Embed presentation
Download as PDF, PPTX













![Repeat: [CA]n
Repeat: [AGAGAT]n[AGAGAT]m](https://image.slidesharecdn.com/mscivsemesterdnaprofiling-strbiologyandartifacts-240515082426-9bd80228/75/MSC-IV-SEMESTER_DNA-Profiling-STR-biology-and-artifacts-pdf-34-2048.jpg)
![Repeat: [TATC]
Repeat: [TCTA], [TCTG] Repeat: [GAAA]](https://image.slidesharecdn.com/mscivsemesterdnaprofiling-strbiologyandartifacts-240515082426-9bd80228/75/MSC-IV-SEMESTER_DNA-Profiling-STR-biology-and-artifacts-pdf-35-2048.jpg)

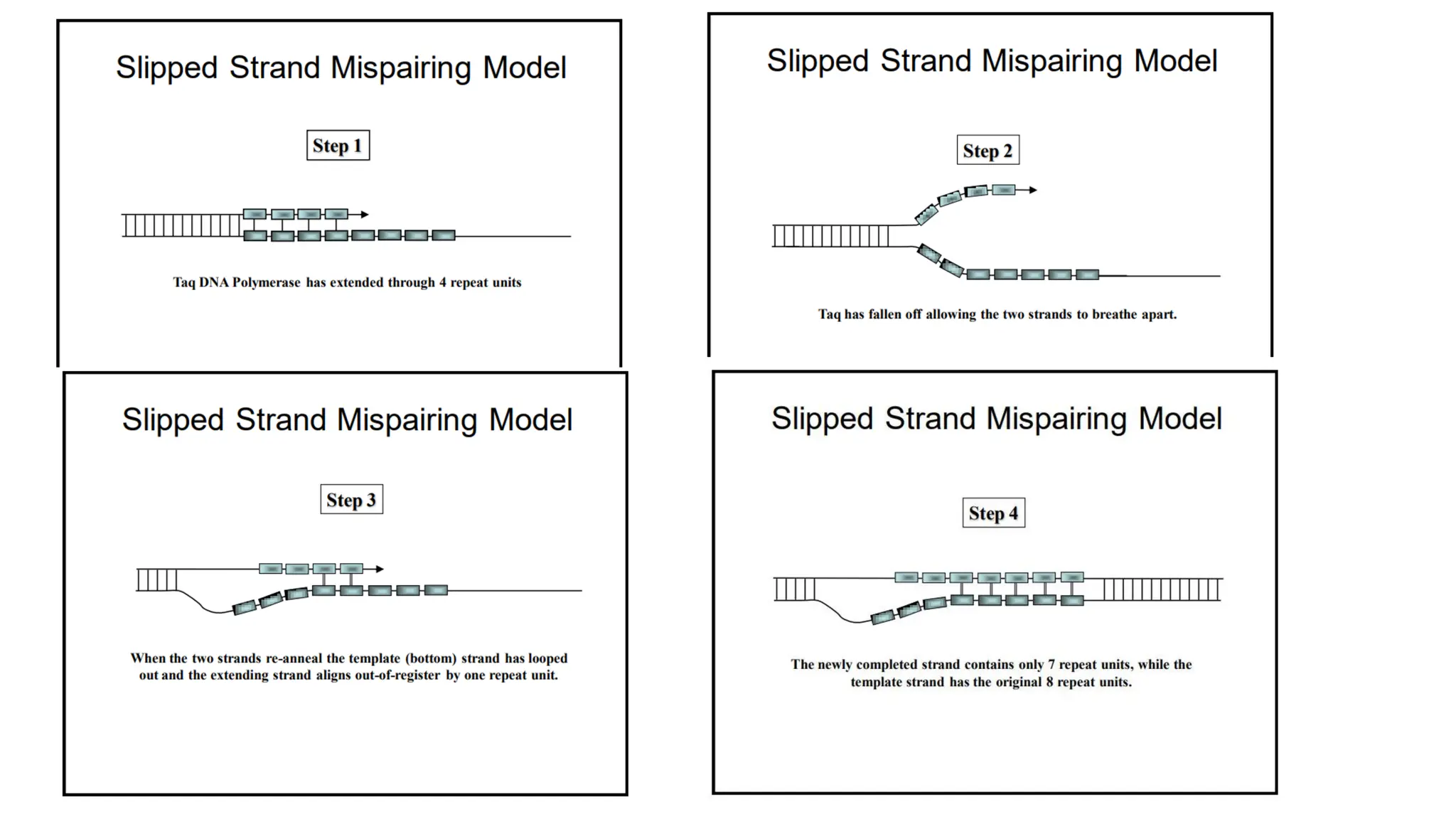
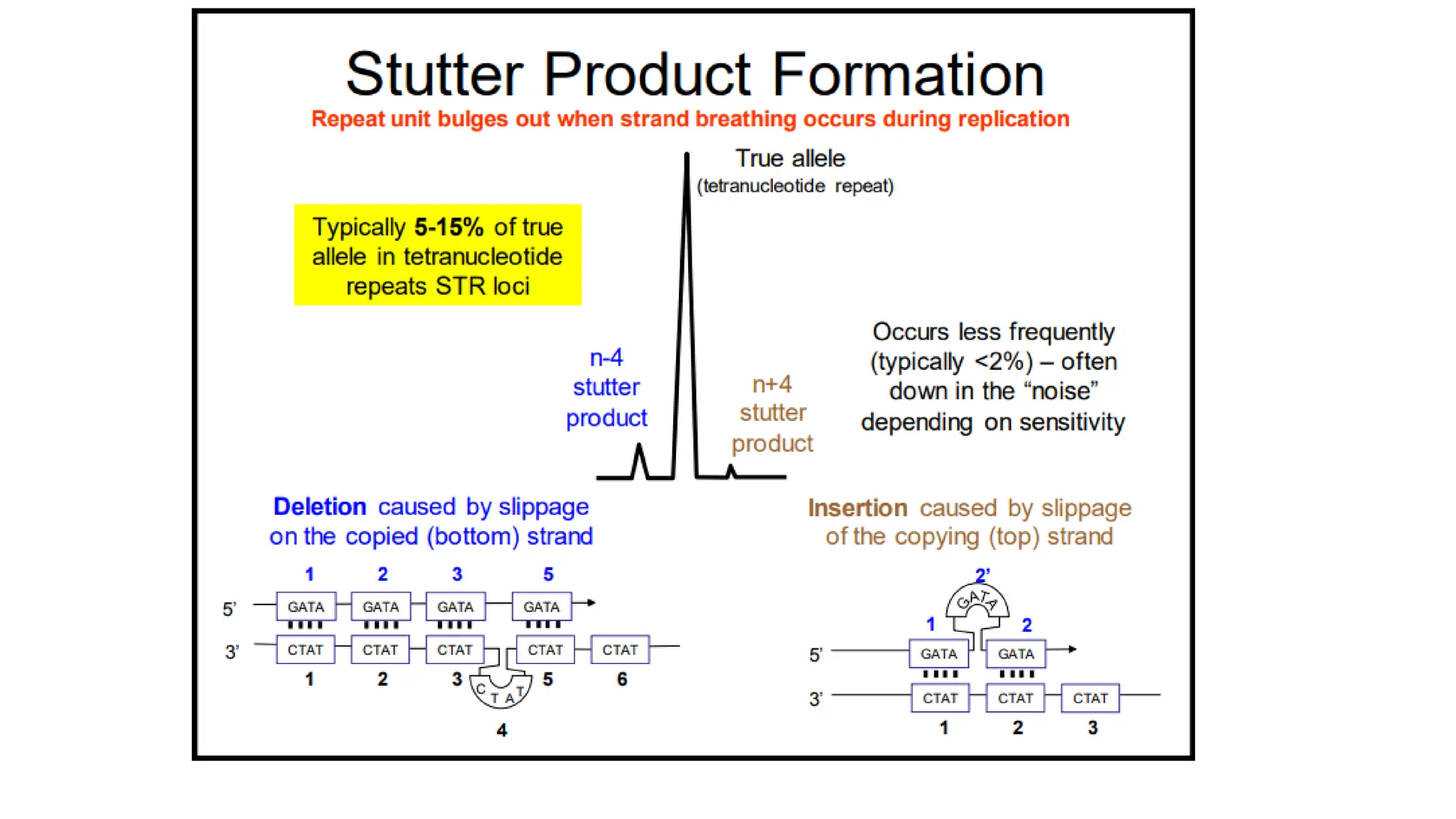


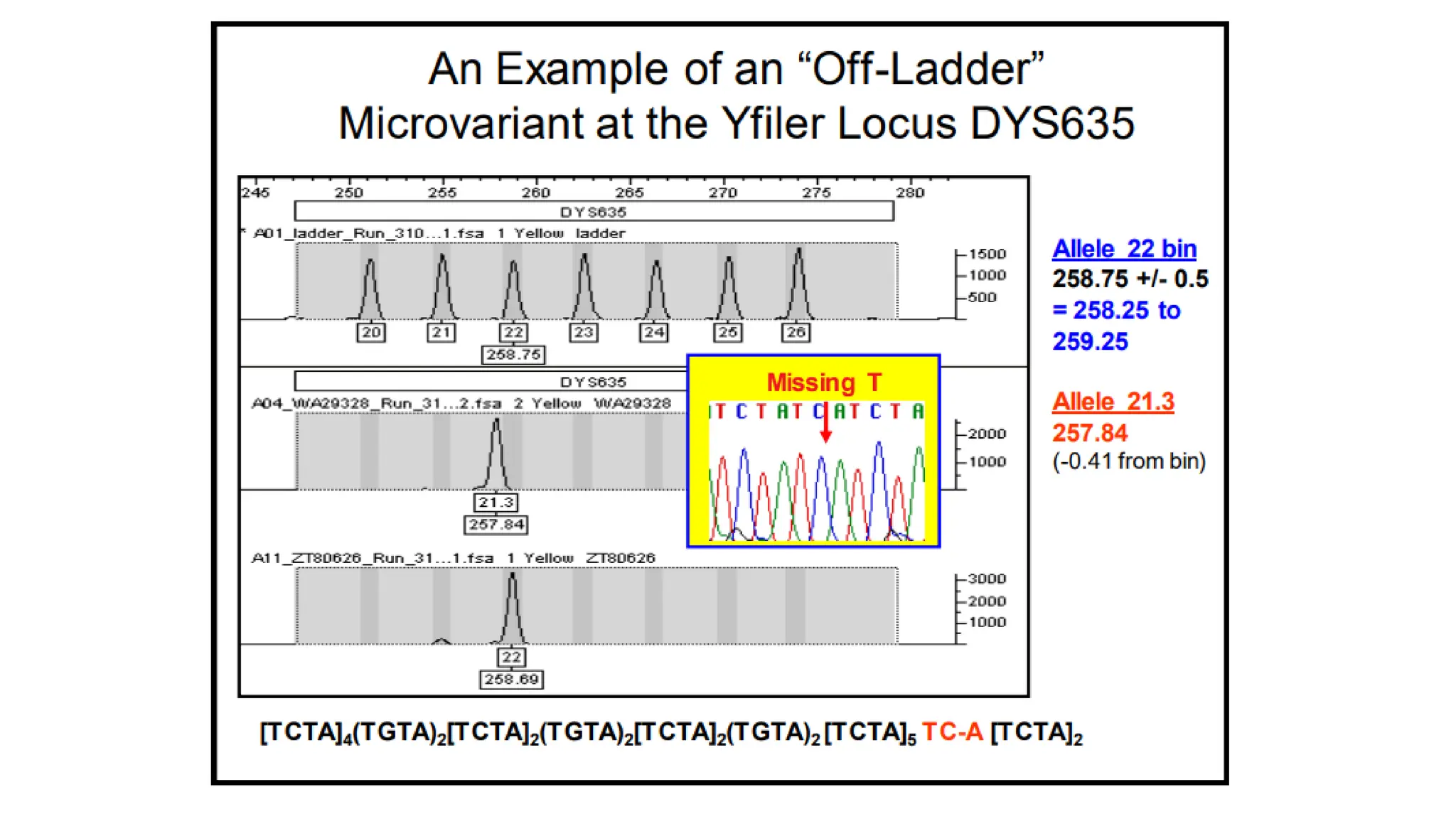

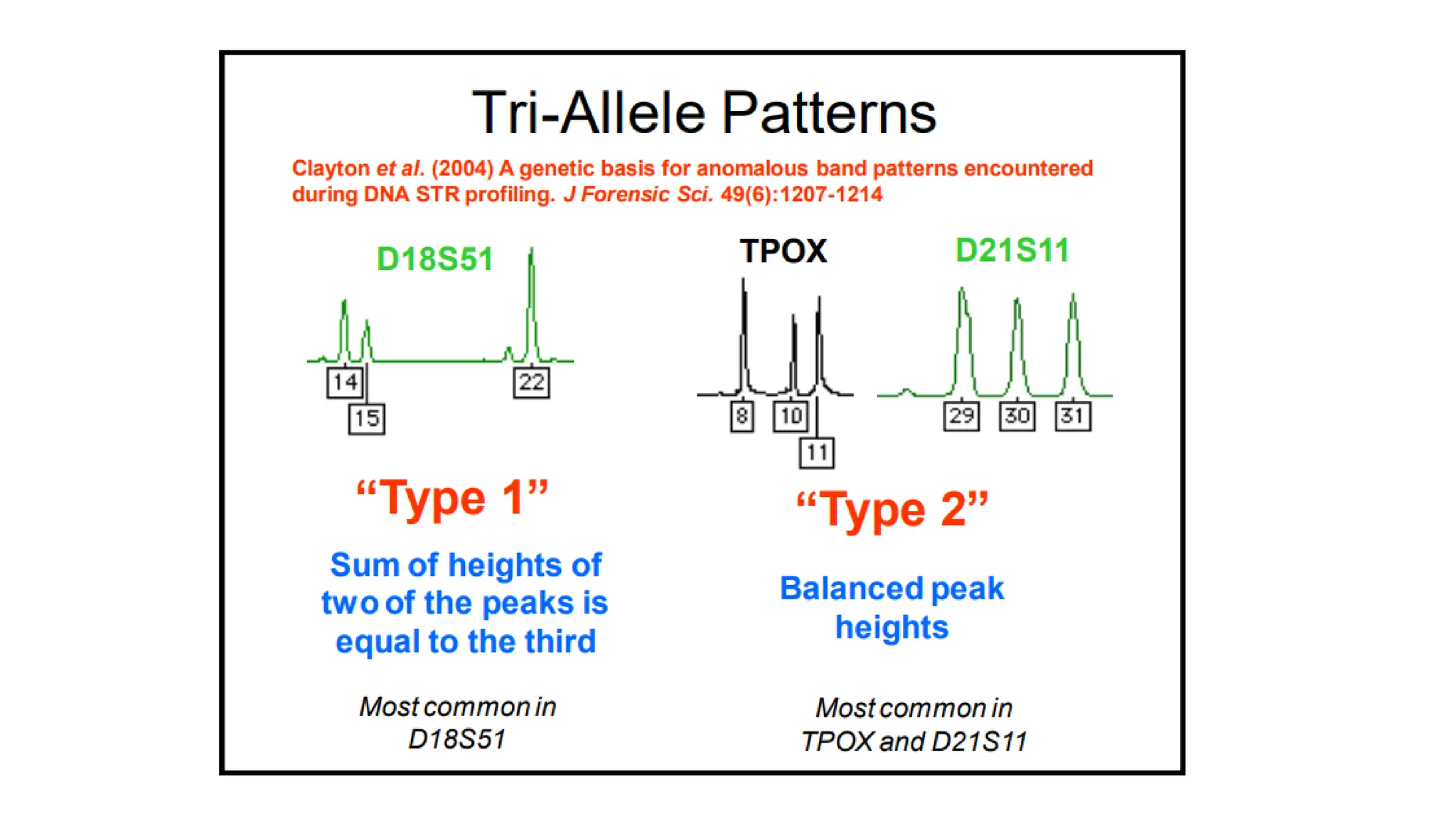

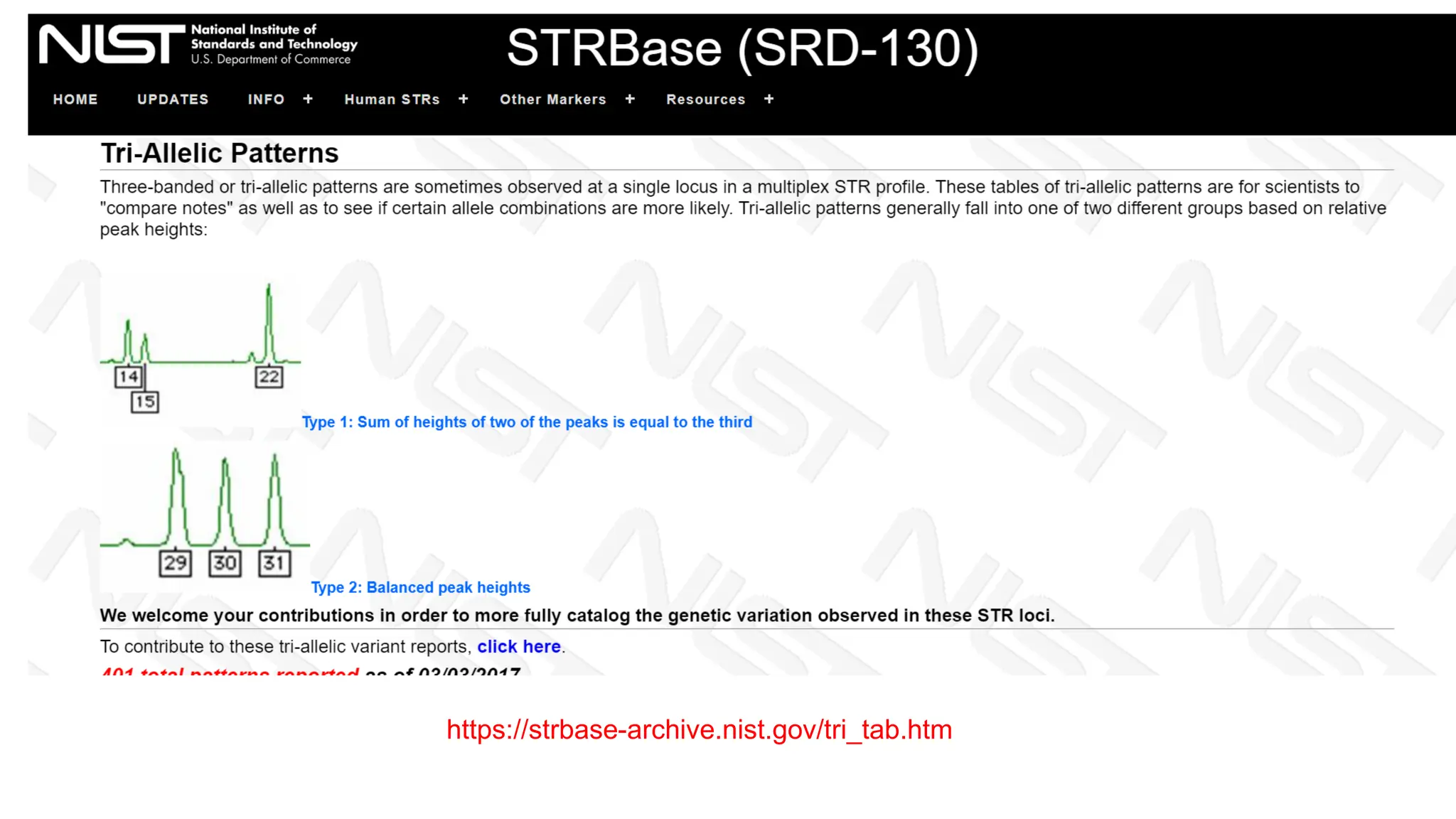
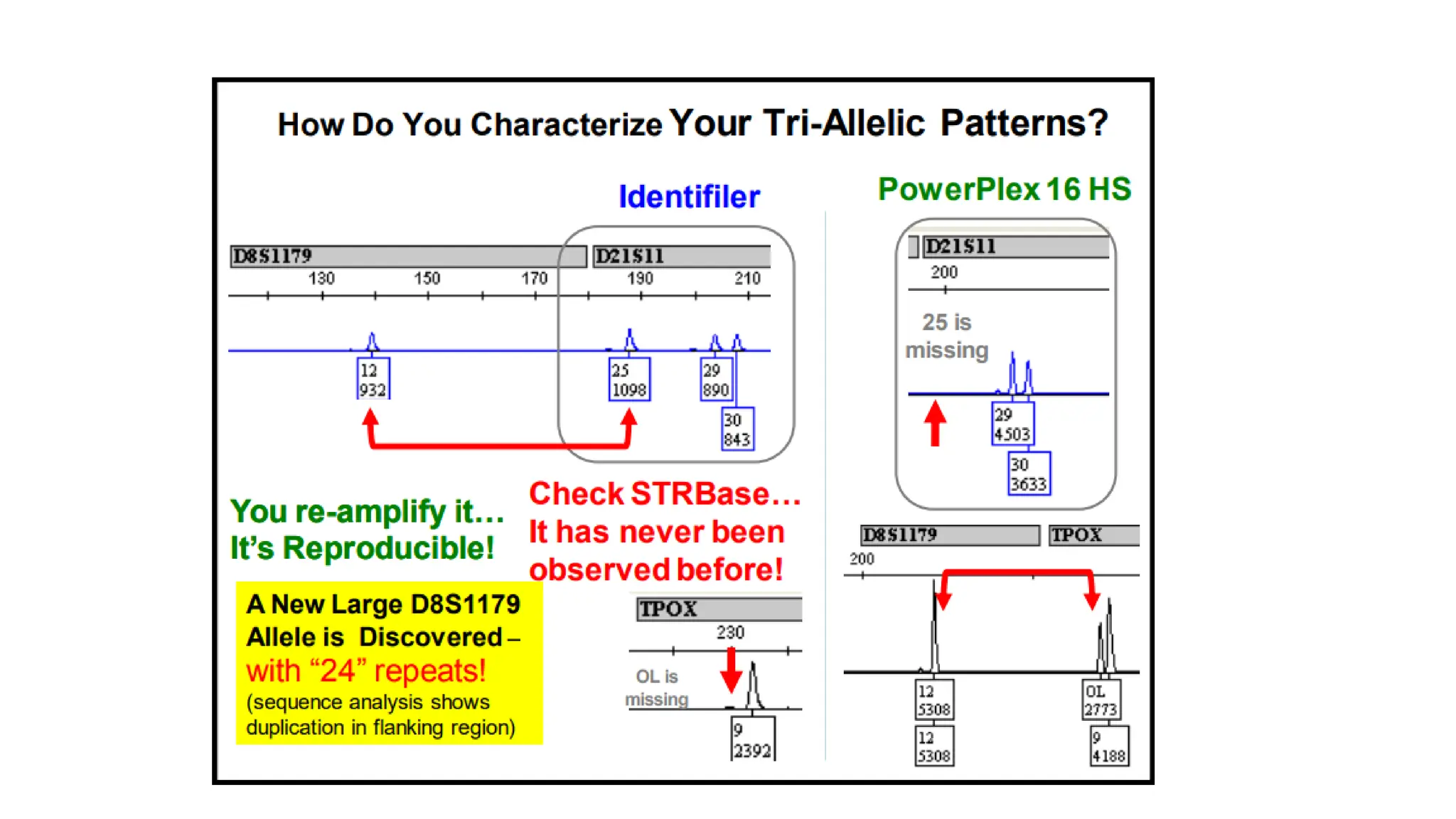


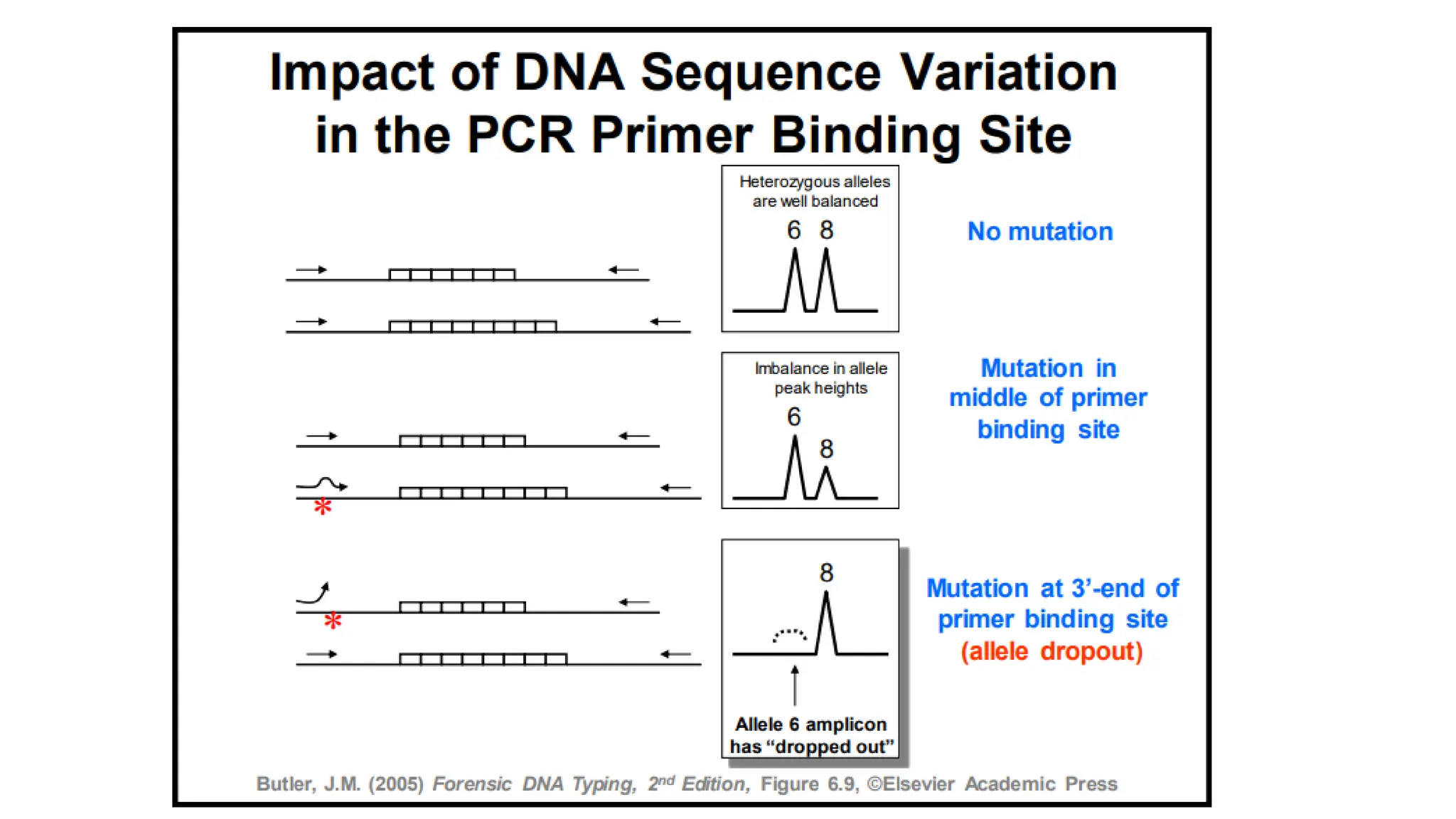
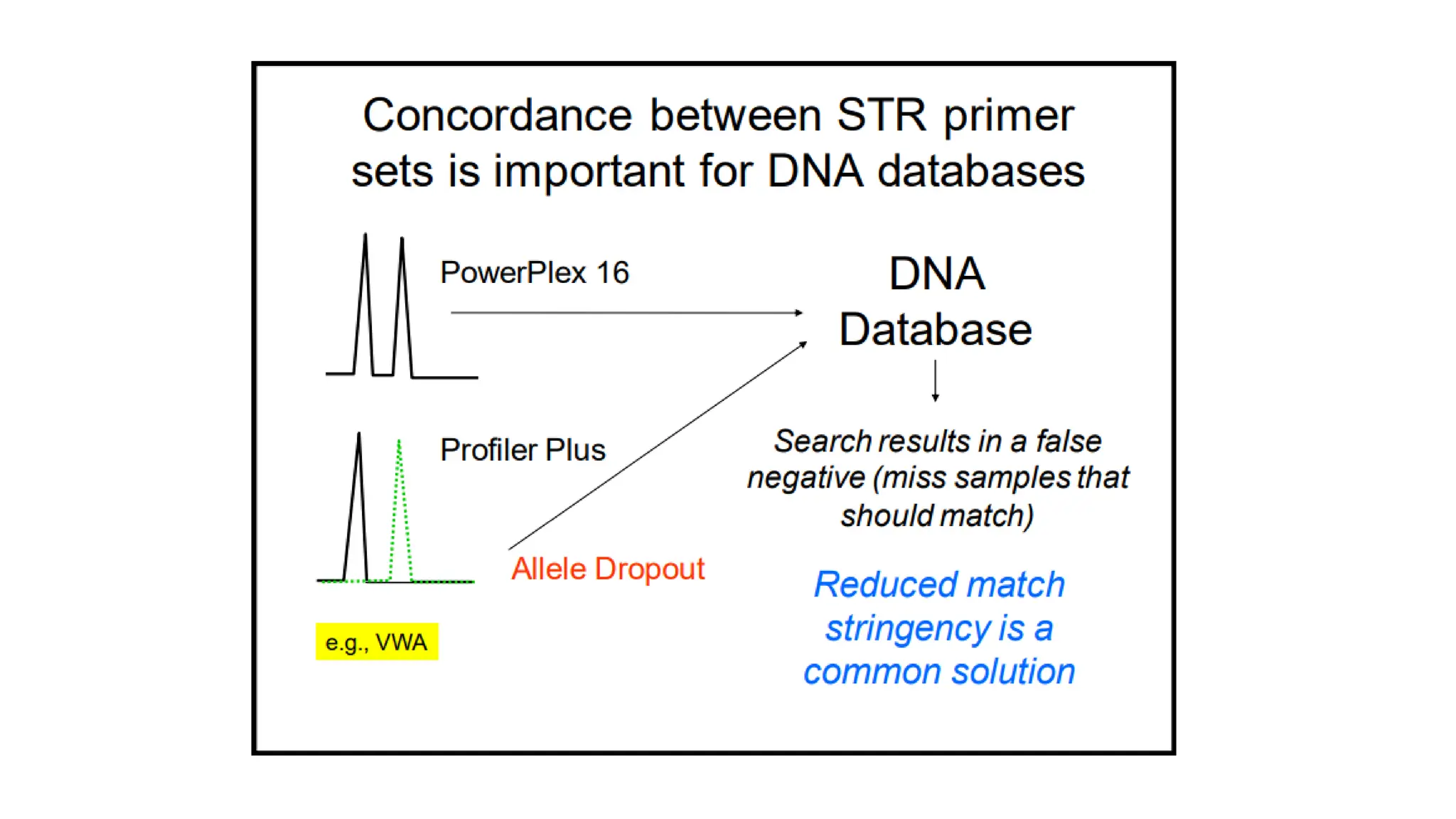

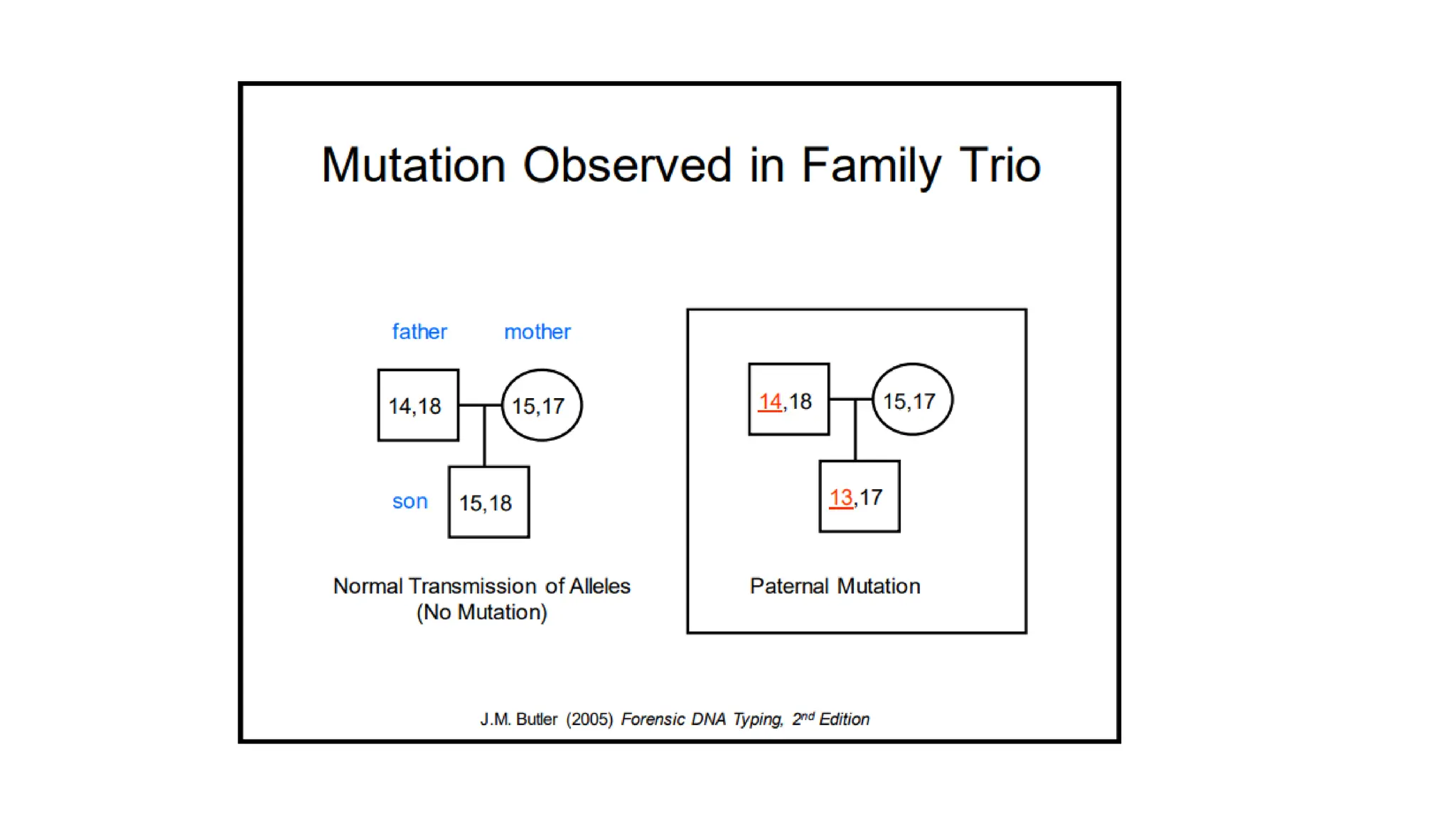
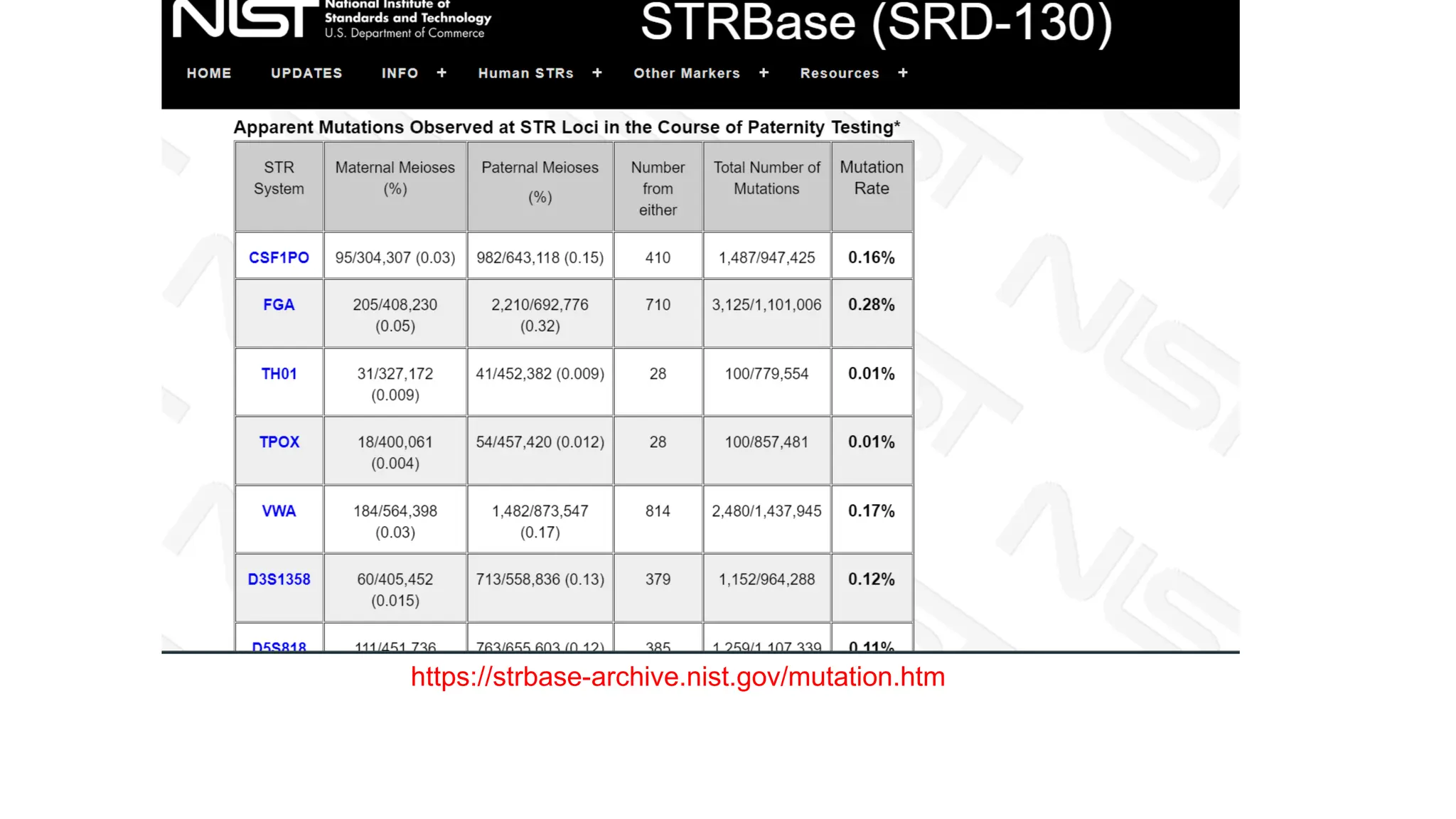


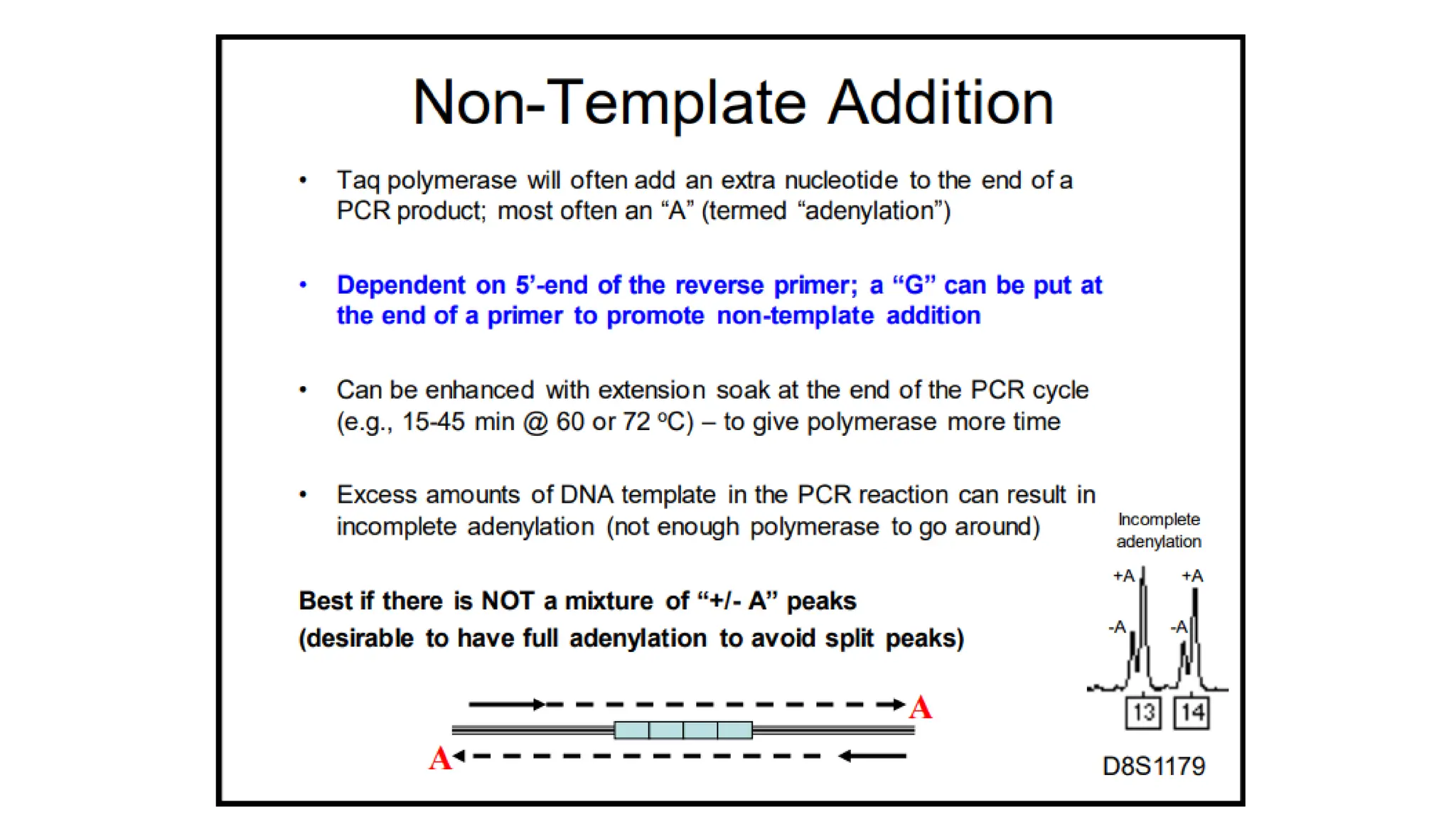
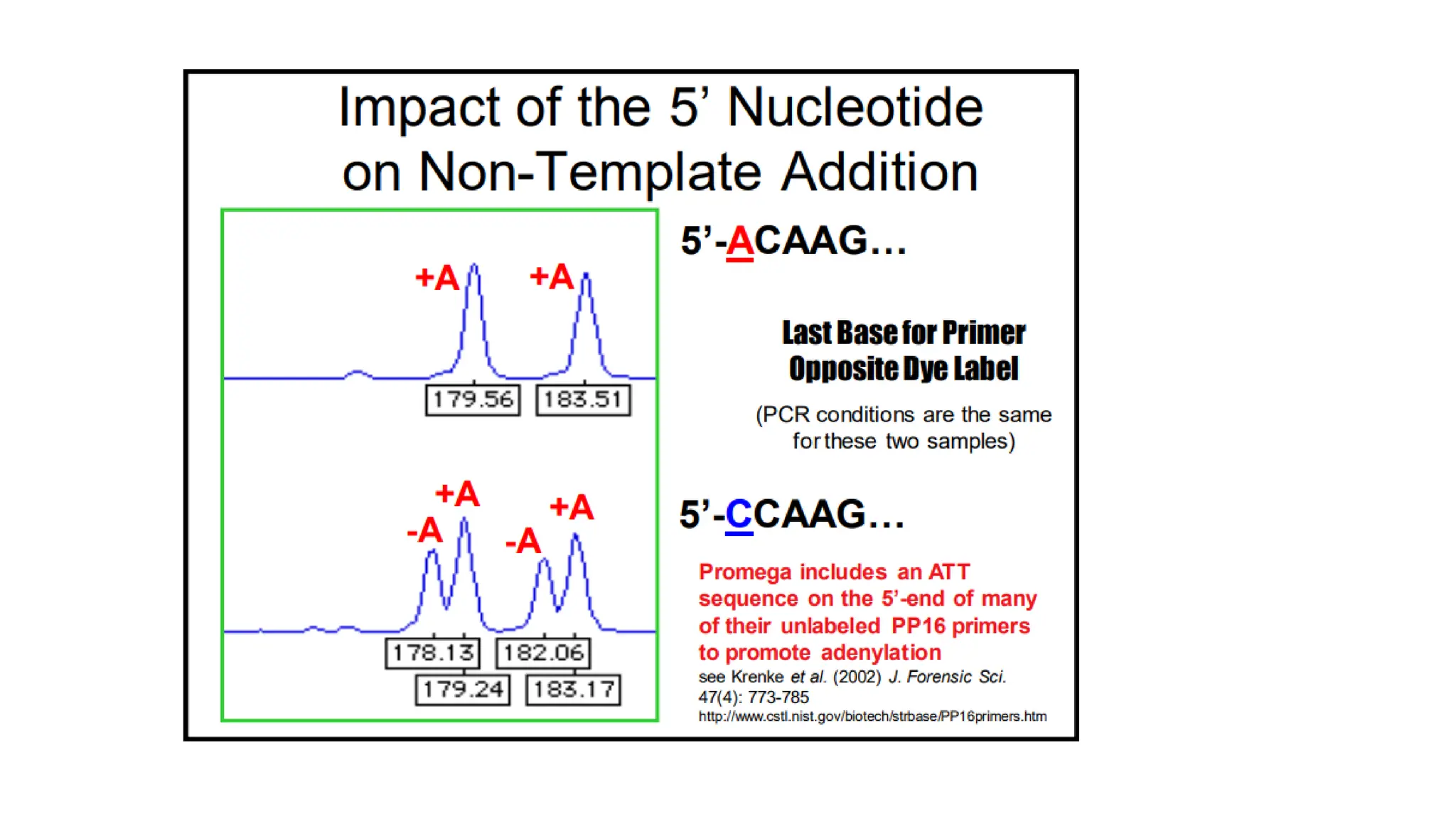
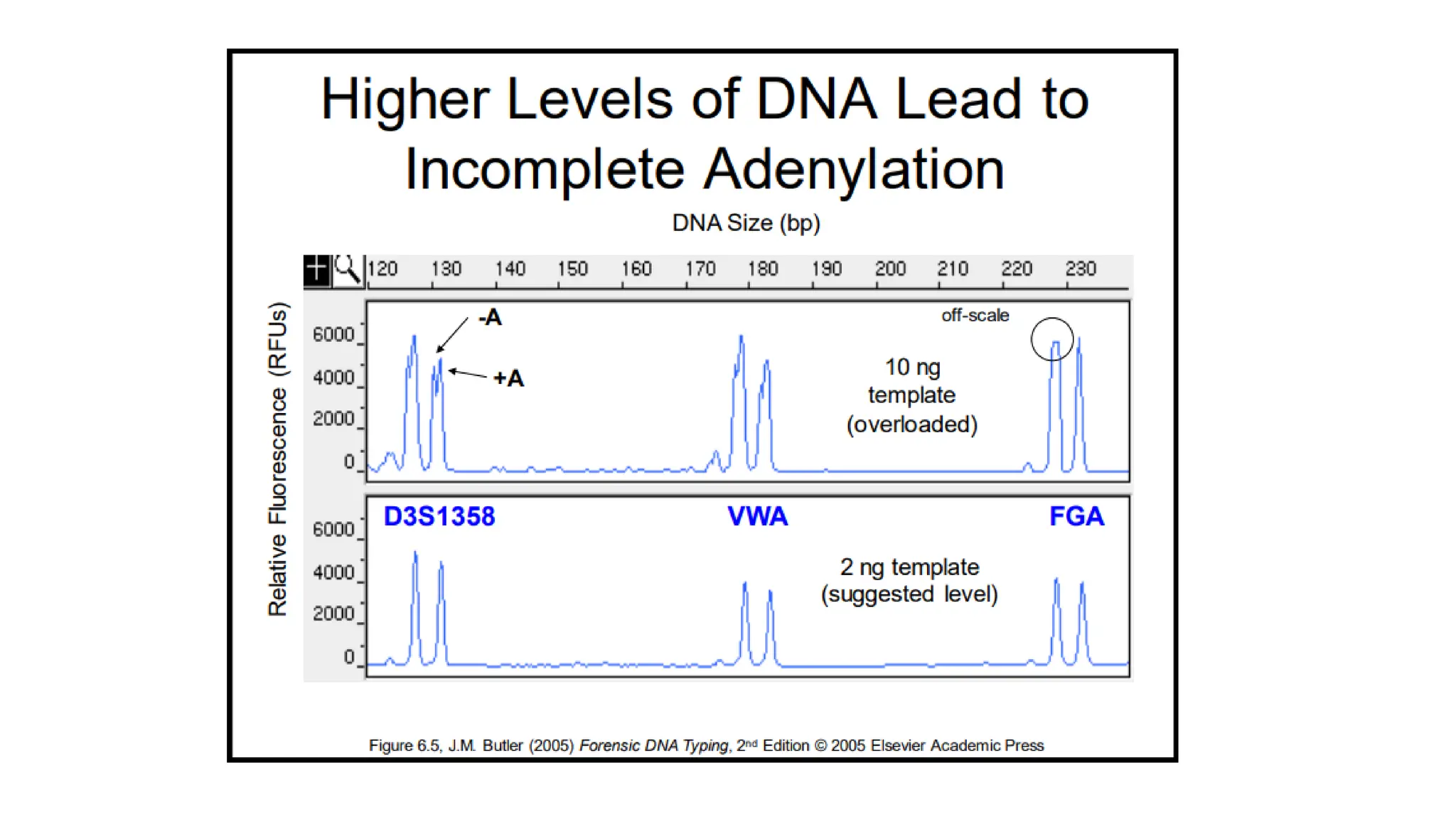
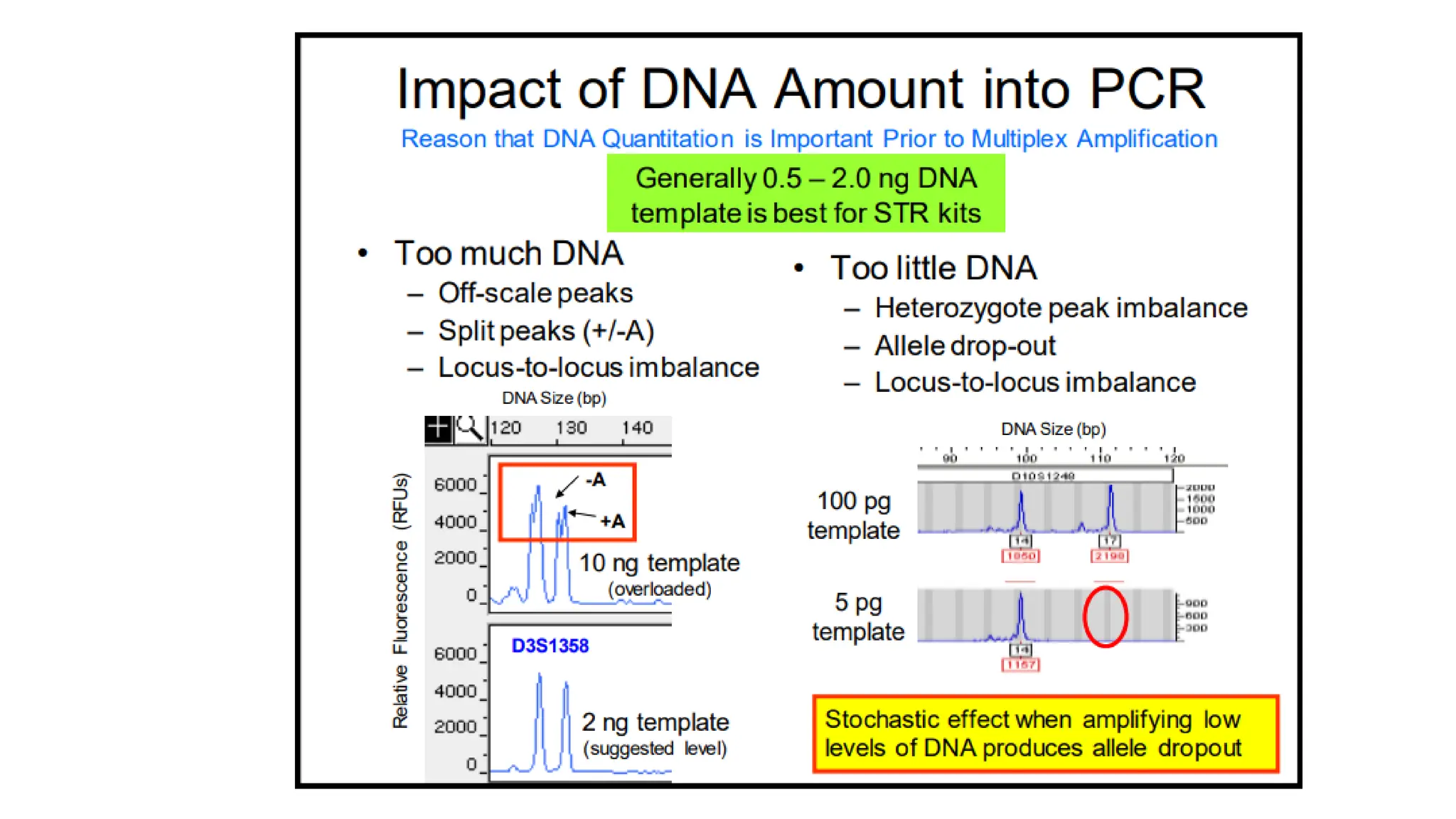


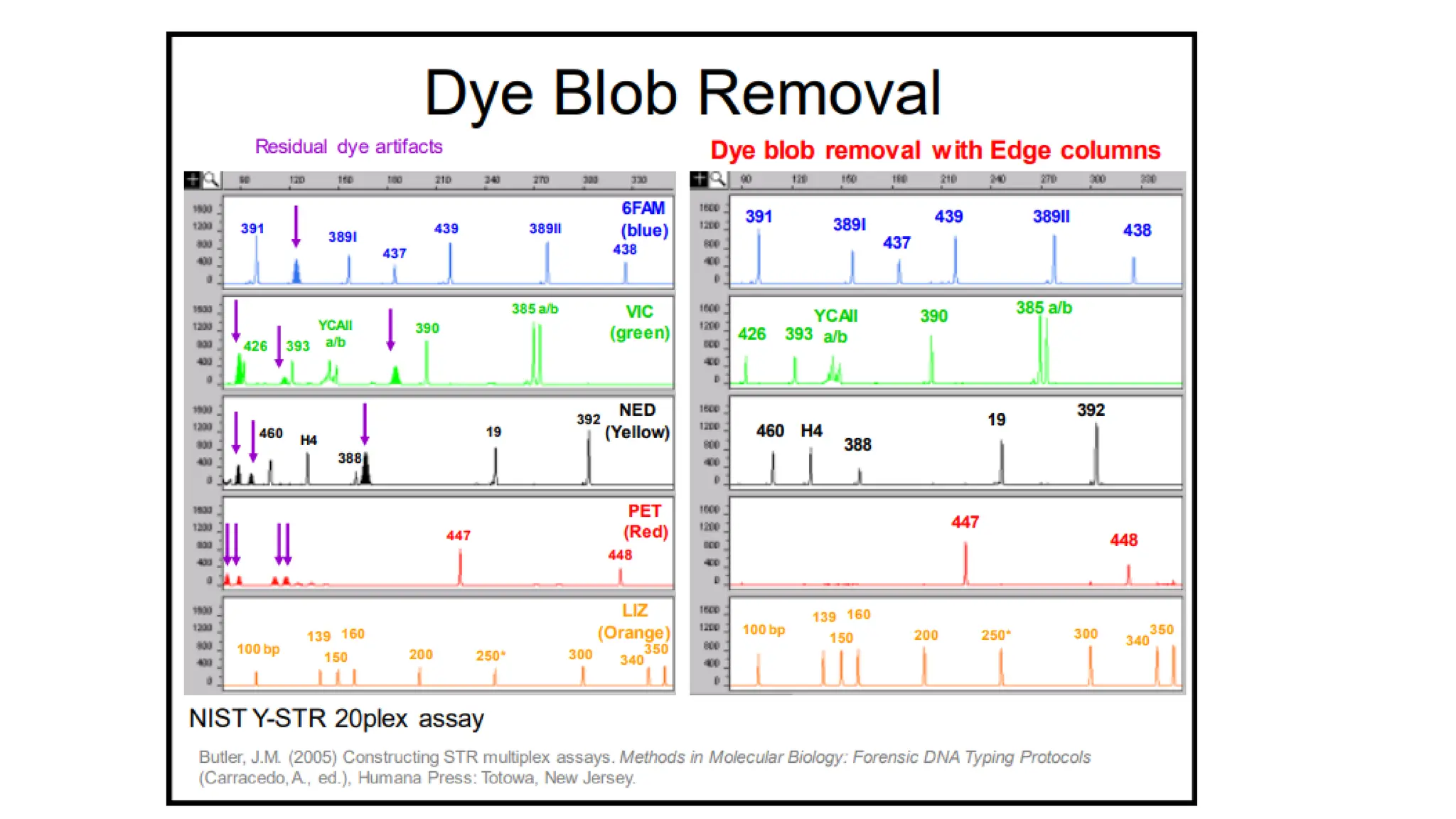

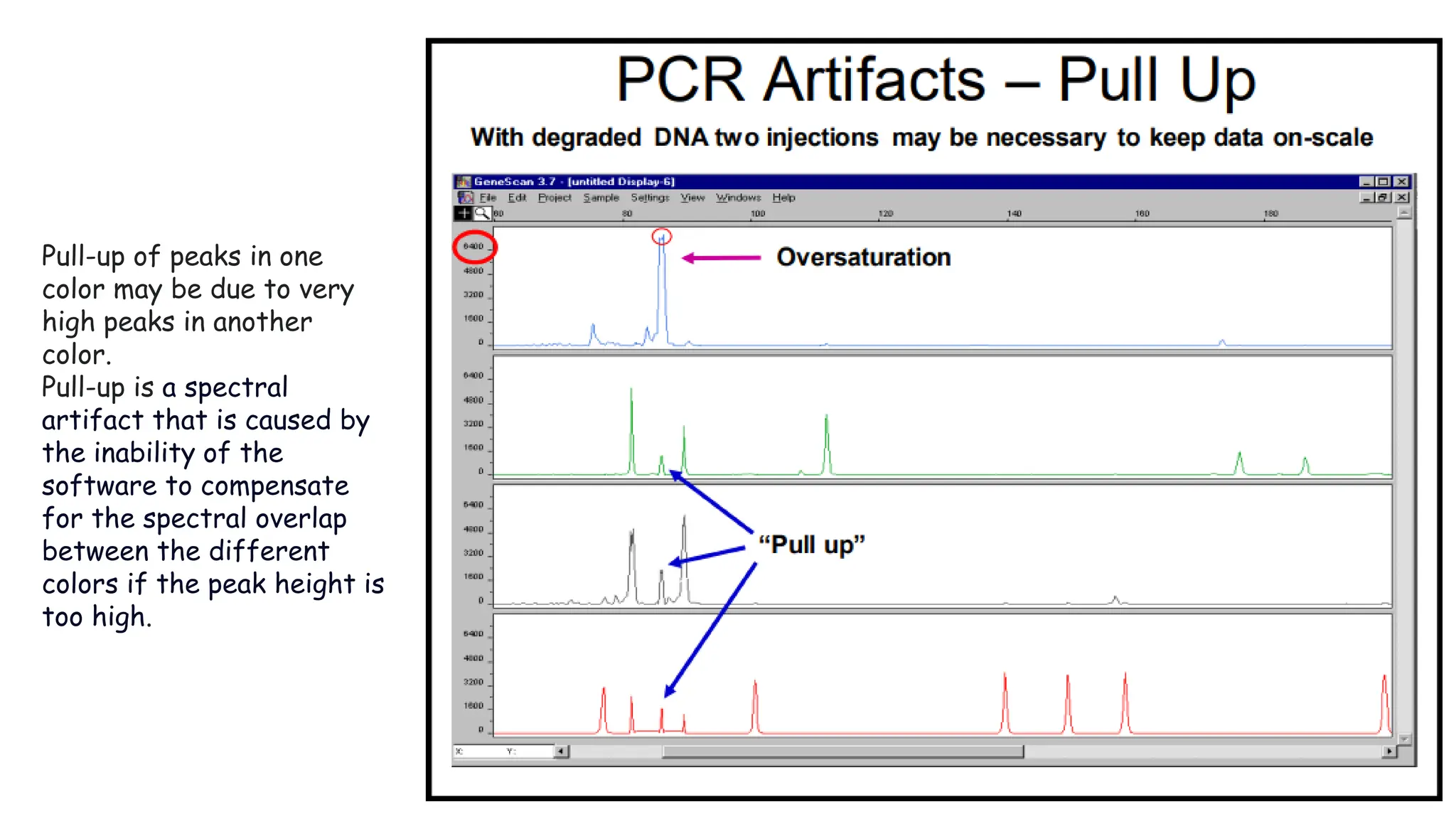
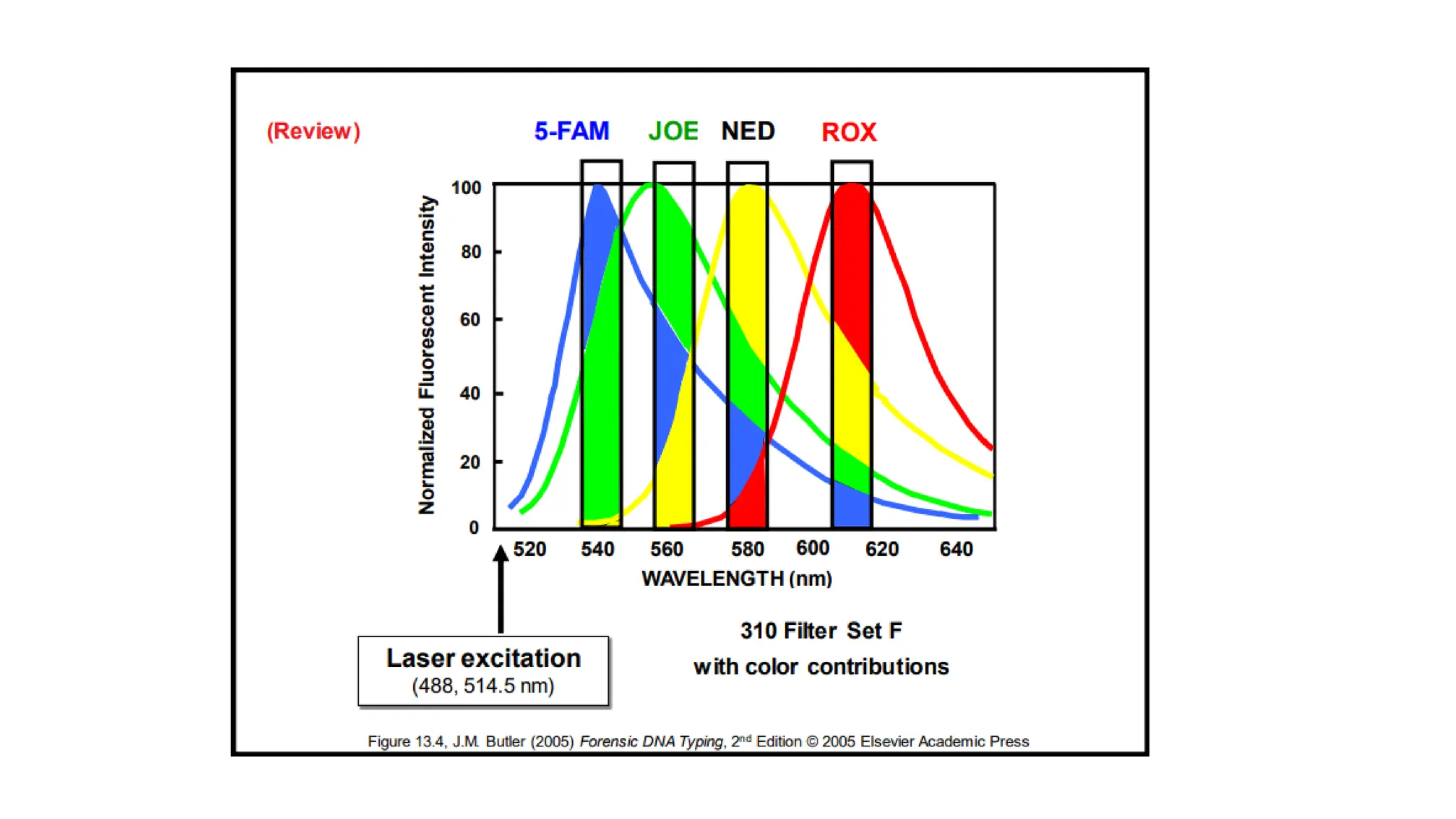


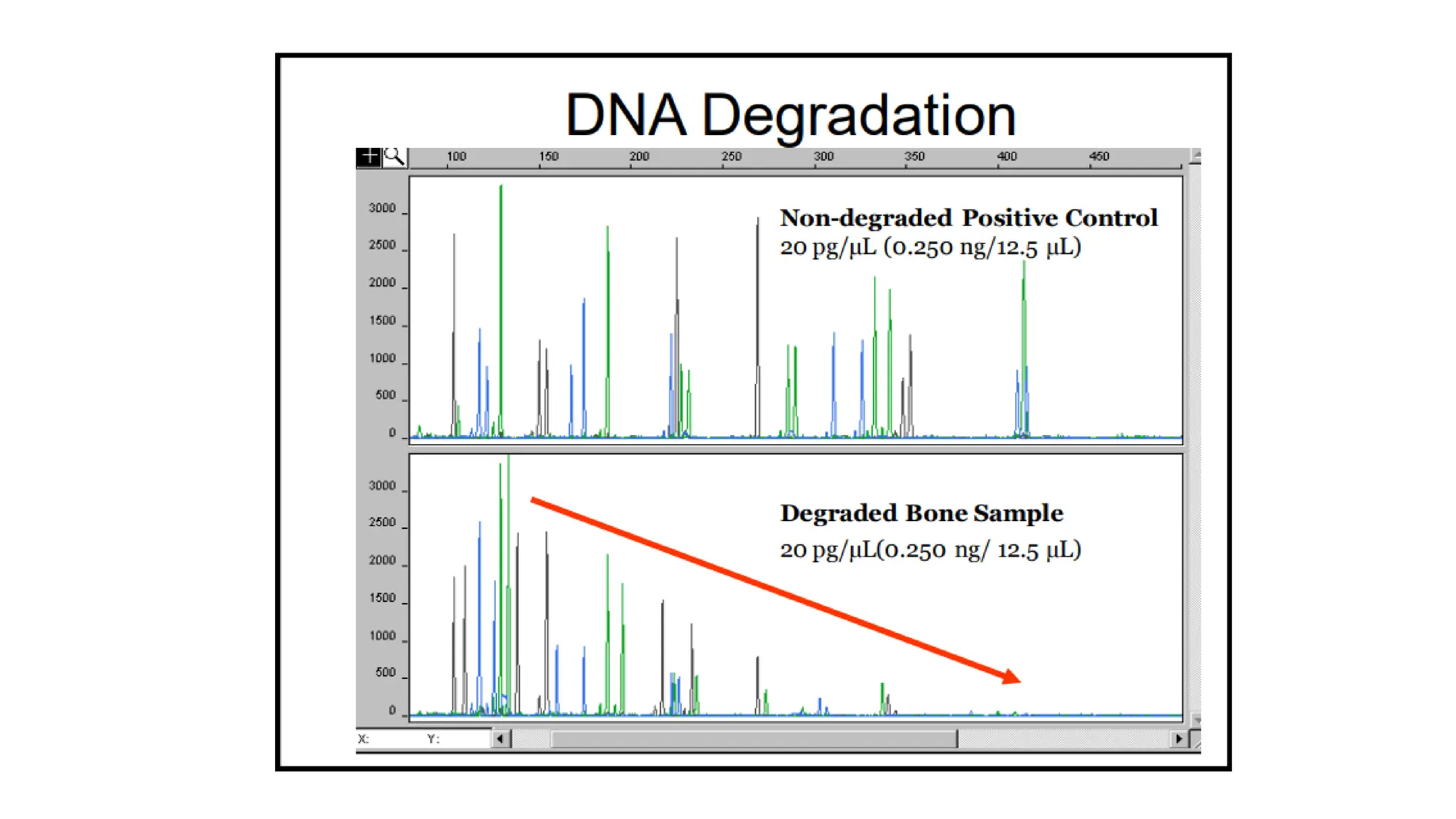
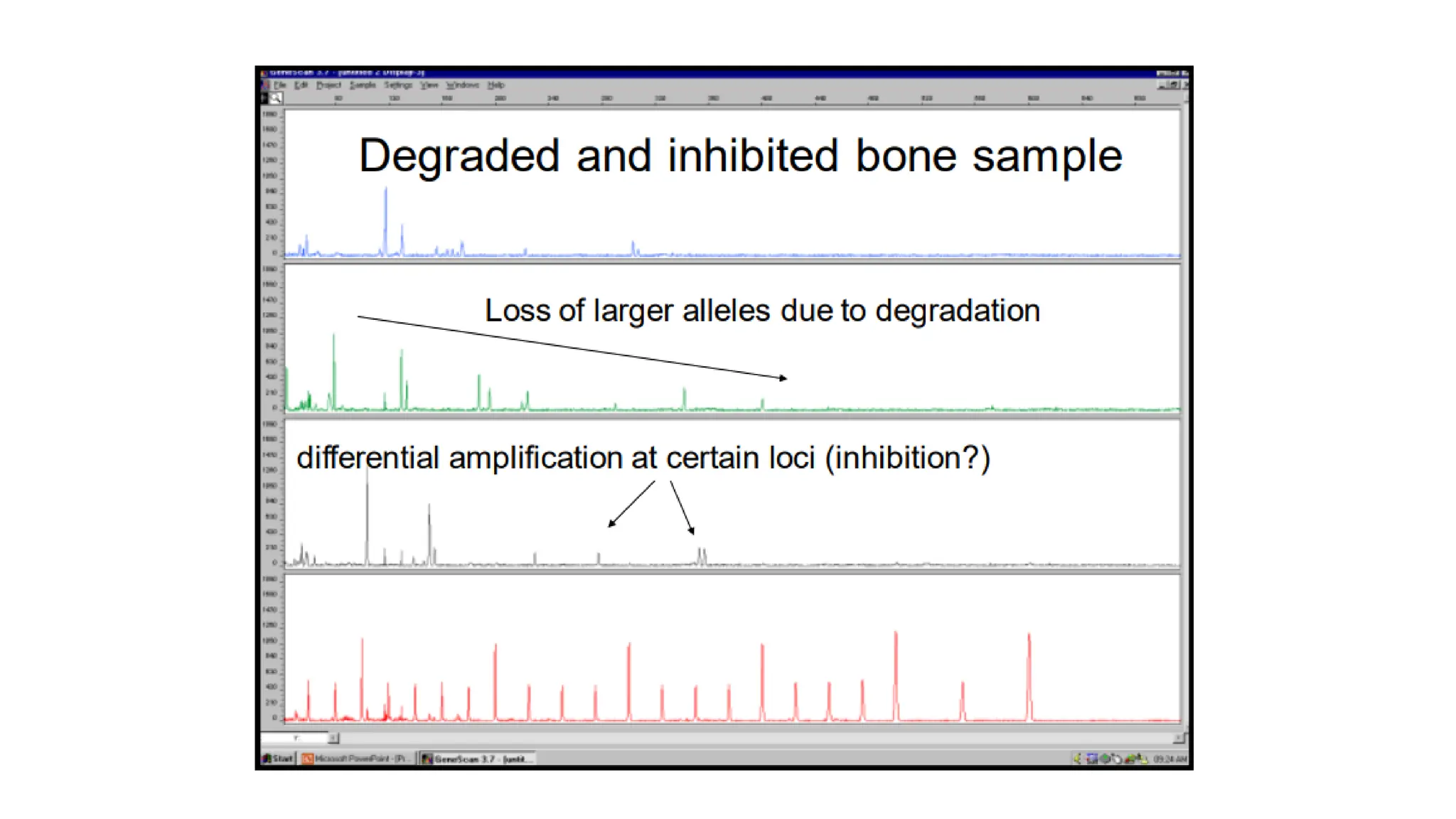



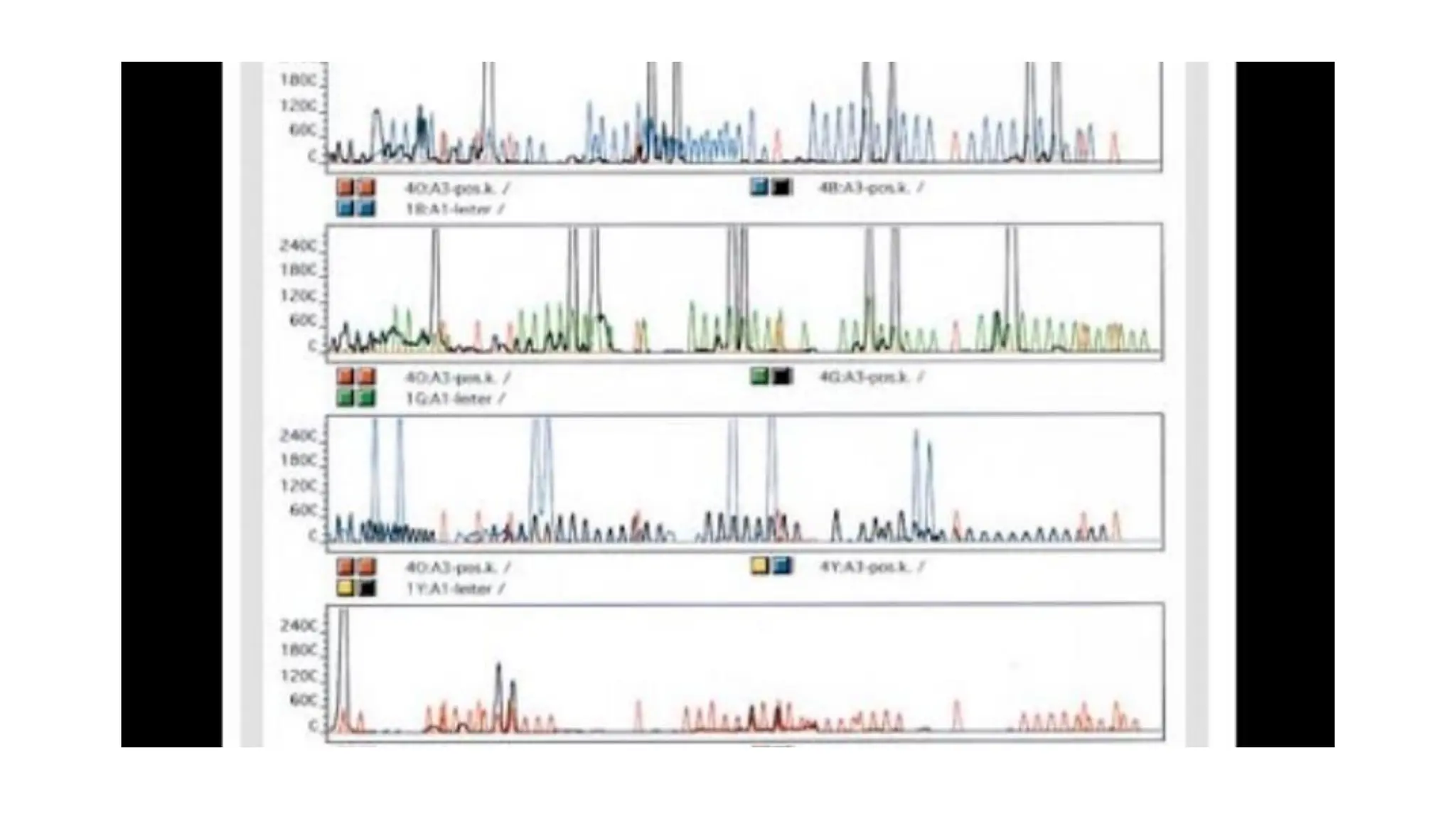

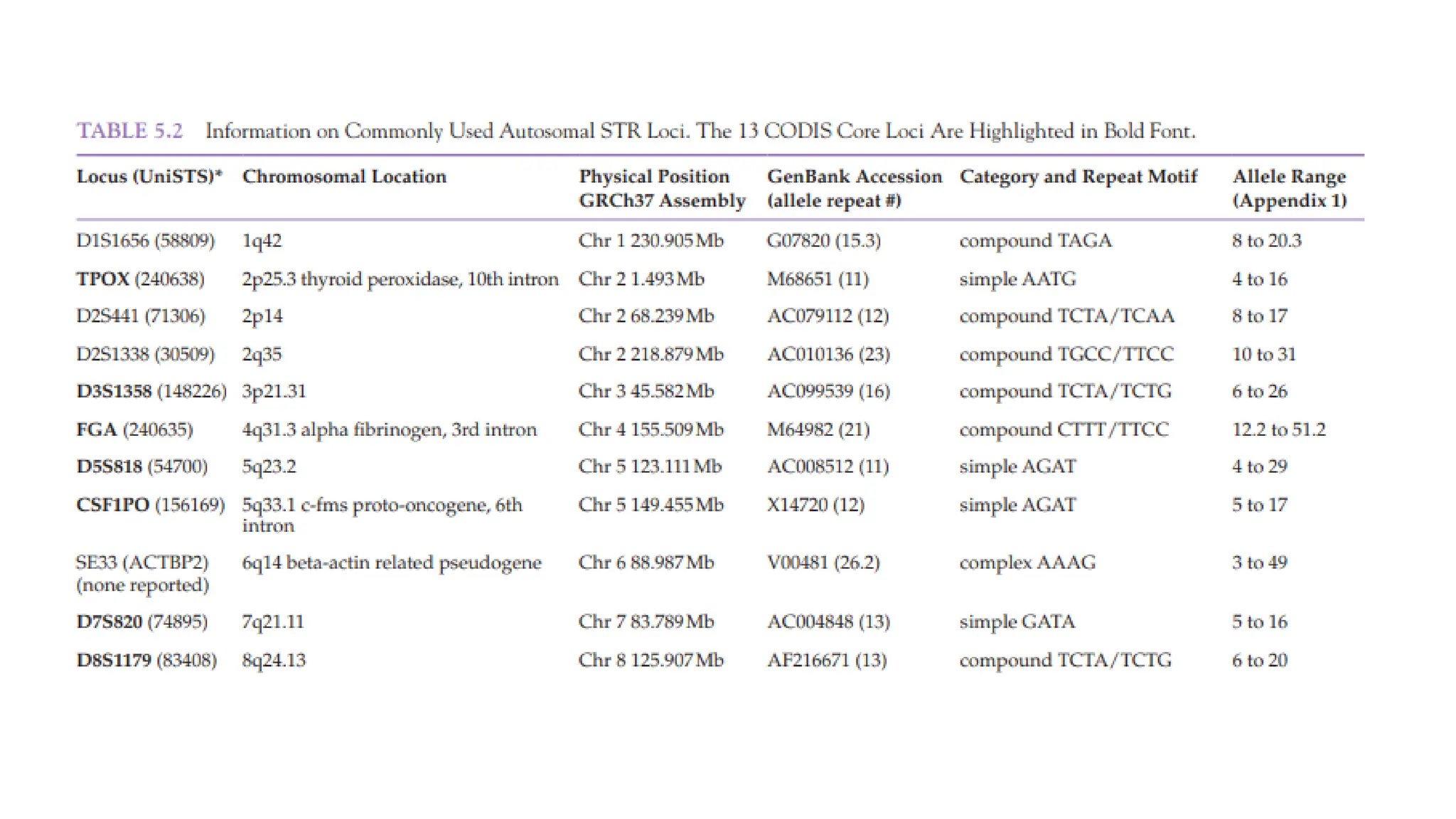
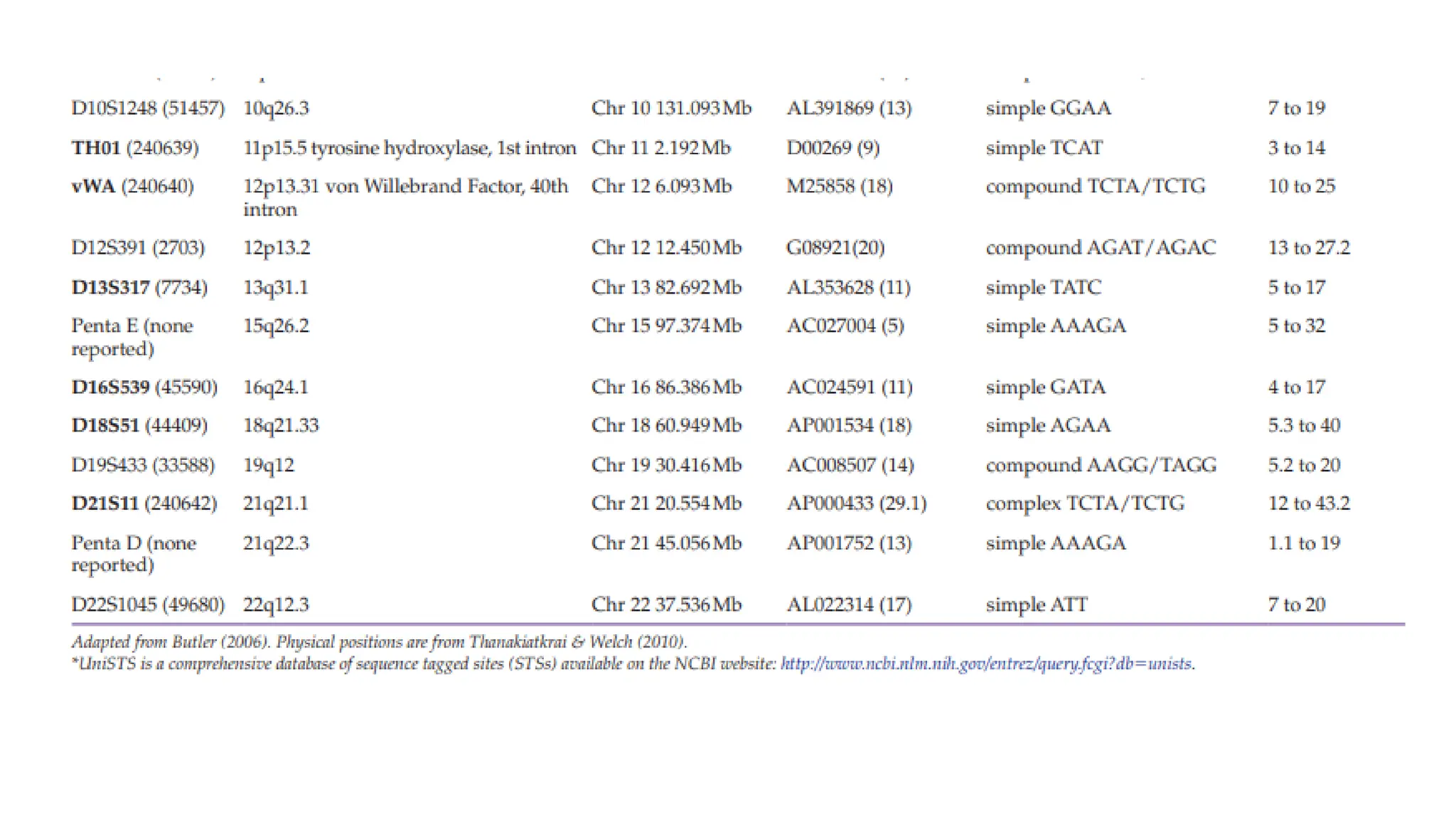
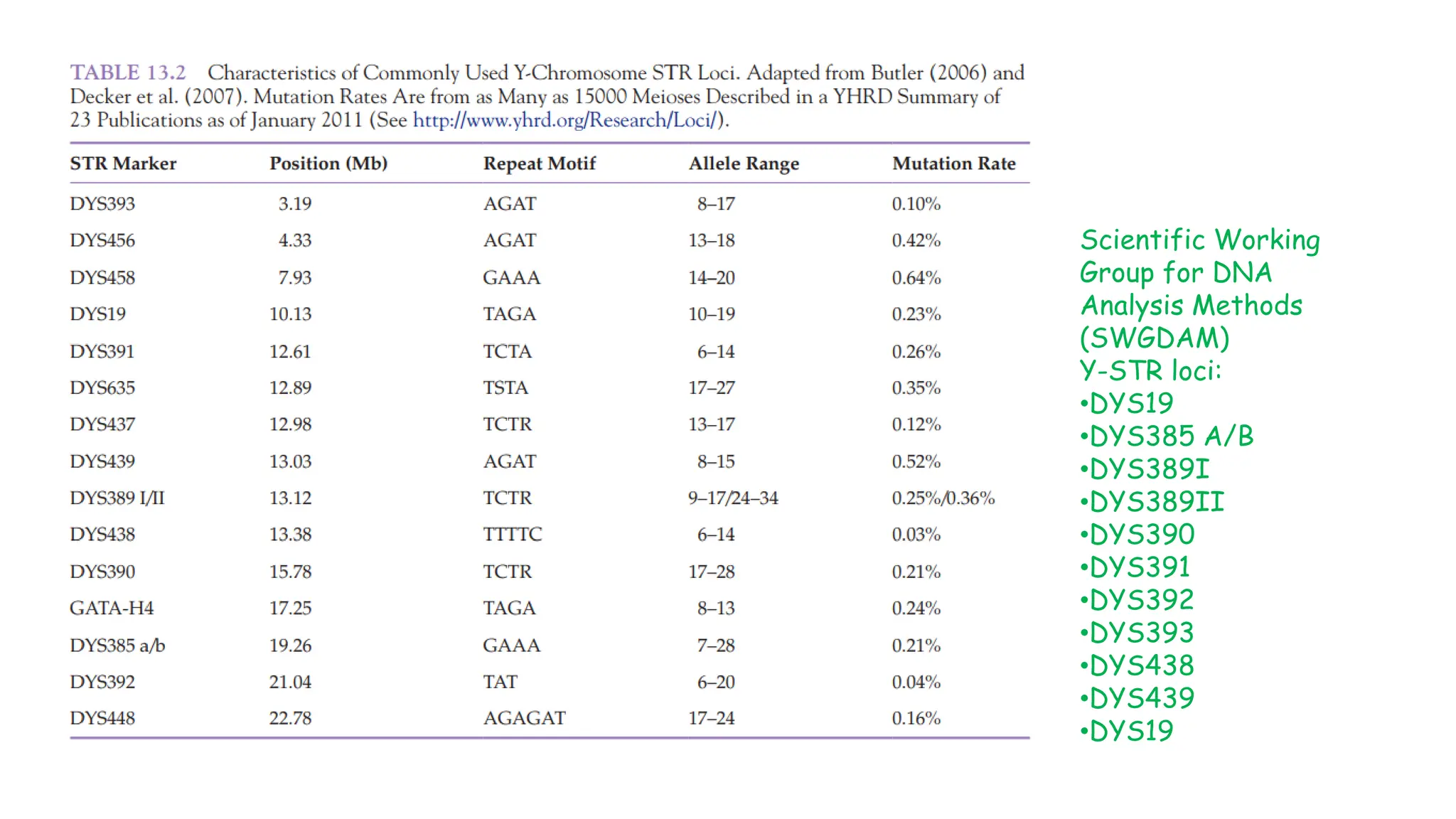
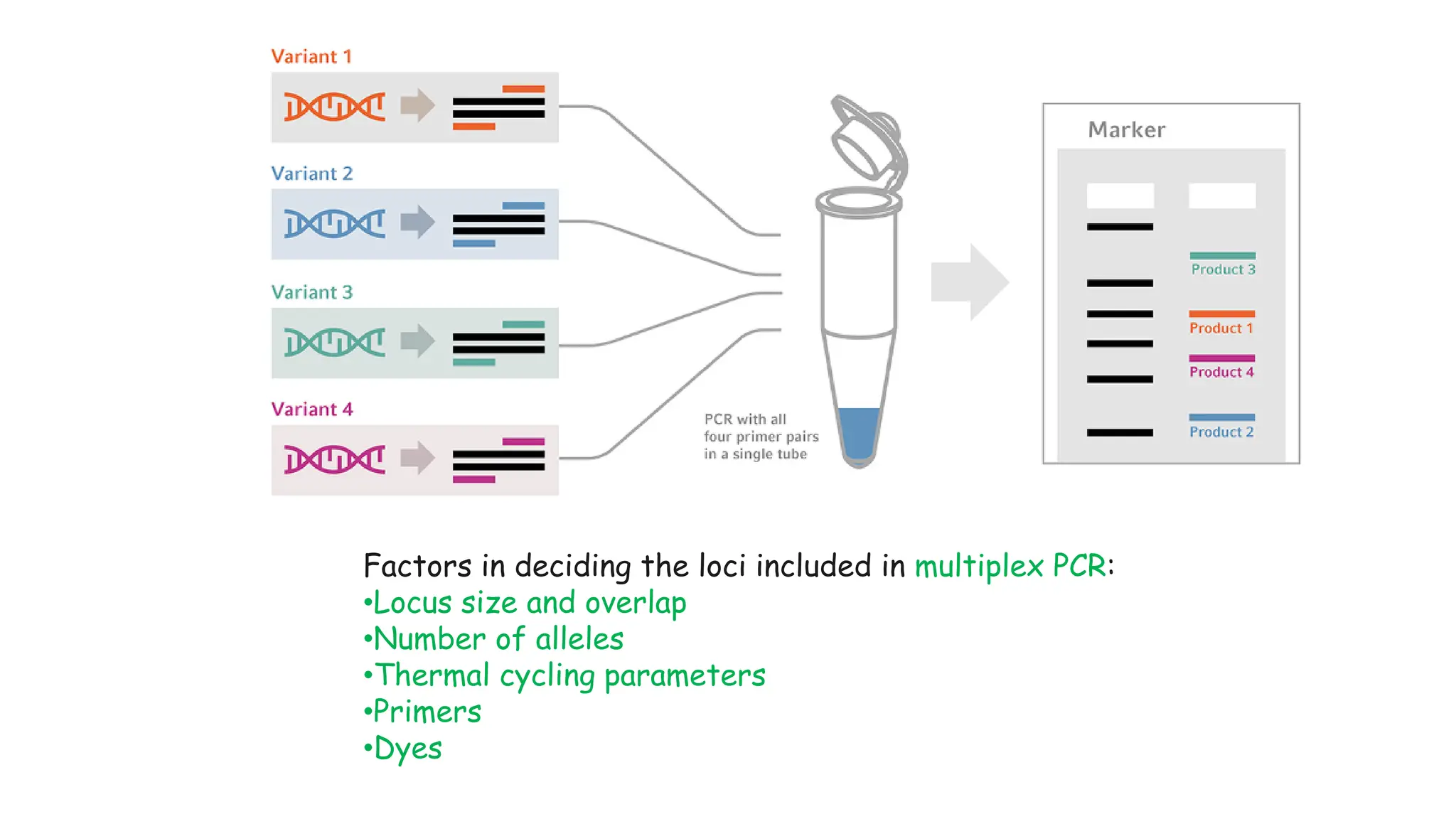
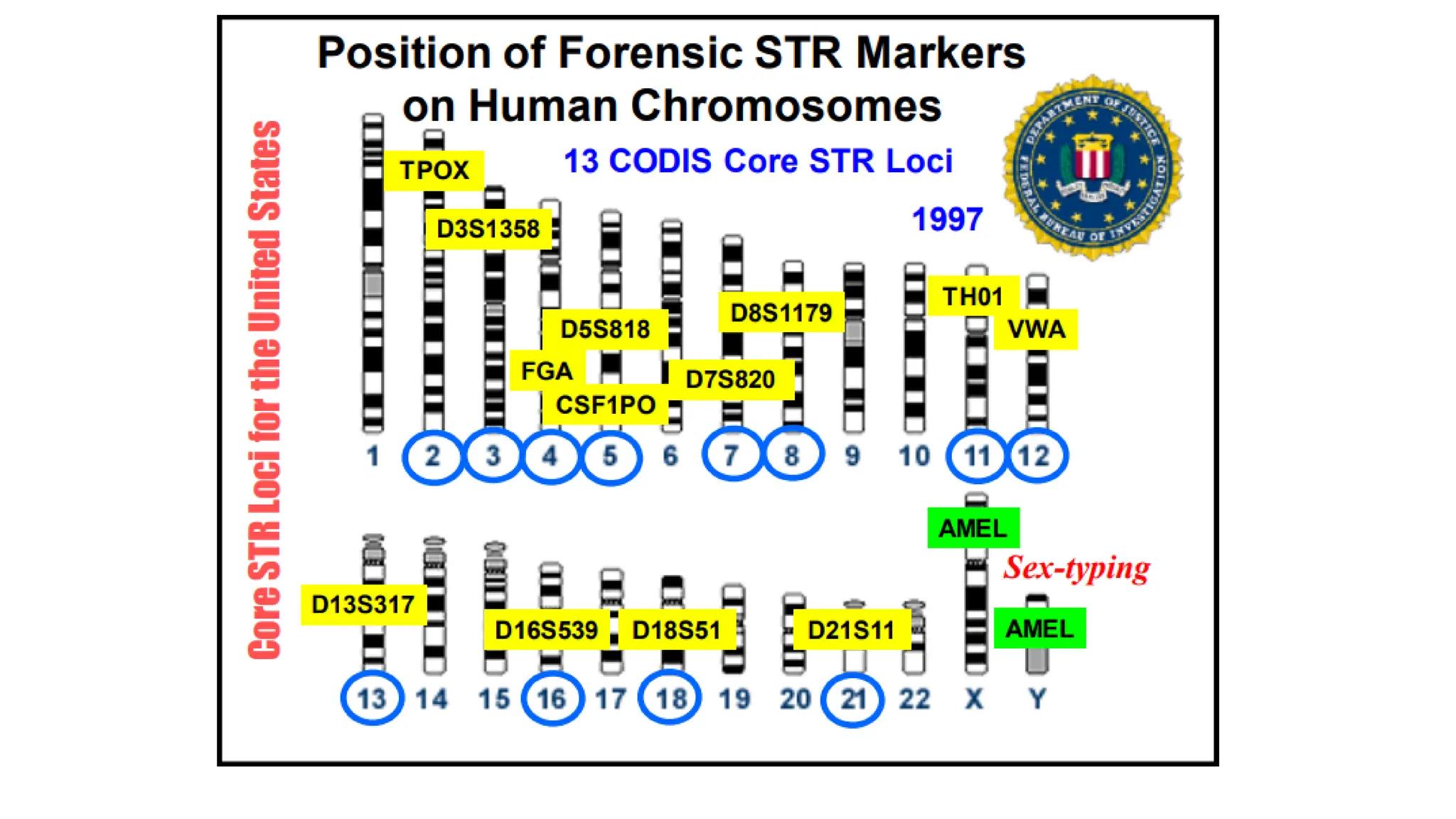
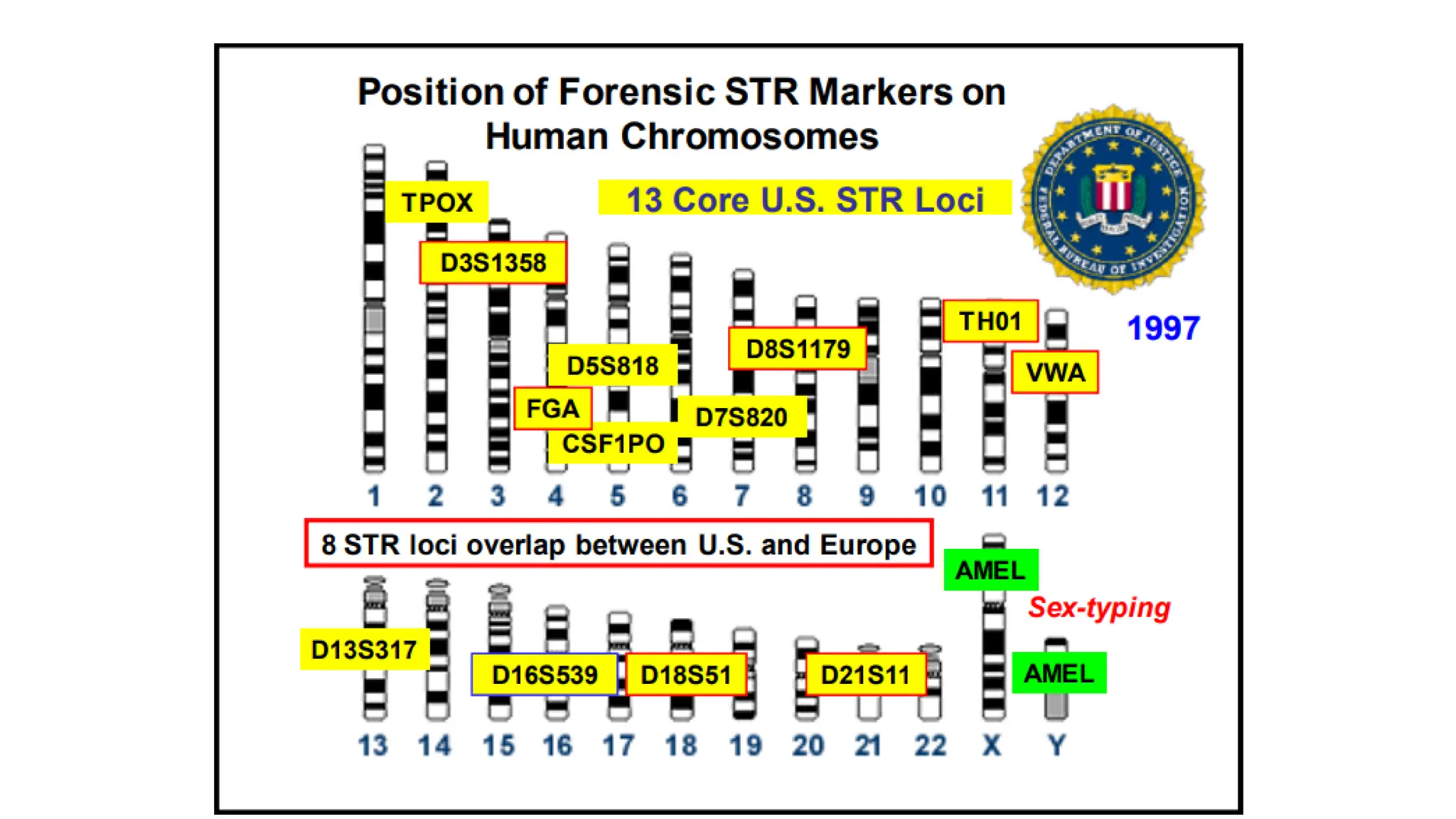
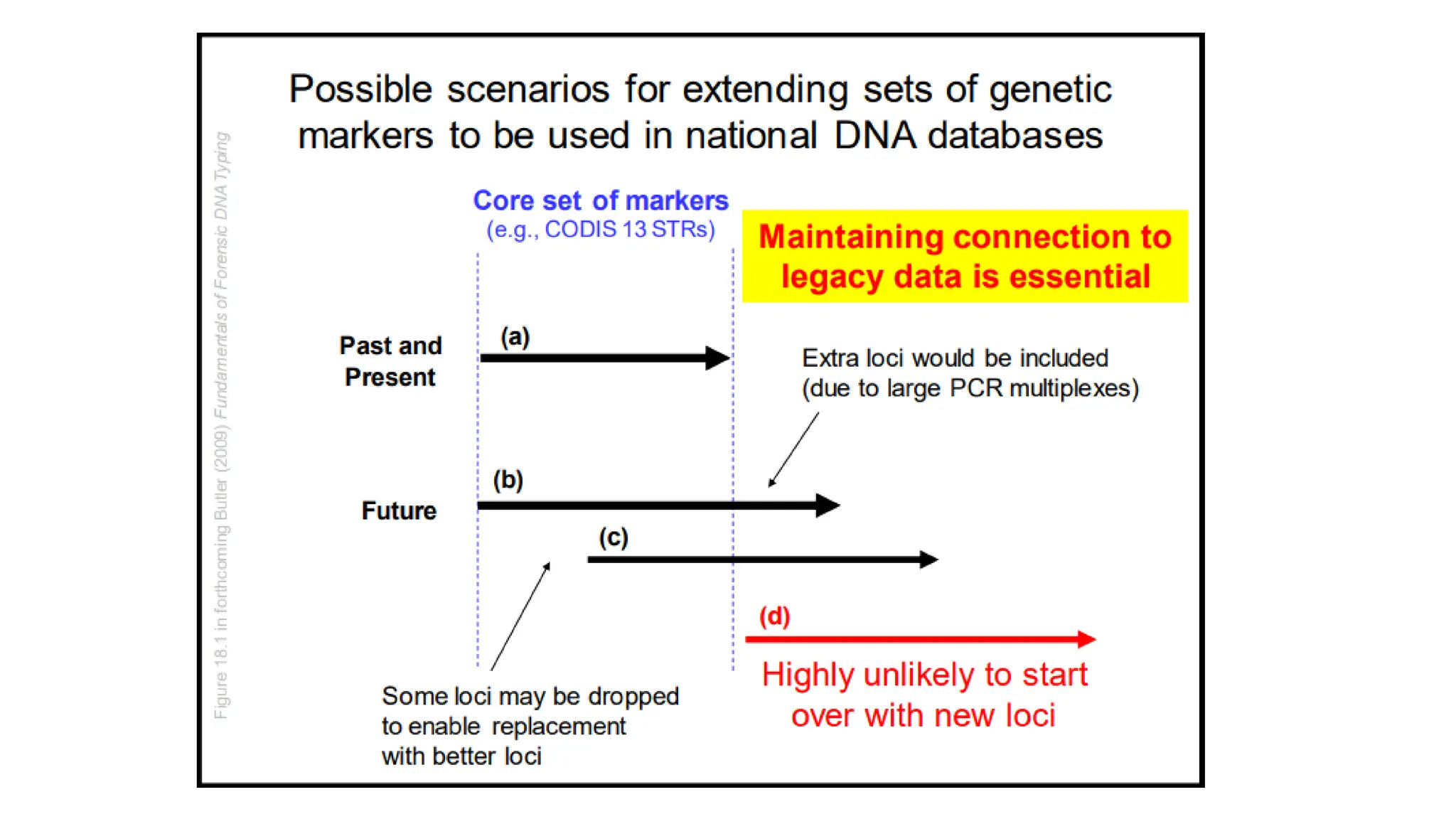
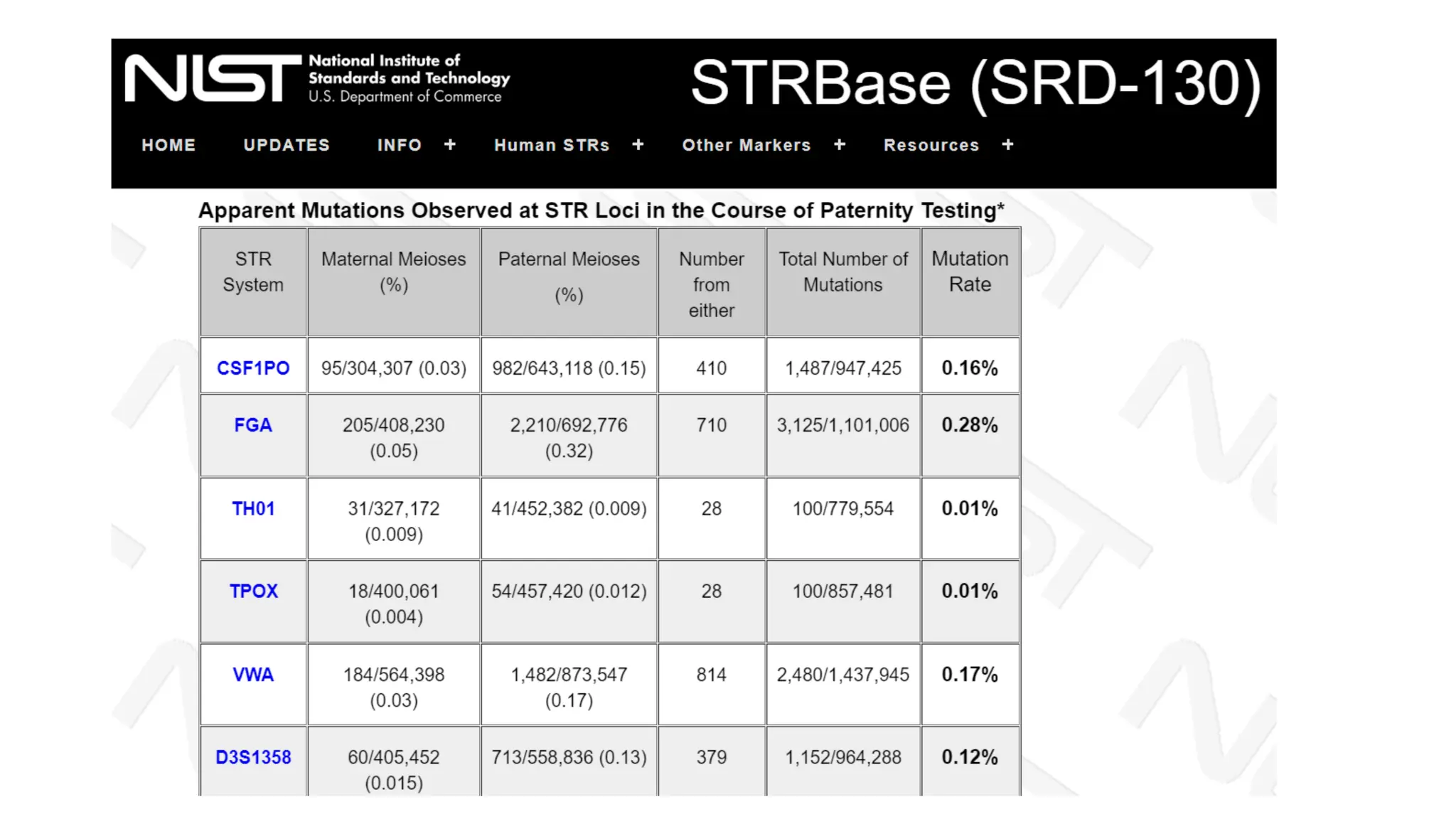
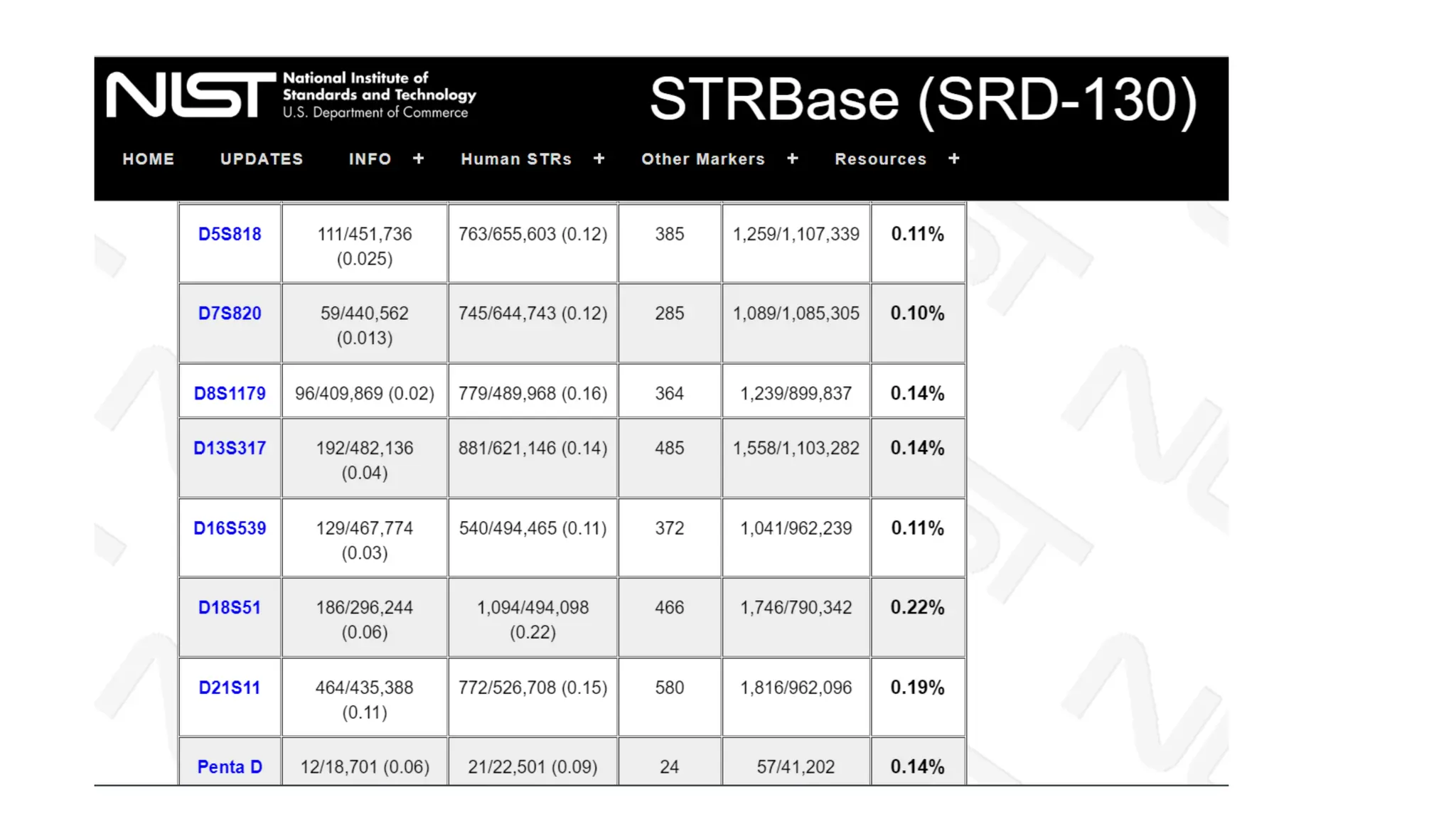
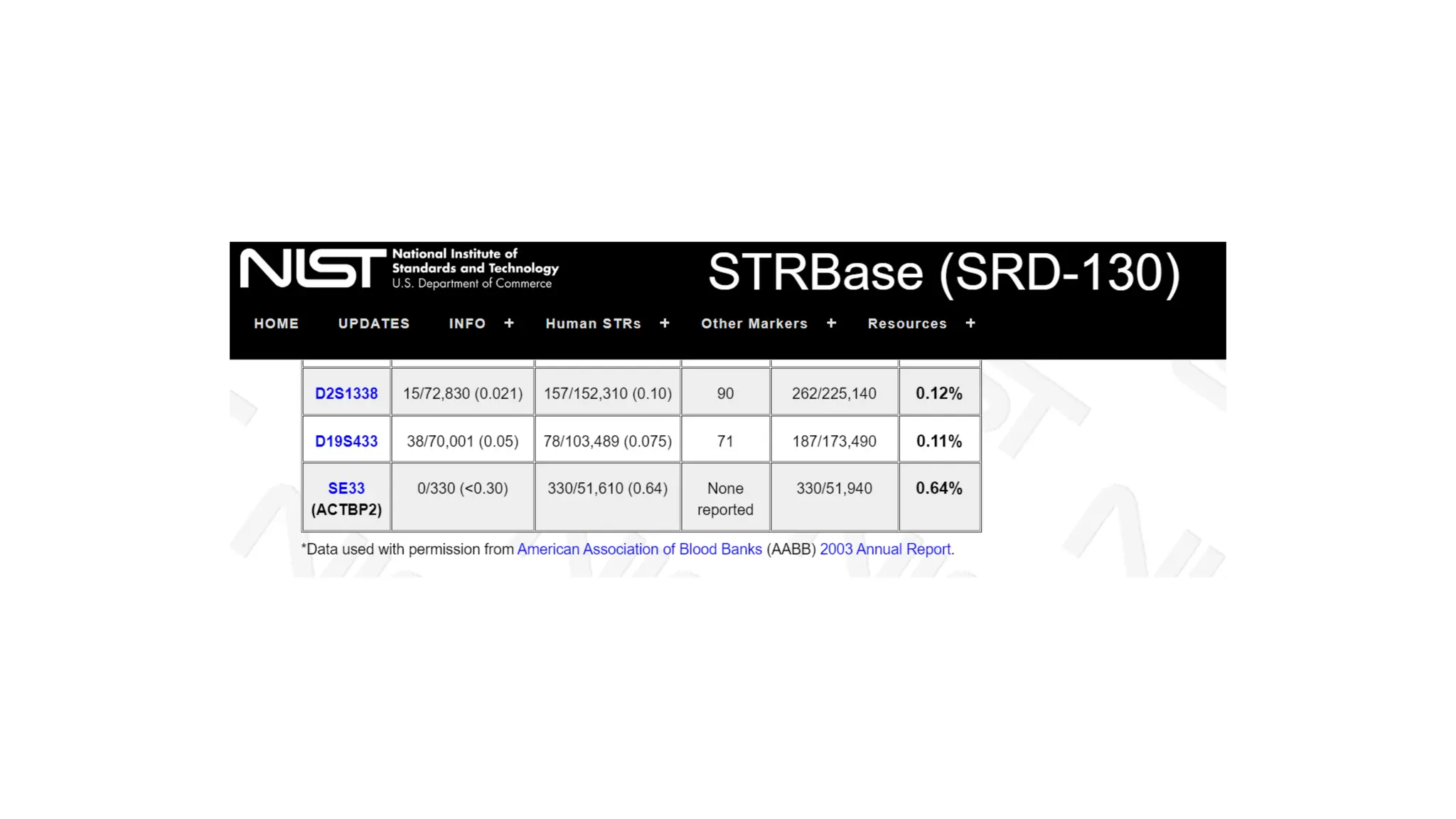
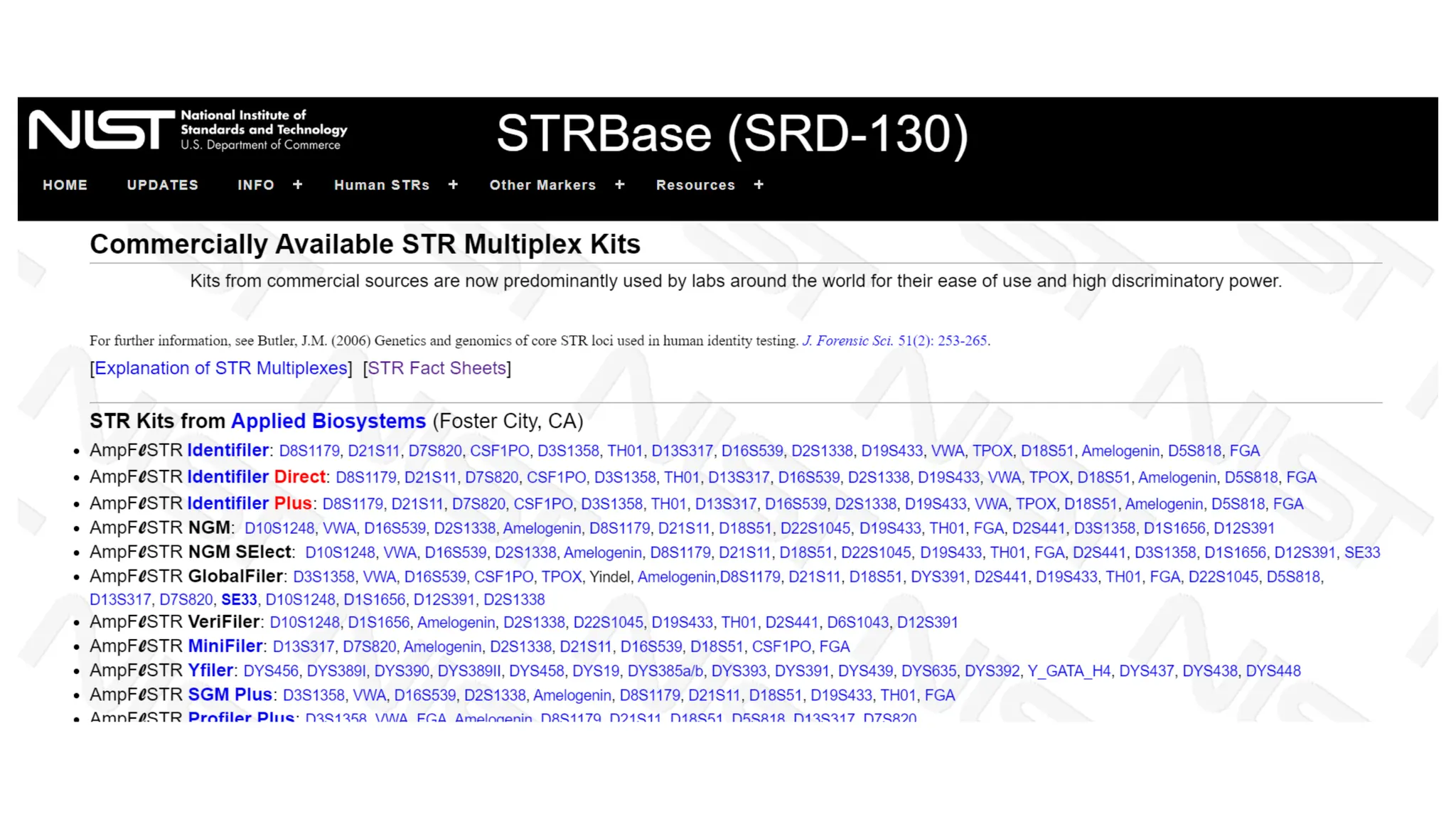
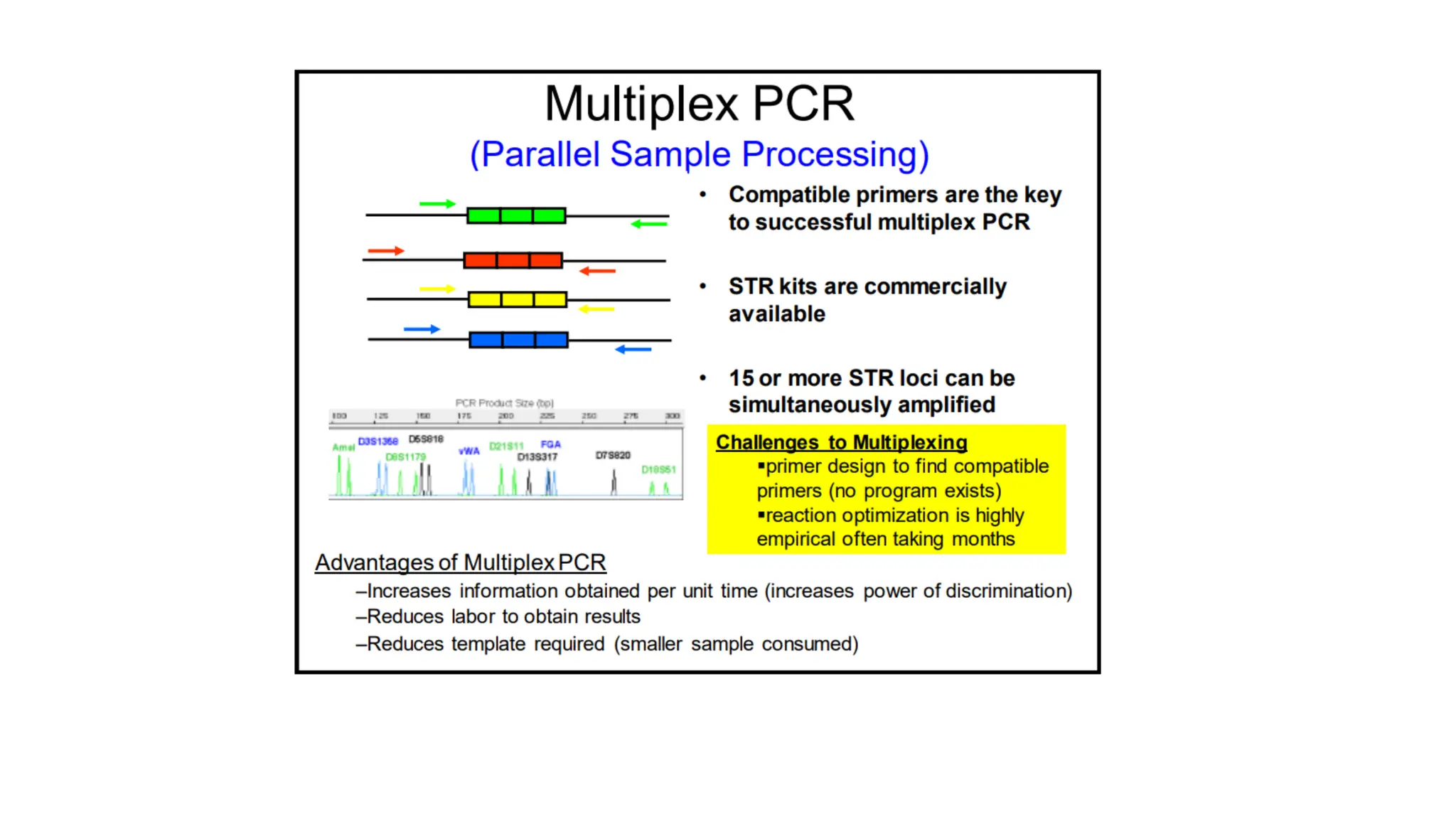
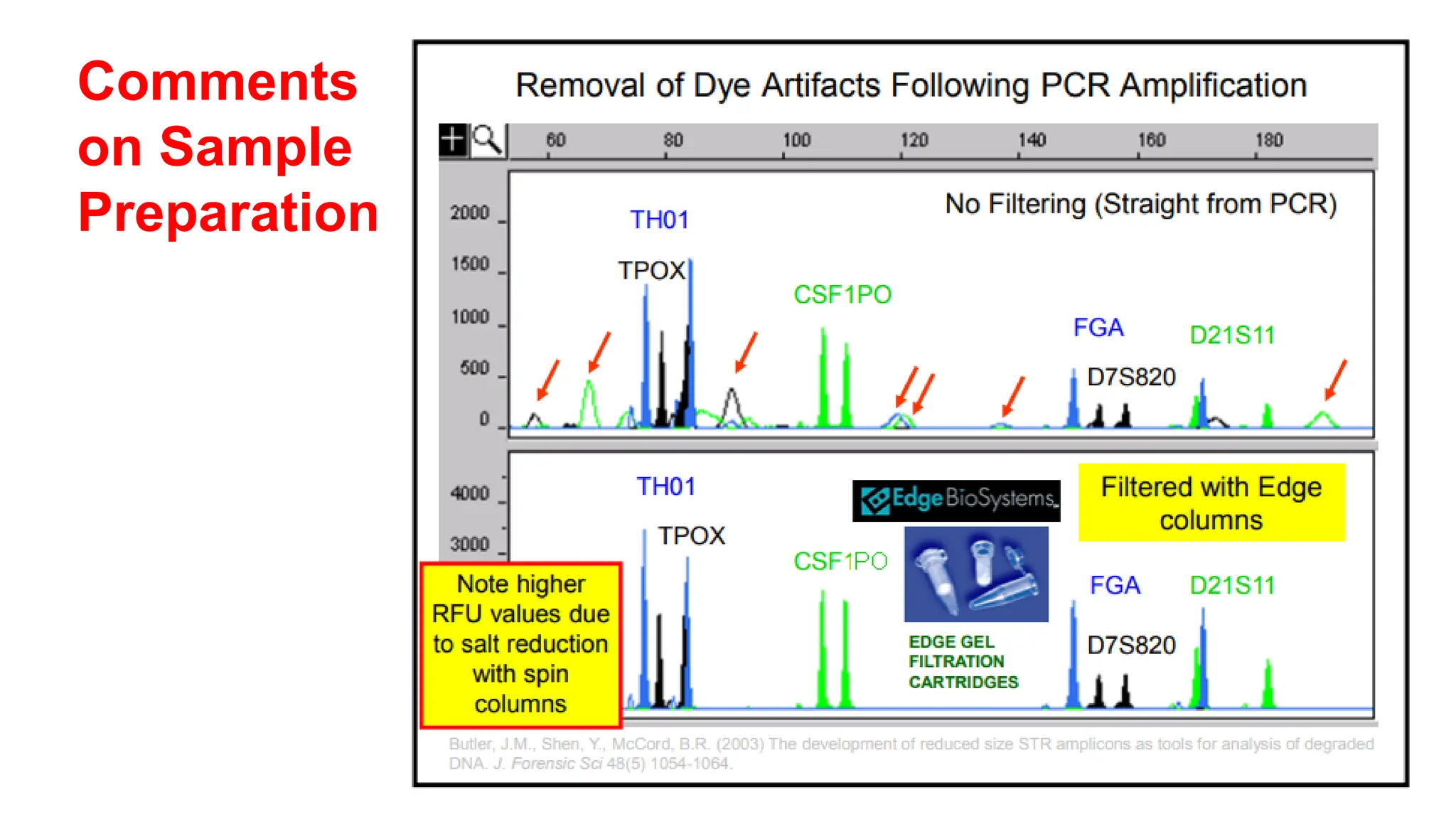
The document discusses DNA profiling using short tandem repeats (STRs) with a focus on selecting loci that have high discriminating power and stability in evidence samples. It highlights a consensus panel of core loci recommended by the Scientific Working Group for DNA Analysis Methods (SWGDAM) and factors influencing the selection of loci for multiplex PCR. Additionally, it addresses the issue of spectral artifacts such as peak pull-up caused by high peak heights in DNA analysis software.

































![Repeat: [CA]n
Repeat: [AGAGAT]n[AGAGAT]m](https://image.slidesharecdn.com/mscivsemesterdnaprofiling-strbiologyandartifacts-240515082426-9bd80228/75/MSC-IV-SEMESTER_DNA-Profiling-STR-biology-and-artifacts-pdf-34-2048.jpg)
![Repeat: [TATC]
Repeat: [TCTA], [TCTG] Repeat: [GAAA]](https://image.slidesharecdn.com/mscivsemesterdnaprofiling-strbiologyandartifacts-240515082426-9bd80228/75/MSC-IV-SEMESTER_DNA-Profiling-STR-biology-and-artifacts-pdf-35-2048.jpg)






































