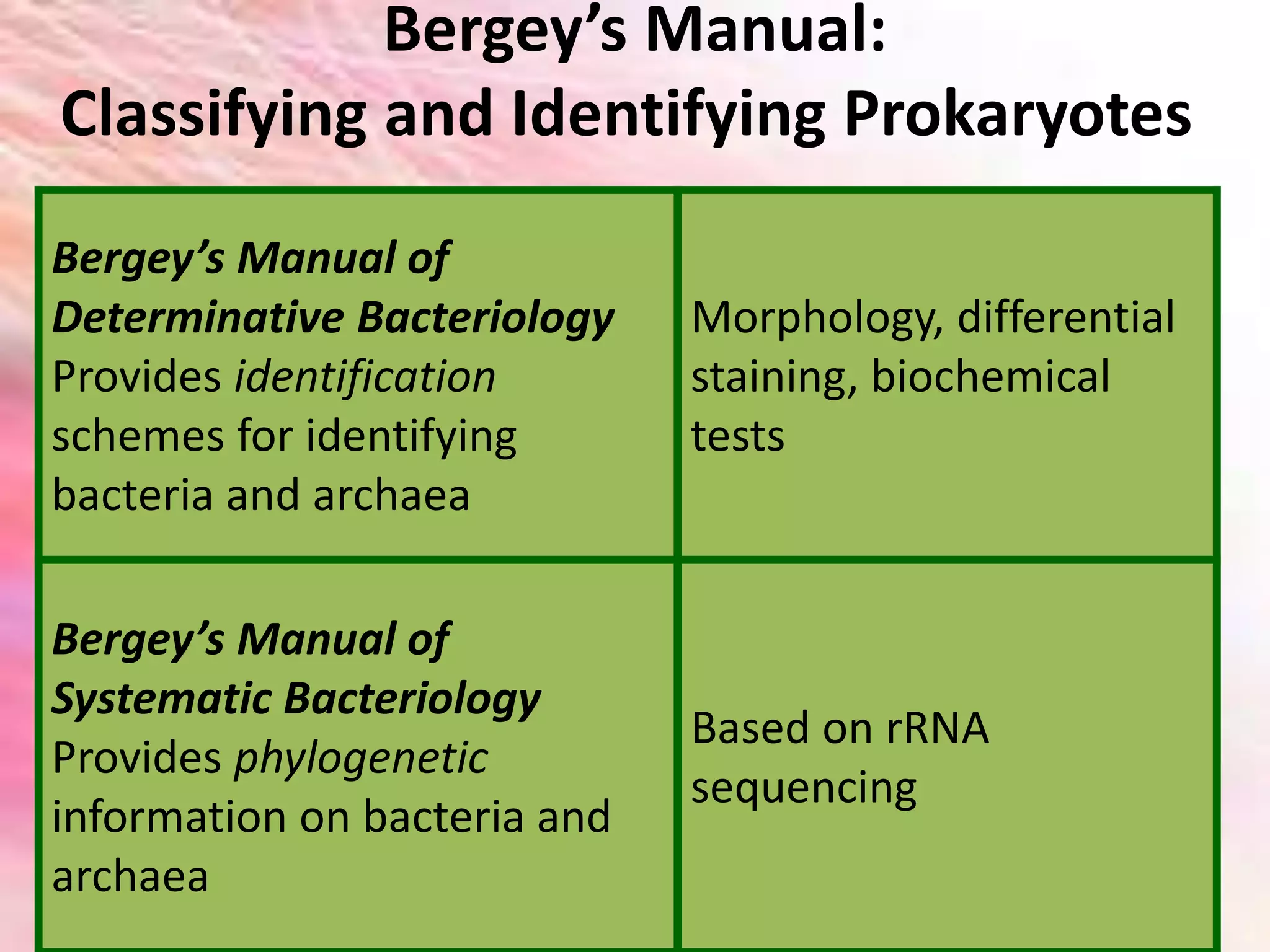
Bergey’s Manual is a key reference work for classifying and identifying prokaryotes. It consists of two parts: Bergey’s Manual of Determinative Bacteriology, which provides identification schemes based on morphology and biochemical tests; and Bergey’s Manual of Systematic Bacteriology, which provides phylogenetic classification based on rRNA sequencing. The first edition in 1923 established bacterial classification for identification. Current editions are based on phylogenetic studies and distribute pathogenic bacteria throughout volumes organized by phylogenetic relationships rather than clinical importance.