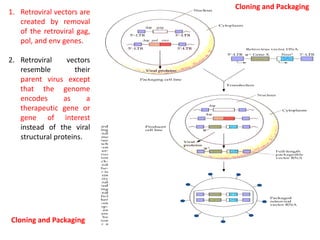
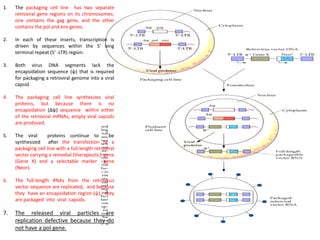
1. Bovine papillomavirus (BPV) vectors utilize the circular, double-stranded DNA genome of BPV. The BPV genome contains early and late regions and can transform cells without integrating.
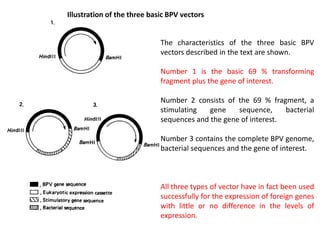
2. There are three main types of BPV vectors. All contain the transforming 69% fragment of BPV and bacterial sequences. One type inserts a gene of interest, another adds a stimulating gene, and the third uses the full BPV genome.
3. Transformation efficiency is highest with the full BPV genome due to an enhancer in the non-transforming region. Stimulating genes can replace this enhancer's function when parts of the genome are removed. BPV vectors provide amplified