A Theoretical Model of Repression of Transcription by Nucleoid Proteins
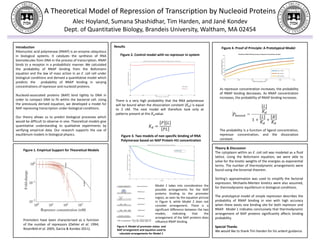
- 1. A Theoretical Model of Repression of Transcription by Nucleoid Proteins Alec Hoyland, Sumana Shashidhar, Tim Harden, and Jané Kondev Dept. of Quantitative Biology, Brandeis University, Waltham, MA 02454 Introduction Ribonucleic acid polymerase (RNAP) is an enzyme ubiquitous in biological systems. It catalyzes the synthesis of RNA biomolecules from DNA in the process of transcription. RNAP binds to a receptor in a probabilistic manner. We calculated the probability of RNAP binding from the Boltzmann equation and the law of mass action in an E. coli cell under biological conditions and derived a quantitative model which predicts the probability of RNAP binding in varying concentrations of repressor and nucleoid proteins. Nucleoid-associated proteins (NAP) bind tightly to DNA in order to compact DNA to fit within the bacterial cell. Using the previously derived equation, we developed a model for NAP repressing transcription under biological conditions. Our theory allows us to predict biological processes which would be difficult to observe in vivo. Theoretical models give quantitative understanding to qualitative experiments by verifying empirical data. Our research supports the use of equilibrium models in biological physics. Promoters have been characterized as a function of the number of repressors (Oehler et al. 1994; Rosenfeld et al. 2005; Garcia & Kondev 2011). As repressor concentration increases, the probability of RNAP binding decreases. As RNAP concentration increases, the probability of RNAP binding increases. Figure 4. Proof of Principle: A Prototypical Model Figure 1. Empirical Support for Theoretical Models There is a very high probability that the RNA polymerase will be bound when the dissociation constant (𝐾 𝑑) is equal to 3 nM. The next model will therefore look only at patterns present at this 𝐾 𝑑value. 𝐾 𝑑 = 𝑃 𝐿 𝑃𝐿 Figure 3. Two models of non specific binding of RNA Polymerase based on NAP Protein HU concentration Figure 2. Control model with no repressor in system Model 1 takes into consideration the possible arrangements for the NAP proteins binding to the promoter region, as seen by the equation picture in Figure 4, while Model 2 does not consider arrangement. There is a significant difference between the two models, indicating that the arrangement of the NAP proteins does influence RNAP binding. Theory & Discussion The cytoplasm within an E. coli cell was modeled as a fluid lattice. Using the Boltzmann equation, we were able to solve for the kinetic weights of the energies as exponential terms. The number of thermodynamic arrangements were found using the binomial theorem. Stirling’s approximation was used to simplify the factorial expression. Michaelis-Menten kinetics were also assumed, for thermodynamic equilibrium in biological conditions. The prototypical model of simple repression describes the probability of RNAP binding in vivo with high accuracy when there exists one binding site for both repressor and RNAP. Model 1 indicates conclusively that thermodynamic arrangement of NAP proteins significantly affects binding probability. Special Thanks We would like to thank Tim Harden for his ardent guidance. The probability is a function of ligand concentration, repressor concentration, and the dissociation constant. Results Figure 4. Model of promoter states and NAP arrangement and equation used to calculate arrangements for Model 1