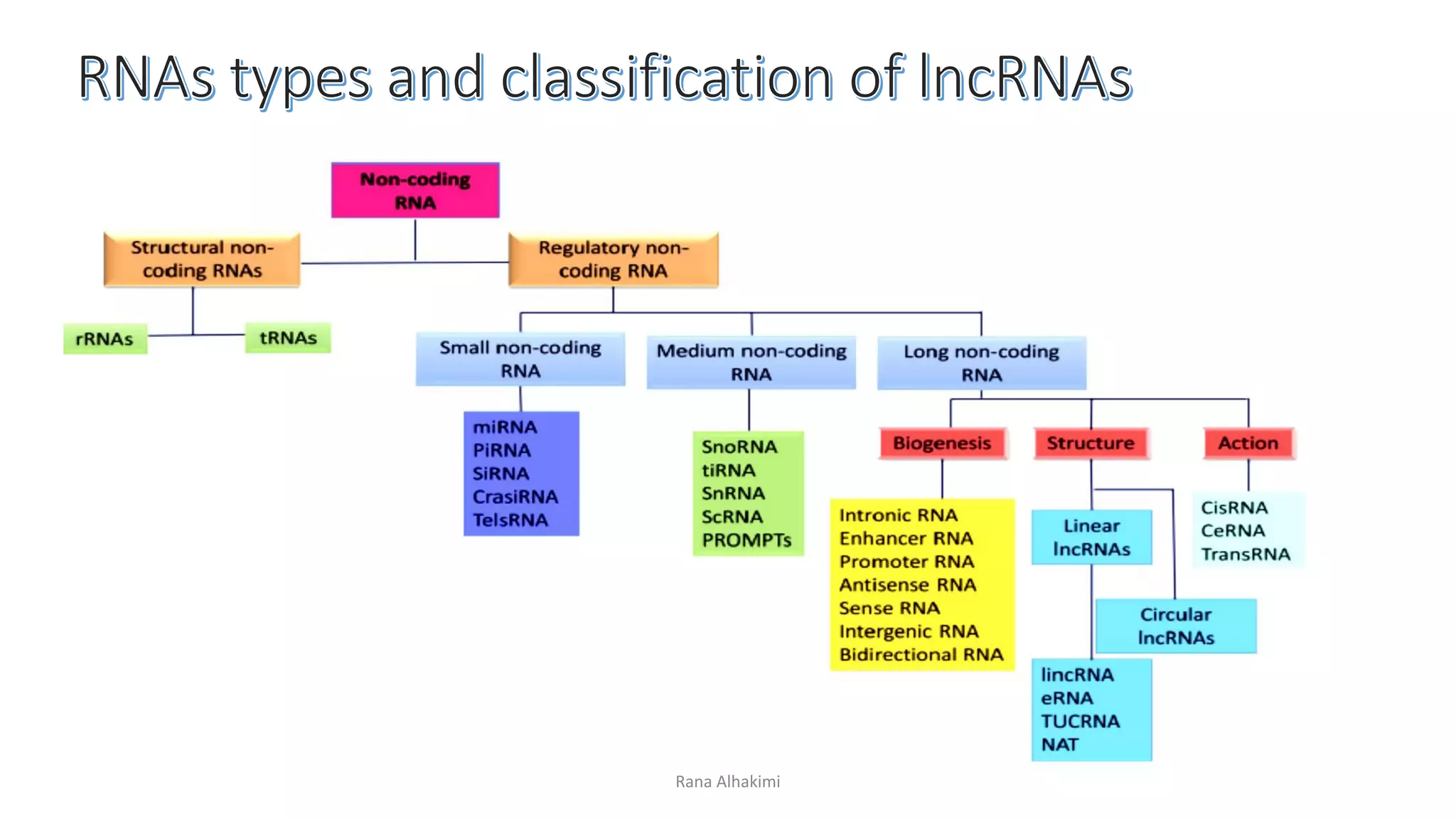
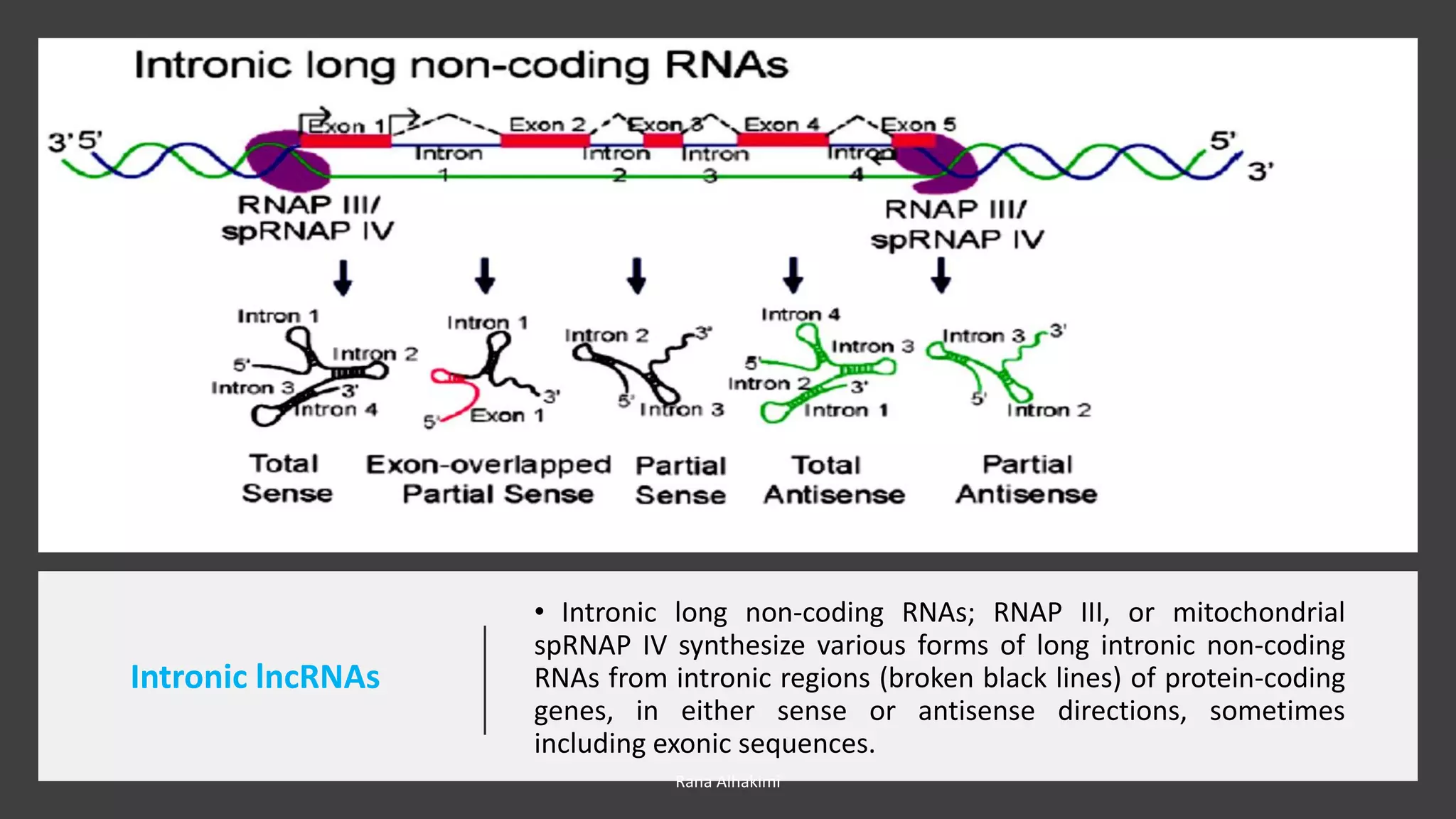
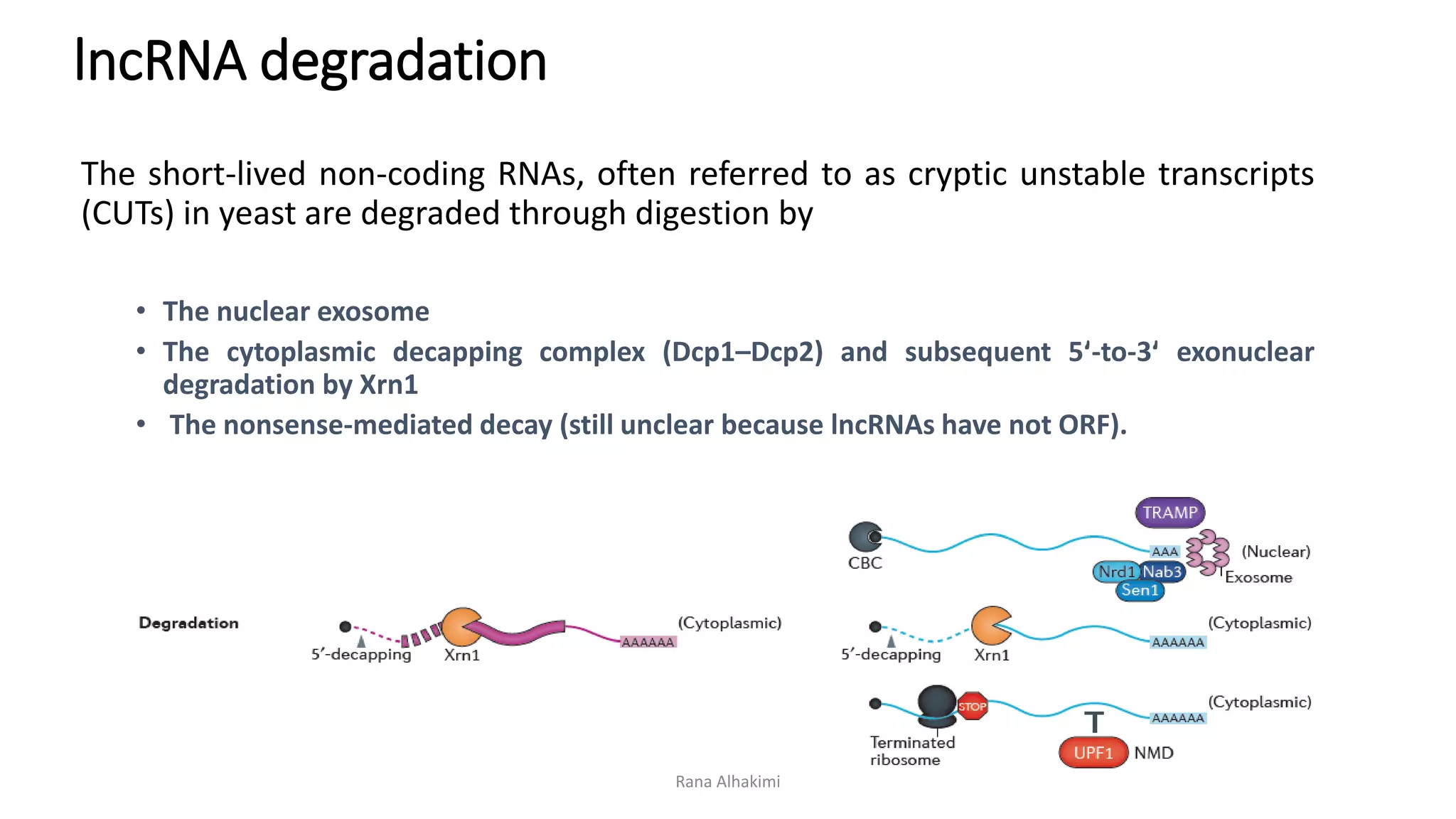
Long non-coding RNAs (lncRNAs) comprise over 200 base pairs of RNA that do not encode proteins and play essential roles in gene expression regulation and cellular processes. They are classified into various categories based on their genomic location and have structural and functional similarities with mRNAs, though they lack open reading frames and exhibit differences in expression and localization patterns. lncRNAs participate in mechanisms such as chromatin remodeling, transcriptional and post-transcriptional regulation, and serve as molecular decoys, signals, and scaffolds in cellular functions.