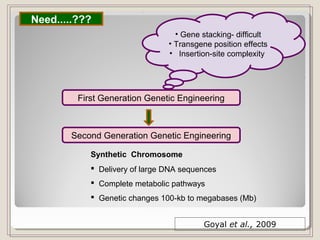
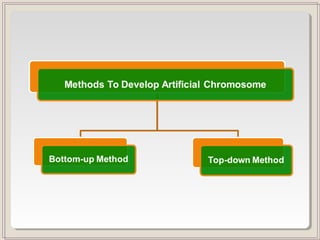
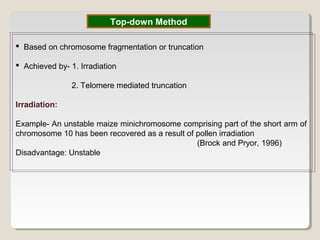
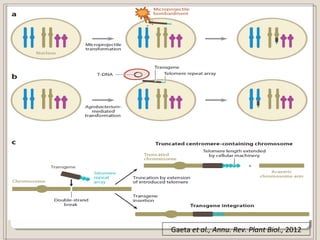
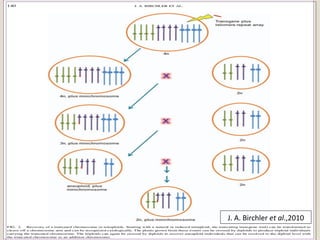
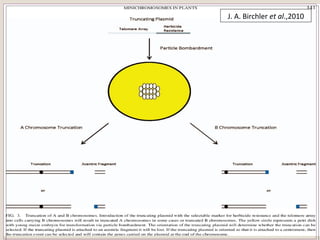
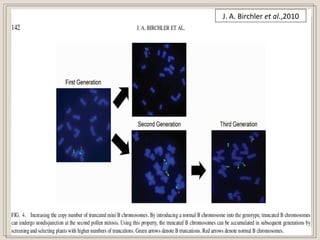
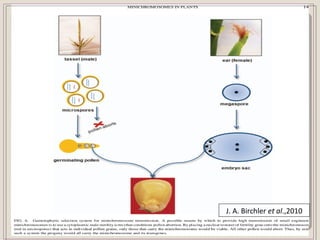
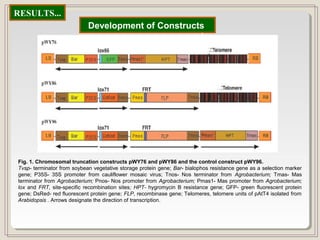
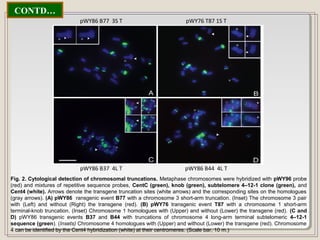
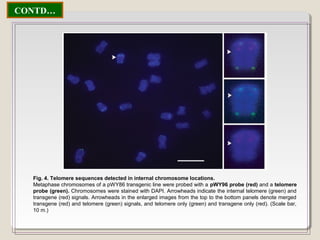
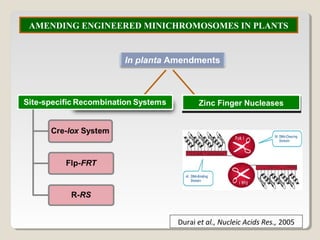
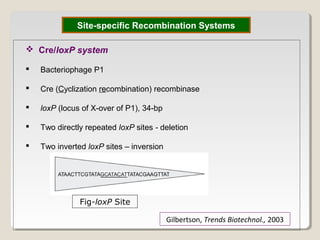
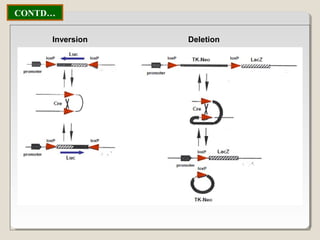
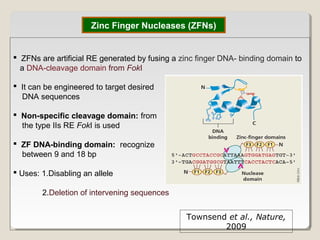
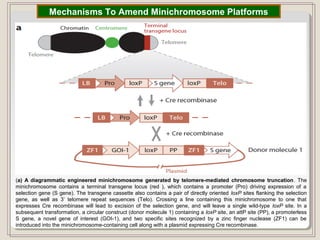
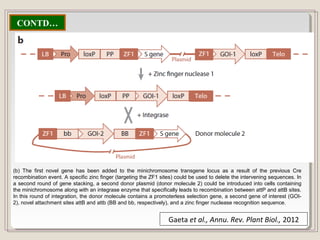
This document outlines a seminar on synthetic/artificial chromosomes. It begins by discussing the need for synthetic chromosomes due to limitations of traditional genetic engineering approaches. It then describes various methods that have been used to develop artificial chromosomes, including the bottom-up and top-down methods. Several case studies are presented where telomere-mediated truncation was used to produce engineered minichromosomes in plants with endogenous centromeres. The document concludes by discussing ways that engineered minichromosomes can be amended in planta, such as through site-specific recombination systems and zinc finger nucleases.