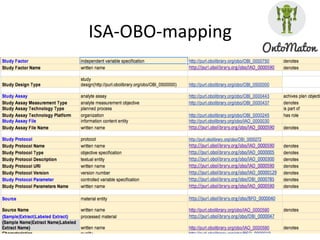
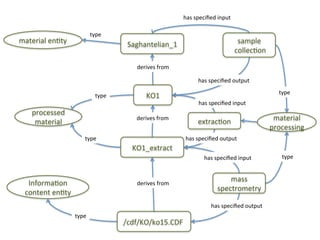
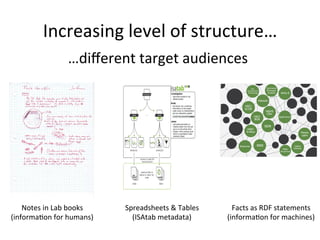
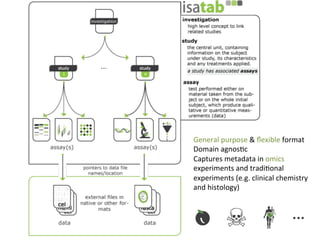
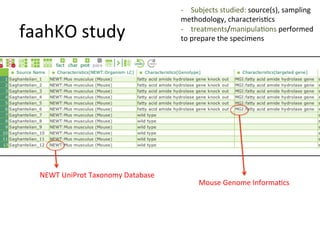
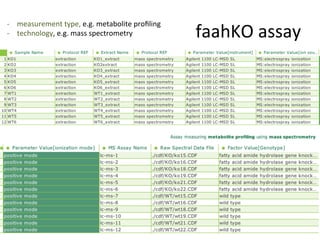
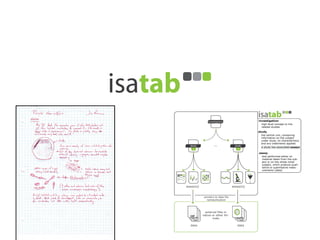
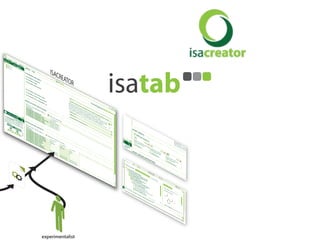
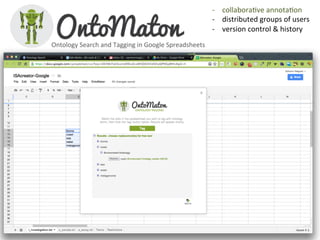
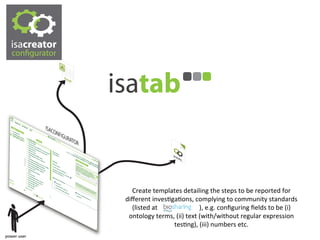
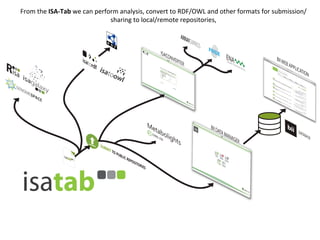
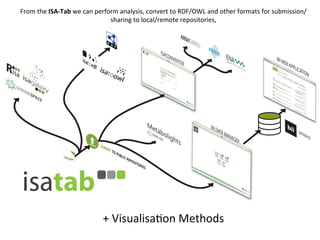
The document discusses knowledge management of experimental data through the ISA ecosystem. It describes the ISA-tab format and software suite that allows annotation and curation of experimental metadata. As a use case, it analyzes a dataset on metabolite profiling from a study of fatty acid amide hydrolase knockout mice. The ISA tools can represent investigations and assays, convert data to standardized formats, and facilitate sharing and analysis of experimental data.













![• faahKO
package
v.
2.12
contains
ISAtab
files
describing
the
experiment
faahkoISA
=
readISAta(find.package("faahKO"))
assay.filename
<-‐
faahkoISA["assay.filenames"][[1]]
xset
=
processAssayXcmsSet(faahkoISA,
assay.filename)
…
updateAssayMetadata(faahkoISA,
assay.filename,"Derived
Spectral
Data
File","faahkoDSDF.txt"
)
• MTBLS2
processing
and
analysis
using
Risa,
xcms
and
CAMERA
BioConductor
packages
Metabolights – an open access general-purpose repository for
metabolomics studies and associated meta-data
Haug et al, 2012
Nucleic Acids Research](https://image.slidesharecdn.com/agb-nettab2012-121116043134-phpapp02/85/NETTAB-2012-25-320.jpg)
![ISA
syntax
&
Underlying
Material/Data
workflows
Input
Material
or
Output
Material
or
Data
Node
Data
Node
Characteris9cs[…]
Factor
Value[…]
Characteris9cs[…]
Factor
Value[…]
Protocol
REF
Parameter
Value
[…]
26](https://image.slidesharecdn.com/agb-nettab2012-121116043134-phpapp02/85/NETTAB-2012-26-320.jpg)