1. Protein is made up of amino acids that are linked together through peptide bonds to form polypeptide chains. The sequence and structure of these chains determines the protein's function.
2. There are four levels of protein structure: primary, secondary, tertiary, and quaternary. The primary structure is the sequence of amino acids. Secondary structures include alpha helices and beta sheets. Tertiary structure involves folding into a 3D shape. Quaternary involves multiple protein subunits.
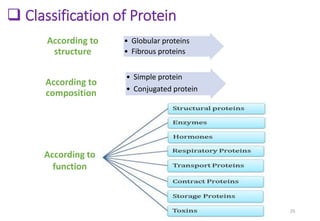
3. Examples of important proteins and their structures include hemoglobin, which has a quaternary structure to transport oxygen in red blood cells, and collagen which has a triple helix structure that provides structure and connective