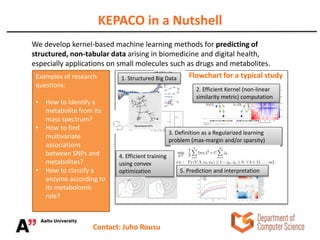
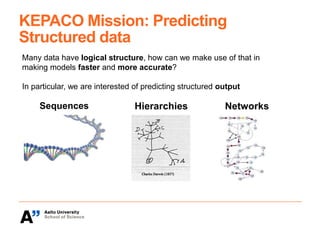
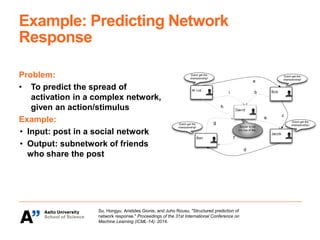
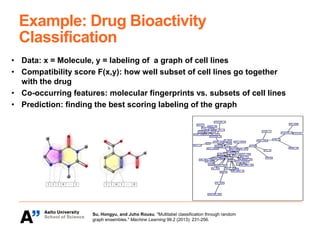
The Kepaco research group, led by Associate Professor Juho Rousu, focuses on developing kernel-based machine learning methods to analyze structured data in biomedicine and digital health, particularly regarding small molecules like drugs and metabolites. They work on various research problems, such as metabolite identification, drug bioactivity classification, and multivariate association meta-analysis, utilizing advanced computational techniques for better prediction and interpretation. Funding for their initiatives comes from multiple sources, including the Academy of Finland and EU projects.