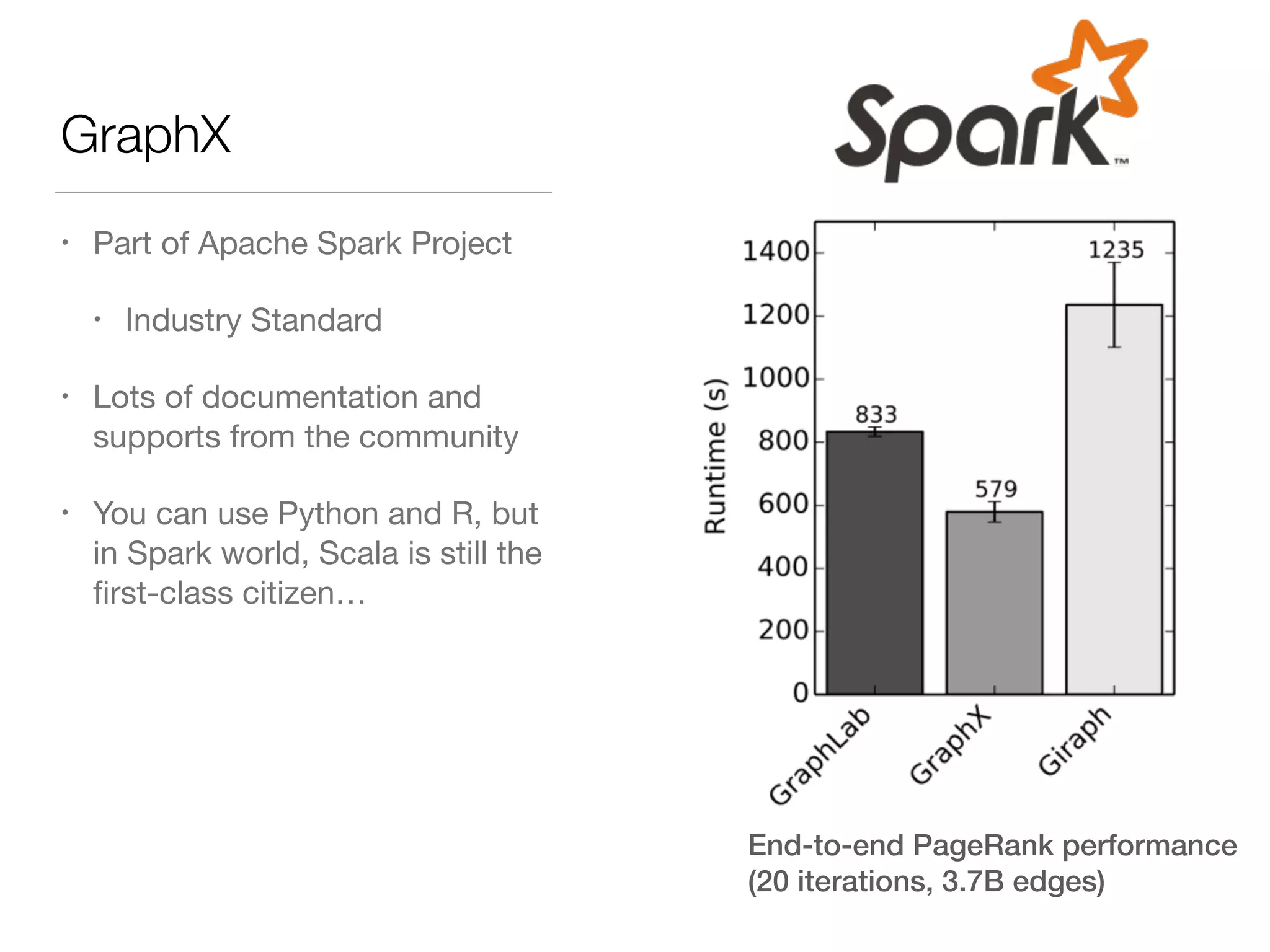
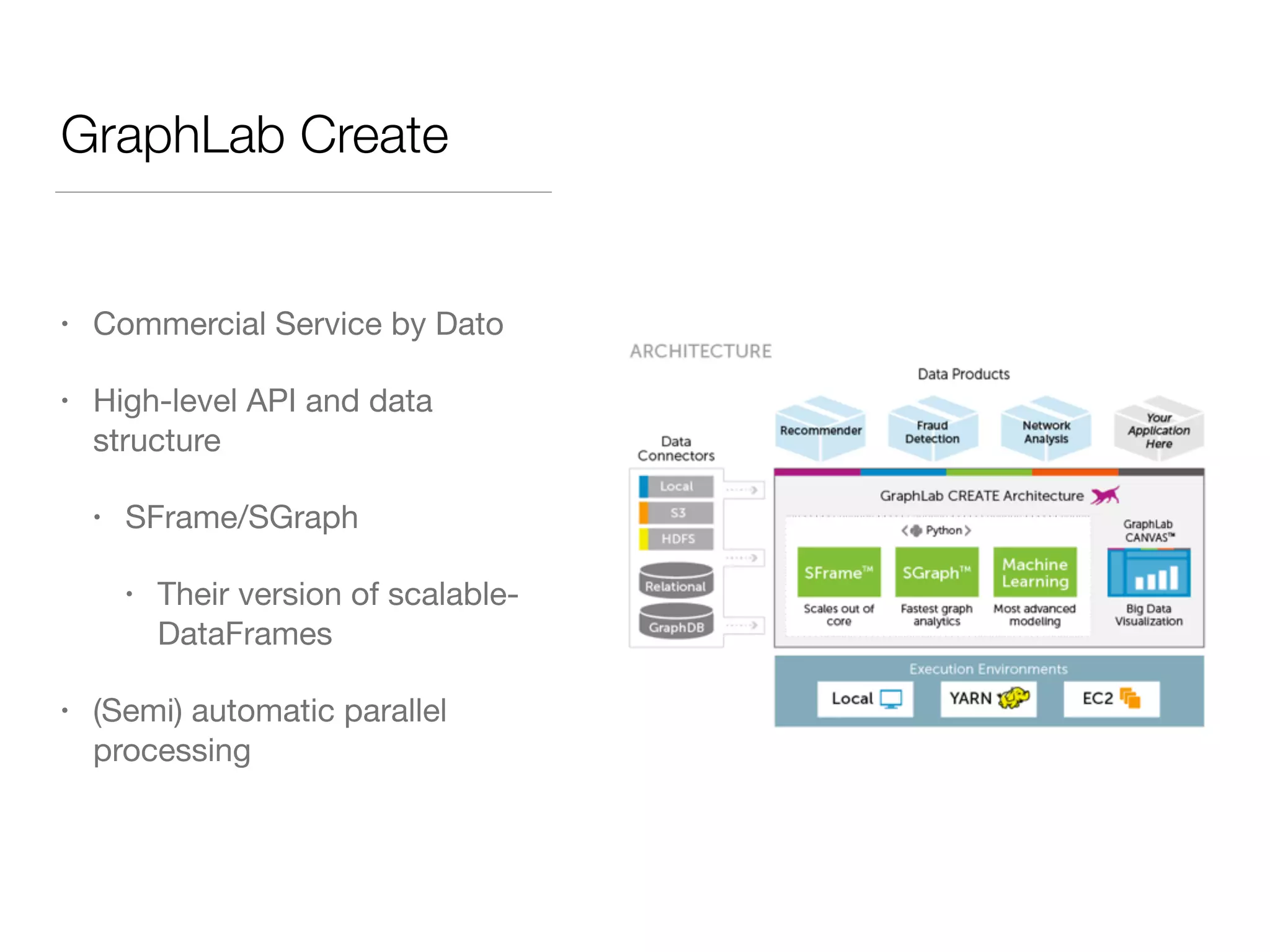
This document discusses modern tools for graph analysis and making graph workflows reproducible. It introduces cyREST, a RESTful API for programmatic access to Cytoscape, and language-specific wrappers like RCy3 and py2cytoscape that provide natural APIs. These tools allow running Cytoscape workflows in notebooks and remote machines. It also covers graph libraries for analysis like NetworkX, igraph, graph-tool, and PGX for smaller graphs, and distributed frameworks like GraphX, GraphLab Create, and Neo4j for extremely large graphs with billions of nodes. The document recommends not using NetworkX for large data and considering cloud-based options for difficult to install tools.