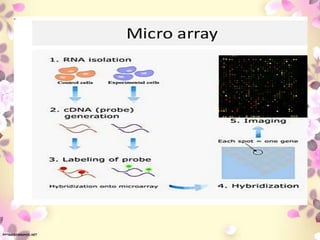
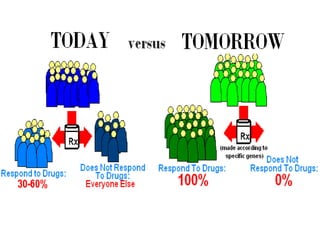
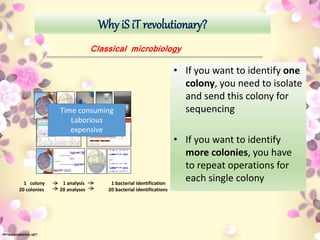
Functional genomics, pharmacogenomics, and metagenomics are methods for determining the roles of genes. Functional genomics examines gene function through techniques like gene knockouts, microarrays, and RNA sequencing. Pharmacogenomics studies how genes affect individual responses to drugs in terms of metabolism and efficacy. Metagenomics analyzes the collective genomes of microbial communities through direct environmental DNA sequencing without requiring isolation of individual species.