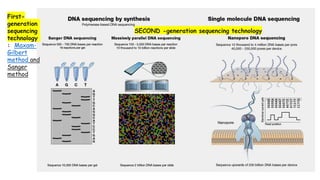
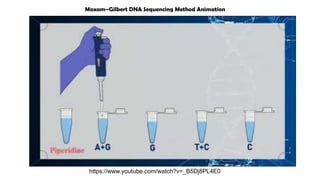
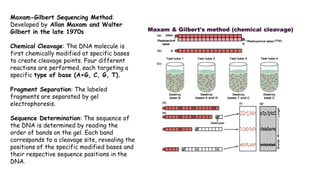
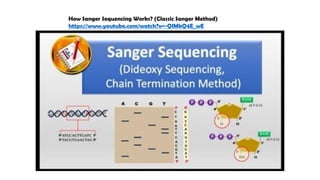
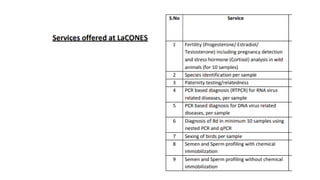
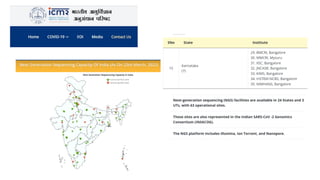
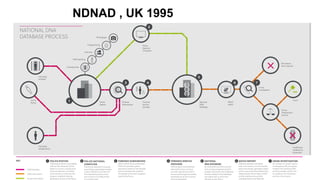
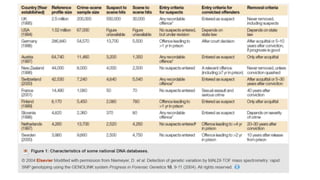
The document discusses DNA profiling and sequencing, detailing first-generation methods like Maxam-Gilbert and Sanger sequencing, and how these have paved the way for next-generation sequencing techniques. It also highlights the importance of DNA databases for crime detection, mentioning regulatory frameworks, the introduction of DNA technology bills in India, and the necessity for proper consent and procedures for DNA data collection and management. Additionally, it notes the advantages of Sanger sequencing over Maxam-Gilbert in terms of automation and throughput.