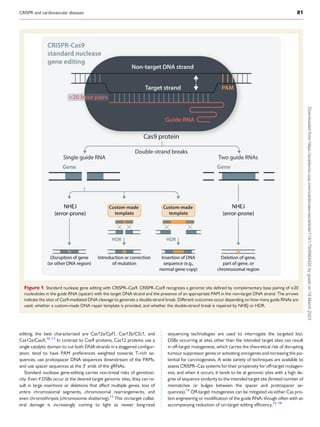
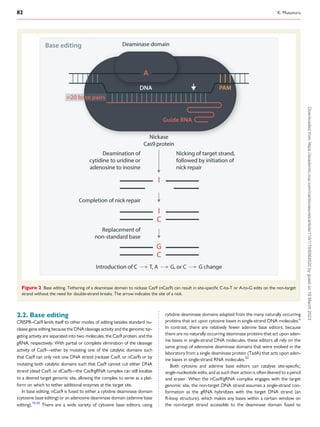
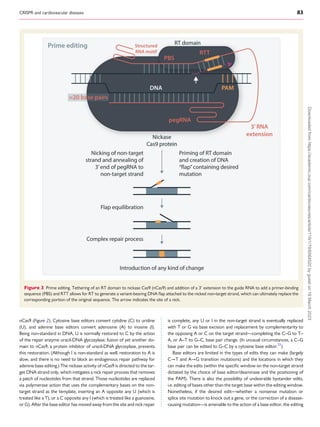
The review discusses advances in CRISPR technologies over the past decade, emphasizing their transformative impact on cardiovascular disease research and emerging therapies. Various CRISPR techniques, such as standard nuclease gene editing and base editing, are detailed, showcasing their applications in the treatment of conditions like hypercholesterolemia and transthyretin amyloidosis. The significance of careful consideration of off-target effects and gene editing precision is also highlighted.