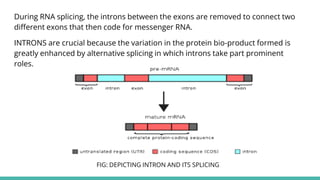
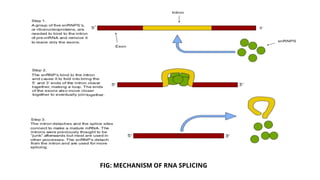
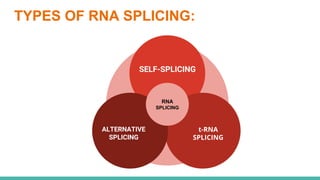
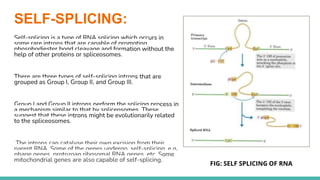
The document provides an extensive overview of RNA splicing, detailing its mechanisms, significance, and historical context, including an explanation of introns and exons. It describes types of RNA splicing such as self-splicing and alternative splicing, highlighting their roles in protein diversity and cellular function. The presentation concludes by emphasizing the importance of RNA splicing in gene expression regulation, evolutionary processes, and its applications in cancer research.