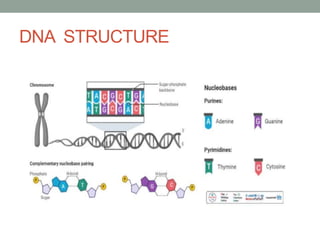
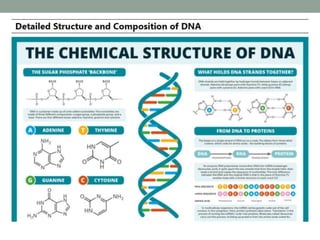
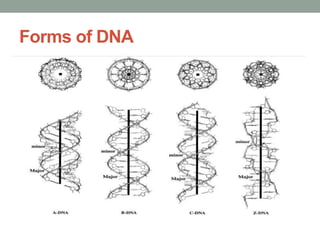
DNA carries genetic instructions in all living things. It is a double-stranded molecule shaped like a twisted ladder. Each strand has a backbone made of sugar and phosphate, with nitrogen bases (A, T, C, G) forming rungs between the strands via hydrogen bonding. The structure was discovered in 1953 by Watson and Crick. DNA is found in the cell nucleus and mitochondria, where it stores and transmits hereditary information from parents to offspring that directs protein synthesis and cell functions.