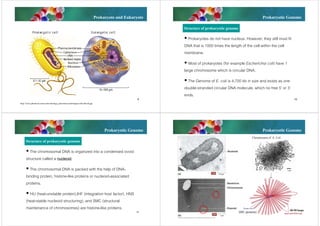
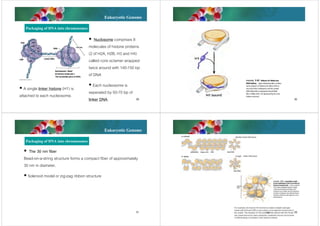
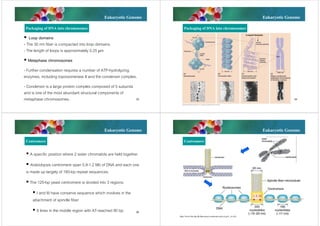
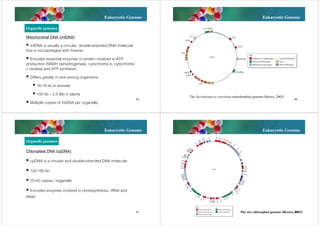
The document discusses genomes of organisms. It describes that genomes vary in size and can be DNA or RNA. It then compares prokaryotic and eukaryotic genomes. Prokaryotic genomes like E. coli are usually circular and highly condensed, organized into loops within the nucleoid. Eukaryotic genomes contain nuclear and organelle genomes. The nuclear genome is linear and packaged into chromatin and chromosomes using histones. Organelle genomes like mitochondria and chloroplasts resemble bacterial genomes. The document also discusses features like centromeres, telomeres, and gene organization within genomes.