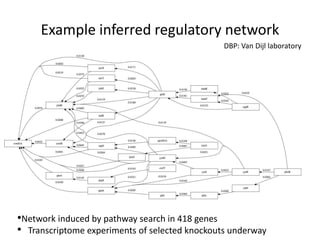
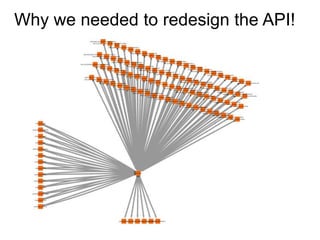
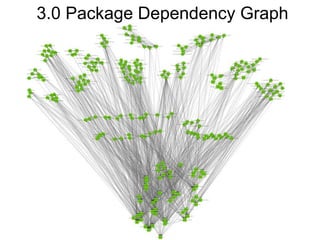
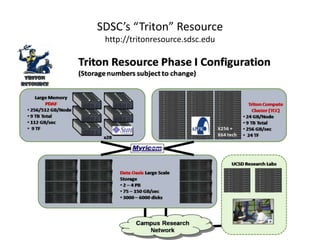
The National Resource for Network Biology aims to provide freely available, open-source software tools to enable researchers to assemble biological data into networks and pathways and use these networks to better understand biological systems and disease; it pursues this mission through technology research and development projects, driving biological projects, collaboration and service projects, training, and dissemination; key components include the Cytoscape software platform, supercomputing infrastructure, and partnerships with over 30 external research groups.