End-point zygosity and CNV determination from crude samples
End-point zygosity and CNV determination based on post PCR Rn/ΔRn reading is expected to reduce the cost and increase throughput as compared to current real time Cq-based measurement. There is an immediate need by AgBio customers to screen seed zygosity and copy number of transgenes through end-point reading. Moreover, Agbio customers expect to determine zygosity and CNV using crude samples in order to stream-line their workflow. For endpoint quantification, the major challenge is to control the saturation of PCR to maintain the segregation of end-point intensity according to the input copy number of the target transgenes in reference to a control gene. We have tried a few approaches such as Asymmetric PCR, ARCS (Amplification Ratio Control System) PCR, Dual Tailing PCR, pyrophosphate removal and controlling plateau of PCR by running a third PCR in the background. Among the tested approaches, controlled plateau of PCR (CoP’ed PCR) showed best separation of copy number through end-point PCR reading. We further optimized CoP’ed PCR conditions using purified DNA and crude plant and blood samples. By CoP’ed PCR, we are able to do zygosity with crude plant samples to satisfy the immediate needs for Agbio customers. Furthermore, since this approach improves the sensitivity for the copy number separation for not only end-point but also real-time PCR for crude blood samples, in the future, it can be used for CNV analysis directly from human blood.
Recommended
Recommended
More Related Content
What's hot
What's hot (20)
Similar to End-point zygosity and CNV determination from crude samples
Similar to End-point zygosity and CNV determination from crude samples (20)
More from Thermo Fisher Scientific
More from Thermo Fisher Scientific (20)
Recently uploaded
Recently uploaded (20)
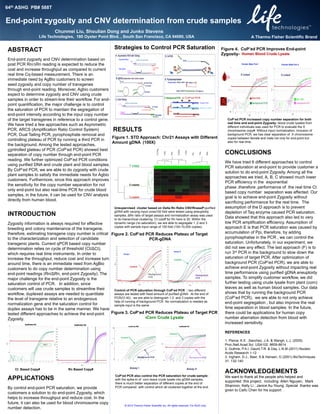
End-point zygosity and CNV determination from crude samples
- 1. A Thermo Fisher Scientific Brand End-point zygosity and CNV determination from crude samples or CoP’ed PCR © 2014 Thermo Fisher Scientific Inc. All rights reserved. For RUO only. 64th ASHG PB# 588T Chunmei Liu, Shoulian Dong and Junko Stevens Life Technologies, 180 Oyster Point Blvd. , South San Francisco, CA 94080, USA ABSTRACT End-point zygosity and CNV determination based on post PCR Rn/ΔRn reading is expected to reduce the cost and increase throughput as compared to current real time Cq-based measurement. There is an immediate need by AgBio customers to screen seed zygosity and copy number of transgenes through end-point reading. Moreover, Agbio customers expect to determine zygosity and CNV using crude samples in order to stream-line their workflow. For end-point quantification, the major challenge is to control the saturation of PCR to maintain the segregation of end-point intensity according to the input copy number of the target transgenes in reference to a control gene. We have tried a few approaches such as Asymmetric PCR, ARCS (Amplification Ratio Control System) PCR, Dual Tailing PCR, pyrophosphate removal and controlling plateau of PCR by running a third PCR in the background. Among the tested approaches, controlled plateau of PCR (CoP’ed PCR) showed best separation of copy number through end-point PCR reading. We further optimized CoP’ed PCR conditions using purified DNA and crude plant and blood samples. By CoP’ed PCR, we are able to do zygosity with crude plant samples to satisfy the immediate needs for Agbio customers. Furthermore, since this approach improves the sensitivity for the copy number separation for not only end-point but also real-time PCR for crude blood samples, in the future, it can be used for CNV analysis directly from human blood. INTRODUCTION Zygosity information is always required for effective breeding and colony maintenance of the transgene, therefore, estimating transgene copy number is critical to the characterization and selection of candidate transgenic plants. Current qPCR based copy number determination relies on cycle of threshold (Ct/ΔCt), which requires real time instruments. In order to increase the throughput, reduce cost and increase turn around time, there is an immediate need from AgBio customers to do copy number determination using end-point readings (Rn/ΔRn, end-point Zygosity). The major challenge for the end-point Zygosity is the saturation control of PCR. In addition, since customers will use crude samples to streamline their workflow, duplexed assays are needed to quantitate the level of transgene relative to an endogenous normalization gene and the saturation control for duplex assays has to be in the same manner. We have tested different approaches to achieve the end-point Zygosity. 3,2,1 Ct Based Copy# Rn Based Copy# APPLICATIONS By control end-point PCR saturation, we provide customers a solution to do end-point Zygosity, which helps to increase throughput and reduce cost. In the future, it can also be used for blood chromosome copy number detection. Figure 4. CoP’ed PCR Improves End-point Zygosity- Human Blood Crude Lysate CoP’ed PCR increased copy number separation for both real time and end-point Zygosity: blood crude lysates from different individuals was used for PCR to evaluate the X chromosome copy#. Without input normalization, inclusion of background PCR, we has clear separation of X chromosome copies between female and male not only for end-point but also for real time. CONCLUSIONS We have tried 6 different approaches to control PCR saturation at end-point to provide customer a solution to do end-point Zygosity. Among all the approaches we tried, A, B, C showed much lower PCR efficiency in the exponential phase ,therefore ,performance of the real time Ct based copy number separation was affected. Our goal is to achieve end-point Zygosity without sacrificing performance for the real time. The assumption of the D approach is to prevent depletion of Taq enzyme caused PCR saturation. Data showed that this approach also led to very low PCR amplification efficiency. Assumption of approach E is that PCR saturation was caused by accumulation of Ppi, therefore, by adding pyrophosphatse in the PCR , we can control the saturation. Unfortunately, in our experiment, we did not see any effect. The last approach (F) is to run 3rd PCR in the background to slow down the saturation of target PCR. After optimization of background PCR (CoP’ed PCR), we are able to achieve end-point Zygosity without impacting real time performance using purified gDNA aneuploidy samples. To simplify customer workflow, we did further testing using crude lysate from plant (corn) leaves as well as human blood samples. Our data shows that by running the background PCR (CoP’ed PCR), we are able to not only achieve end-point segregation , but also improve the real time separation in blood samples. In the future, there could be applications for human copy number aberration detection from blood with increased sensitivity. REFERENCES 1. Pierce, K.E. ,Sanchez, J.A. & Wangh, L.J. (2005) Proc.Natl.Acad.Sci. USA102, 8609-8614 2. Guthrie, P.A.I, Gaunt,T.R. & Day, L.N.M (2011) Nucleic Acids Research 1-12 3. Ingham, D.J., Beer, S.& Hansen, G (2001) BioTechniques 31: 132-140 ACKNOWLEDGEMENTS We want to thank all the people who helped and supported this project, including Allen Nguyen, Mark Shannon, Kelly Li , Janice Au-Young .Special thanks was given to Caifu Chen for his support . Strategies to Control PCR Saturation RESULTS Figure 1. STD Approach: Chr21 Assays with Different Amount gDNA (100X) Unsupervised cluster based on Delta Rn Ratio CNV/RnaseP:purified gDNA with varying input cross100 fold were tested using aneuploidy samples. ΔRn ratio of target assays and normalization assay was used to do hierarchical clustering. Ct cutoff for Rn here is 30. Within the dynamic range (no saturation), we are able to segregate 1 ,2 and 3 copies with sample input range of 100 fold (100-10,000 copies). Figure 2. CoP’ed PCR Reduces Plateau of Target PCR-gDNA Control of PCR saturation through CoP’ed PCR : two different assays are tested with fixed amount of purified gDNA. At the end of PCR(Ct 40) , we are able to distinguish 1,2 and 3 copies with the help of running of background PCR. No normalization is needed as sample input is the same . Figure 3. CoP’ed PCR Reduces Plateau of Target PCR -Corn Crude Lysate CoP’ed PCR also control the PCR saturation for crude sample: with the spike-in of corn leave crude lysate into gDNA samples, there is much better separation of different copies at the end of PCR compared with control which all clustered together at the end.
