Genetic Engineering presentation
- 1. Chloroplast Transformation MBB 602 – Advances in Genetic engineering (3+0)
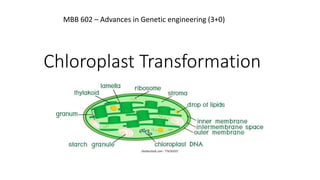
- 2. INTRODUCTION • Chloroplast word derived from Greek word : chloros – green and plast – organelle or cell. • One of the form of a plastid such as Amyloplast for storing starch, Elaioplast for fat storage, Chromoplast for pigment synthesis and storage. • Derived from proplastid. • Present in plant cell and capture light energy from sun to produce free energy via photosynthesis. (Bock, 2001)
- 3. • Transplastomic technologies allow precise targeted integration of trait genes into chloroplasts without marker genes. • Industrial and therapeutic proteins expressed in chloroplasts accumulate to extraordinarily high levels providing an attractive production platform for manufacture of high-value products for industry and health ,which is both sustainable and carbon-neutral. • Maternal inheritance of chloroplast genes prevents the pollen-mediated spread of transgenes providing a natural form of gene containment for the next generation of Biotech crops .
- 5. HISTORY • Theory of chloroplast inheritance, 1909, by Erwin baur, in Pelargonium and Mirabilis. • Presence of unique DNA in chloroplast, 1963, by Ruth sagar and Masahiro R. Ishida , in Chlamydomonas. • Circular DNA molecule in chloroplast,1971, by Manning et al., in Euglena. • Physical map of chloroplast,1976, by Jhon R. Bedbrook & Lawrence Bogorad, in Maize. • Physical map of Tobacco chloroplast DNA,1980, by Jurgenson & Bourque.
- 6. • First chloroplast transformation, 1988, by Boynton and Gillham , in Chlamydomonas. • First chloroplast transformation in higher plants, 1990, by Pal Maliga & coworkers, in Tobacco. • Plants with transformed plastid genomes are termed transplastomic.
- 8. Genome organization • Gene content highly conserved. • Semiautonomous organelles. • Highly polyploid genome. • Circular double stranded DNA. • Prokarotically organized. • 120-200kb size with 100–250 genes • tetrapartite genome organization, with a large single-copy region (LSC) and a small single-copy region (SSC) separating two inverted repeat regions. • Most plant genome have two identical copies of 20 to 30 kb inverted repeats separating a large single copy and small copy region. (Barbrook et al. 2010)
- 9. • High-level of Transgene Expression, • Multi-gene Engineering in a Single Transformation Event • Transgene Containment via Maternal Inheritance • Lack of Gene Silencing • No Position and Pleiotropic Effects • No Undesirable Foreign DNA ADVANTAGES OF CHLOROPLAST TRANSFORMATION
- 11. Different Transformation system • Direct DNA delivery methods such as particle gun or Biolistic method. • PEG mediated • Microinjection techniques. • Embryogenesis. • Organogenesis via protoplast or leaves.
- 16. UNIVERSAL VECTOR VERSUS SPECIES-SPECIFIC CHLOROPLAST VECTORS • “Universal vector’’ contains the trnA and trnI genes from the IR region of the tobacco chloroplast genome as flanking sequences for homologous recombination to transform several other plant species (of unknown genome sequence) was suggested several years ago (Daniell et al., 1998). • Vectors designed for transformation of the tobacco plastid genome have been successfully used for potato and tomato plastid transformation, because the homologous flanking sequences present in these vectors showed adequate homology to the corresponding sequences of potato and tomato plastid DNA but the efficiency of transformation is significantly lower than tobacco. • Concept of a universal vector is applicable when a higher level of homology exists among plant species but will be less efficient than species-specific chloroplast vectors.
- 17. Selectable Markers • Historically, chloroplast transformation was developed by using endogenous chloroplast sequences as selectable markers. • At the beginning, selection of plastid transformants was carried out by spectinomycin resistance encoded in the mutant 16S ribosomal RNA (rRNA) gene. • Stable integration and expression of the aadA gene was first reported in the chloroplast genome of Chlamydomonas (Goldschmidt-Clermont, 1991). • The neo gene is another alternative marker for plastid transformation that confers kanamycin resistance (Carrer et al. 1993) • “Double barrel’’ vector was used for cotton transformation where explant for transformation was nongreen cells (Kumar et al. 2004)- at least 8-fold more efficient than single gene based chloroplast vectors.
- 18. aadA gene • This gene encodes an aminoglycoside 3-adenylyltransferase, an enzyme that inactivates several antibiotics of the aminoglycoside type through covalent modification (i.e., attachment of an AMP residue). • Importantly, spectinomycin and streptomycin, two aminoglycoside antibiotics that act as potent inhibitors of plastid translation, are efficient substrates of the aadA enzyme. • Spectinomycin selection turned out to be particularly effective because of the high specificity of the drug to the chloroplast 70S ribosome and its low mutagenic and other side effects (Figure 3). • Although several alternative selectable marker genes have been developed over the years, the aadA gene is still superior to all of them.
- 19. Site of integration • Identification of spacer regions to integrate transgenes and endogenous regulatory sequences that support optimal expression is the first step in construction of chloroplast vectors. • Most commonly used site of integration is the transcriptionally active intergenic region between the trnI-trnA genes, within the rrn operon, located in the IR regions of the chloroplast genome. • The foreign gene expression levels obtained from genes integrated at this site are among the highest ever reported (De Cosa et al., 2001). • oriA is present within the trnI flanking region and this facilitate replication of foreign vectors within chloroplasts (Daniell et al., 1990), enhance the probability of transgene integration, and achieve homoplasmy even in the first round of selection (Guda et al. 2000) • This is further confirmed by the first successful Rubisco engineering obtained by integrating the rbcS gene at this site (Dhingra et al., 2004).
- 20. Integration of Foreign DNA into the Plastid Genome
- 21. Confirmation of Chloroplast transformation • The confirmation is done by testing Homoplasmy or Heteroplasmy. • For complete transformation it is necessary to achieve homoplasmy in the plant. • To achieve homoplasmy, incorporation of multiple rounds of selection markers are done in DNA until wild type plastid genomes are replaced with newly modified genomes. Figure 1: Chloroplast transformation in Chlamydomonas (Blowers et al., 1989)
- 22. Generation of transplastomic potato (Solanum tuberosum) plants – selection pressure a. Preparation of leaves for biolistic transformation. b. Exposure of biolistically bombarded leaf explants to a spectinomycin containing regeneration medium. c. Selection of primary transplastomic lines d. Additional regeneration round under spectinomycin selection to purify the transplastomic line to homoplasmy e. Additional regeneration round initiated from stem sections. The efficiency of regeneration and purification to homoplasmy from stem explants is similar to that from leaf explants f. Growth of homoplasmic transplastomic plants under aseptic conditions on a synthetic medium.
- 24. AGRONOMIC TRAITS EXPRESSED VIA THE PLASTID GENOME • Plastomes offer an ideal place for agronomic gene expression because of their higher levels of expression, ease of transformation, and gene containment via maternal inheritance. • Many important crops have already been genetically modified via the nuclear genome, however, nuclear transgenic plants have drawbacks inherent to their method of transformation. • Although more than half of the soybean planted worldwide is genetically modified, restrictions have to be placed on the location of crops because of the risk of transgenes spreading to wild relatives. • Additionally, low expression levels of Bt toxins engineered via the nuclear genome increases the risk of developing Bt resistant pests
- 26. Herbicide Resistance • The most commonly used herbicide, glyphosate, is a broad spectrum, nonselective, systemic herbicide. When it is applied to crops it is absorbed by the leaves and rapidly moves throughout the plant. • Glyphosate inhibits the enzyme 5-enolpyruvylshikimate-3-phosphate synsthase (EPSPS), a nuclear-encoded gene involved in the biosynthesis of aromatic amino acids (Devine and Daniell, 2004).
- 27. • Transgenic chloroplast plants were engineered to express a wild-type petunia EPSPS by integration of the aroA gene between the trnI and trnA genes in the inverted repeat region or rbcL and accD genes in the large single copy region (Daniell et al., 1998). • Homoplasmic integration of aroA was confirmed via Southern blot analysis and resulted in resistance to up to 5 mM glyphosate, 10-fold greater than the lethal concentration
- 28. Insect Resistance • Crop plants genetically engineered to produce functional insecticides may reduce or omit the spraying (often six to eight times a growing season) of costly and hazardous chemical pesticides. • The biological pesticides typically engineered in plants are highly specific towards their target insects and are non- hazardous to humans and other higher animals. • Most approaches make use of the insecticidal protoxins produced by a variety of Bacillus thuringiensis subspecies (Daniell et al., 2002). • The Cry2Aa2 operon consists of the Cry2Aa2 gene, orf 1 and orf 2 genes. • The unique ability for Cry2Aa2 protein to fold into cuboidal crystals is a result of chaperone activity by orf 2.
- 29. • Tobacco chloroplasts were shown to correctly process a bacterial operon and express the Cry2Aa2 proteins at levels up to 46% tsp, the highest levels ever recorded (De Cosa et al., 2001). Tobacco budworms (Heliothis virescens) died within three days when fed leaves expressing the Bt operon. • Chloroplast transgenic plants were able to kill insects that could withstand insecticidal protein concentrations 40,000 times higher than normal, even at lower levels of Cry2Aa2 protein expression. • The high levels of Cry2Aa2 protein accumulation in transgenic chloroplasts thus offer complete resistance to insects that feed on transgenic plants.
- 31. TRANSGENIC PLASTIDS AS PHARMACEUTICAL BIOREACTORS • Mammalian cells - used for production of recombinant proteins; however, they are costly to culture and have no current option for large-scale setup. • E. coli is commonly used for larger scale industrial applications of recombinant protein production. • This method introduces several disadvantages such as the lack of glycosylation, proper folding, and formation of disulfide bonds. • Nuclear expression of foreign genes is often disappointingly low.
- 32. Human Somatotropin – Therapeutic protein • Human Somatotropin (hST) - to treat hypopituitary dwarfism in children, Turner’s syndrome, chronic renal failure, and HIV wasting syndrome. • The hST gene was integrated between the trnV gene and rps7/3-rps12 operon located within the inverted repeat region of tobacco chloroplasts (Staub et al. 2000). • This expression cassette resulted in high levels of hST expression, reaching 7 percent tsp. • Additionally, it was shown to contain proper disulfide bond formation and was identical to the native hST protein. • These results demonstrate that plastids have the proper machinery for folding eukaryotic proteins requiring disulfide bonds, possibly through the use of the chloroplast protein disulfide isomerase.
- 33. Human Serum Albumin • Human serum albumin (HSA) comprises approximately 60 percent of the protein found in blood serum at concentrations of up to 45 mg/ml, and is one of the most widely used intravenous proteins in many human therapies. • More than 500 tons of HSA is used annually, representing a global market value of more than $1.5 billion. • attempts using nuclear transformation for the expression of HSA have resulted in far below the levels needed for cost-effective production. • Under the control of the psbA regulatory sequences, the HSA gene was integrated into the chloroplast genome between the trnA and trnI genes of the chloroplast rrn operon, located within the inverted repeat regions of the tobacco chloroplast (Fern´andez-San Mill´an et al., 2003). • The amount of HSA was measured using ELISA and found to be up to 8.2% tsp in mature potted plants and at a maximum of 11.1% tsp after 50 hours of continuous illumination
- 34. Electron micrographs of immunogold-labeled HSA in tissues from untransformed (A) and transformed (B) mature leaves. Magnifications are 10,000× (A) and 5000× (B).
- 36. PLASTIDS AS BIOMATERIAL BIOREACTORS • Besides vaccine antigens, biomaterial and amino acids have also been expressed in chloroplasts. • p-hydroxybenzoic acid (pHBA) is produced in small quantities in all plants. • In E. coli, the ubiC gene encoding chorismate pyruvate lyase catalyzes the direct conversion of chorismate to pyruvate and pHBA. • However, in chloroplasts, chorismate is converted to pHBA by 10 consecutive enzymatic reactions due to lack of chorismate pyruvate lyase. • Stable integration of the ubiC gene into the tobacco chloroplast resulted in hyperexpression of the enzyme and accumulation of this polymer up to 25% of dry weight (Viitanen et al., 2004). • In another study, the gene for thermostable xylanase was expressed in the chloroplasts of tobacco plants (Leelavathi et al., 2003). Xylanase accumulated in the cells to approximately 6% of tsp. • Zymography assay demonstrated that the estimated activity was 140,755 units kg-1 fresh leaf tissue
- 37. Enhancing Photosynthetic Efficiency via the Chloroplast Genome • Rubisco (ribulose-1,5-bisphosphate carboxylase/oxygenase), the key enzyme in the Calvin cycle, has attracted attention for as a means to enhance carbon fixation efficiency, increase catalytic activity, and/or reduce photorespiration. • More recent studies have focused on the expression of heterologous Rubisco subunits in chloroplasts. • Most recently, a breakthrough was made by introducing the CO2- concentrating mechanism (CCM) from cyanobacteria into transplastomic plant. • The native tobacco gene encoding the large subunit of Rubisco was knocked out by inserting the large and small subunit genes of the Synechococcus elongates Se7942 enzyme. • Se7942 Rubisco and CcmM35 (a beta-carboxysomal protein) hybrid assembly within chloroplasts resulted in higher rates of CO2 fixation efficiency but slowed growth.
- 38. Marker gene excision by Phage Cre or Int site-specific recombinases
- 39. RNA editing in plastids • In plastid transformation, three approaches were used to test mRNA editing: minigenes, translational fusion with a reporter gene, and incorporation of an editing segment in the 3'-untranslated region (3'-UTR). • The most simple design was the construction of minigenes, which involved insertion of an editing fragment in a plastid expression cassette linked to a marker gene (Zoubenko et al., 1994). • The second approach, translational fusion with a reporter gene, was used to study the psbL and ndhD editing events that create an AUG translation initiation codon by editing of an ACG codon at the mRNA level ( Chaudhuri and Maliga, 1996). • The third approach to test editing was incorporation of editing segments in the 3'-UTR of the aadA marker gene where the editing status of the segment does not affect the expression of the marker gene. In plasmid pRB51, the editing segment can be conveniently cloned in an Xba I-BamH I fragment (Bock et al., 1996). • Future studies will focus on understanding the role of RNA editing in plastids. Since both edited and nonedited mRNAs are translated, RNA editing may serve as a regulatory mechanism leading to yielding multiple proteins from the same gene.
- 40. Engineering the Chloroplast Genome to Confer Biotic/Abiotic Stress-Tolerance or Expression of High-Value Products. A. Antimicrobial peptide retrocyclin-101 fused with GFP expressed in chloroplasts displays strong green florescence in contrast to untransformed leaves showing chlorophyll red fluorescence (B,C) The transplastomic leaf expressing Pinellia ternata agglutinin shows high-level tolerance when challenged with bacterial (Erwinia) or viral (tobacco mosaic virus, TMV) pathogens (D) Gel diffusion assay shows the zone of clearance of chloroplast-derived endo-b-mannanase in crude plant extracts similar to purified recombinant commercial enzyme (E) Tobacco chloroplast genome and integration of the expression cassette (F) Enhanced accumulation of astaxanthin and carotenoids in transplastomic lettuce (G) Transplastomic plants expressing the lectin gene show broad-spectrum resistance to lepidopteran and homopteran (sap-sucking) insects as well as antibacterial (Erwinia) and antiviral (TMV) activities. (H,K) Chloroplast expression of b-glucosidase results in elevated phytohormone levels associated with significant increase in biomass and trichome density. (I) Cotton bollworm larvae with normal pupation or dead larvae when fed with transplastomic tobacco leaves expressing dsRNAs. (J) Overexpression of g-tocopherol methyl transferase chloroplasts confers abiotic stress tolerance and nutritional enhancement.
- 41. Conclusion • Chloroplast genomes are extraordinary; they can express more than 120 foreign genes from different species and organisms like bacteria, fungi, virus animals even humans too. • There are many traits that can be commercially beneficial such as, herbicide and insect resistance, insertion of therapeutic genes in plants etc. • In future, chloroplast genome sequence can be proved very useful in the creation of herbal medicines, therapeutic plants biofuel production and many more. • Chloroplast transformation is proved to be a more eco-friendly and accurate technique for genetic modification in plants where nuclear transmission can cause severe ecological damage through dissemination of the transgene in the environment.
- 42. Thank you Balaji S 2020603002 I-Ph.D Plant Biotechnology