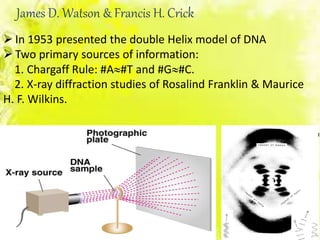
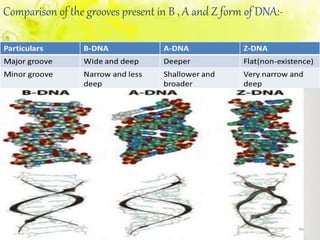
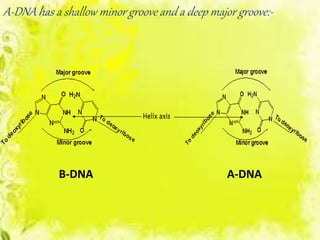
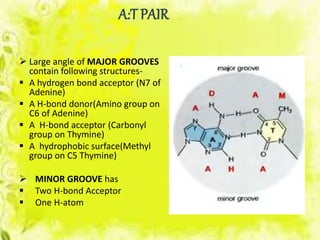
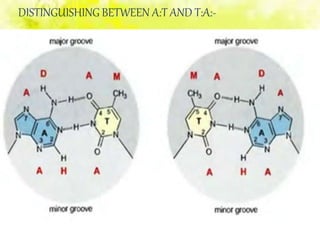
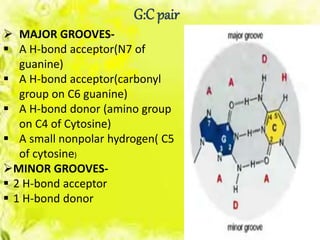
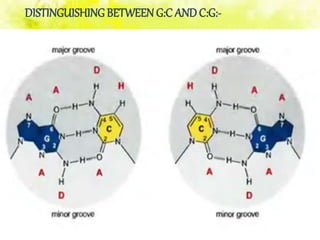
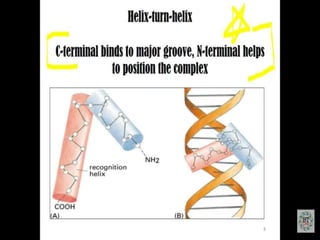
The document discusses the major and minor grooves of DNA, highlighting their significance in genetic processes and the binding of proteins. Major grooves contain more chemical information and are preferred by specific DNA-binding proteins, while minor grooves are involved in non-specific binding. Additionally, the structural characteristics and differences between the grooves in various DNA forms (A, B, and Z) are examined.