The document provides a comprehensive overview of Wikipedia's evolution as a major online encyclopedia that enables collaborative editing and curation of knowledge. It discusses the integration of linked open data through projects like Wikidata and WikiCite, emphasizing their roles in supporting scientific research and enhancing citation practices. Additionally, it addresses the challenges of bias and coverage while highlighting the potential for distributed fact-checking and new forms of knowledge discovery.












![The backbone of the linked open data ecosystem
Schmachtenberg et al
(2014)
http://lod-cloud.net [CC BY SA]](https://image.slidesharecdn.com/vivo2016keynoteverifiablelinkedopenknowledge-160819162835/75/Verifiable-linked-open-knowledge-that-anyone-can-edit-15-2048.jpg)






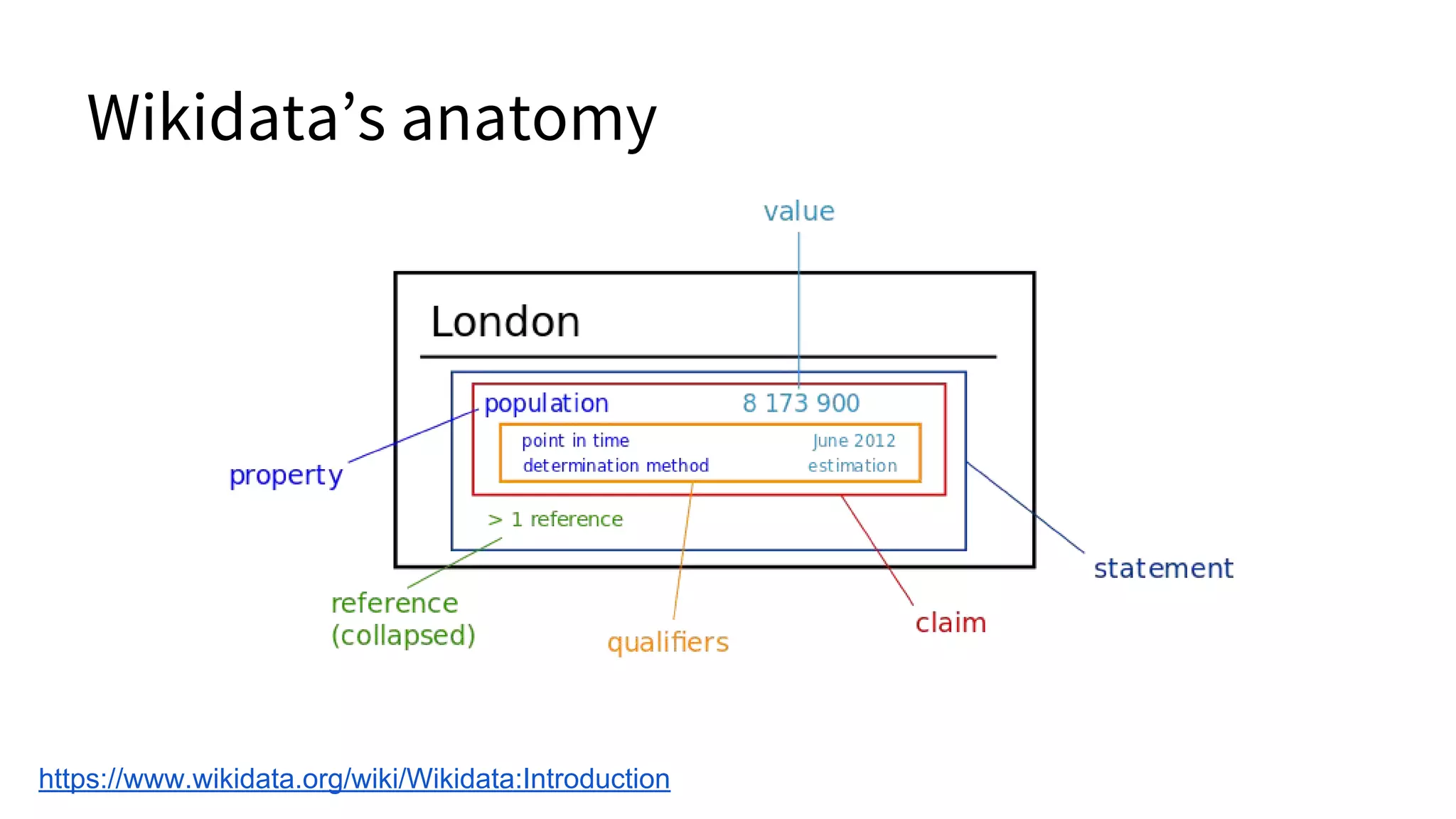
![Wikidata’s anatomy
Linked data, San Francisco, Jeblad
https://commons.wikimedia.org/wiki/File:Linked_Data_-_San_Francisco.svg [CC BY SA]](https://image.slidesharecdn.com/vivo2016keynoteverifiablelinkedopenknowledge-160819162835/75/Verifiable-linked-open-knowledge-that-anyone-can-edit-25-2048.jpg)


![WikiCite
Building the sum of all human citations
Randall Munroe, Wikipedian protester http://tinyurl.com/p3rodlb [CC BY]](https://image.slidesharecdn.com/vivo2016keynoteverifiablelinkedopenknowledge-160819162835/75/Verifiable-linked-open-knowledge-that-anyone-can-edit-31-2048.jpg)
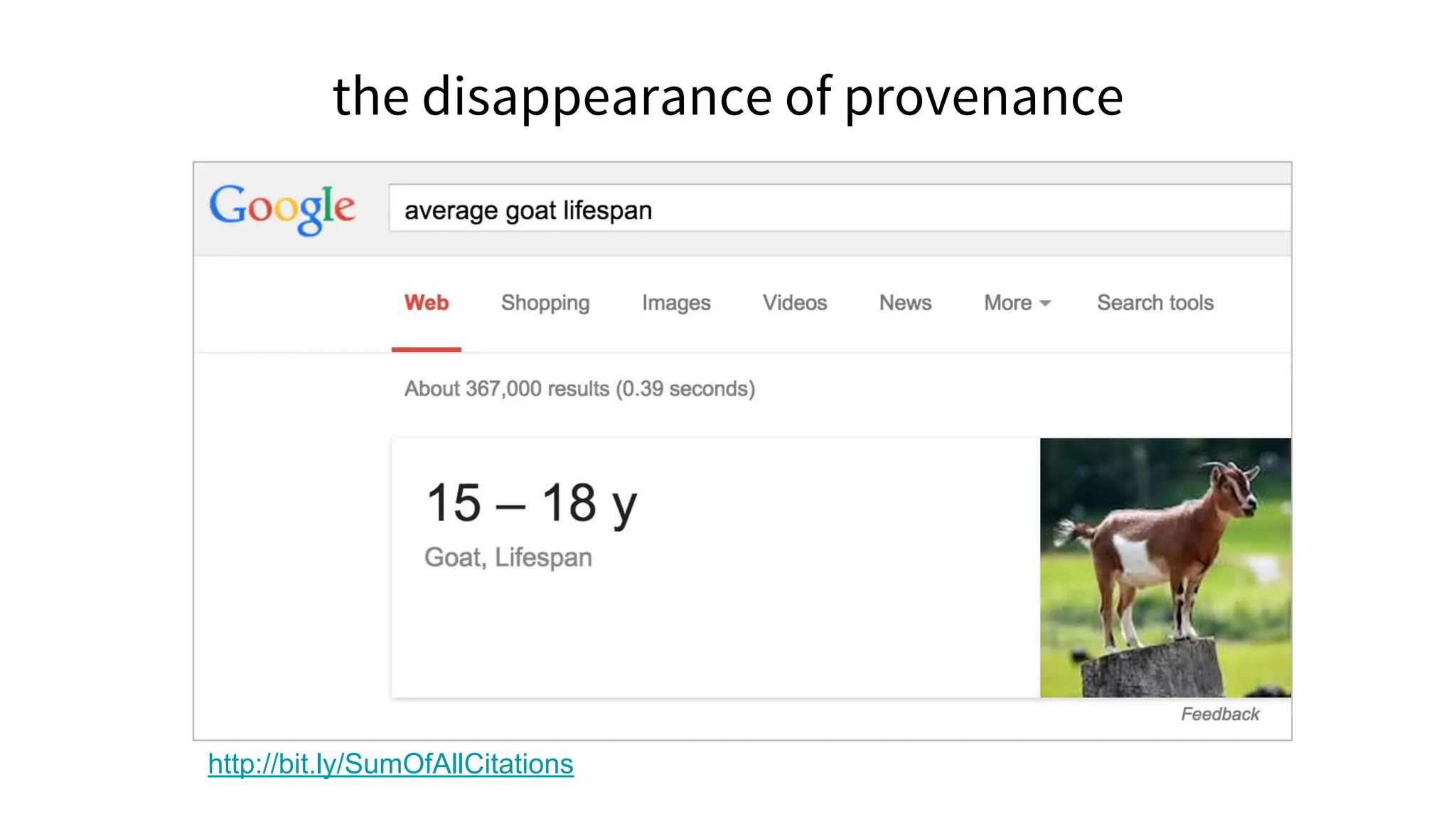
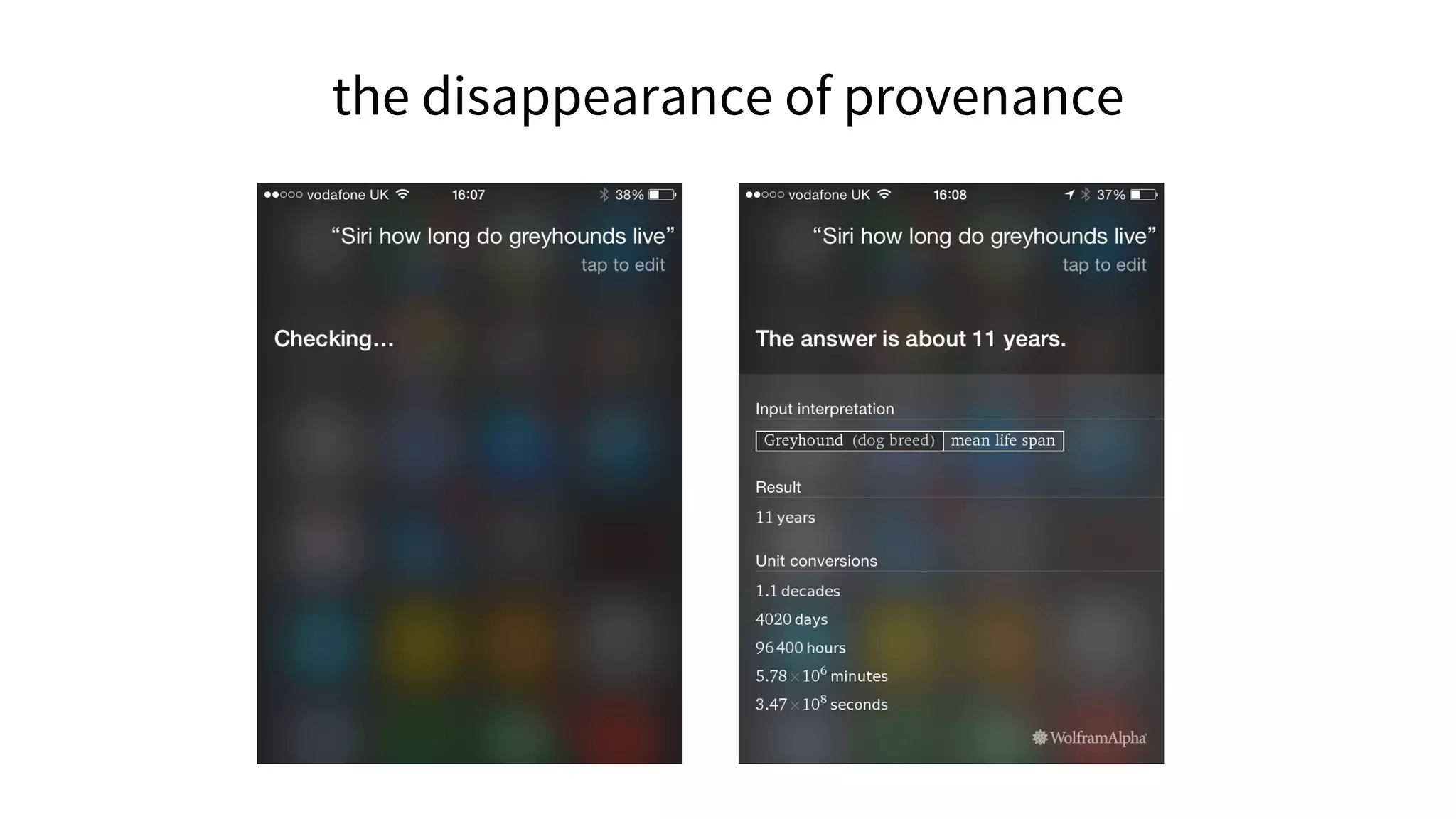


![Linking is a small act of generosity that sends people away from
your site to some other that you think shows the world in a way
worth considering. [...]
[Sources] that are not generous with linking [...] are a stopping
point in the ecology of information. That’s the operational
definition of authority: The last place you visit when you’re
looking for an answer. If you are satisfied with the answer, you
stop your pursuit of it. Take the links out and you think you look
like more of an authority.
D. Weinberger (2012) Linking is a public good
http://www.hyperorg.com/blogger/2012/02/26/2b2k-linking-is-a-public-good/](https://image.slidesharecdn.com/vivo2016keynoteverifiablelinkedopenknowledge-160819162835/75/Verifiable-linked-open-knowledge-that-anyone-can-edit-36-2048.jpg)

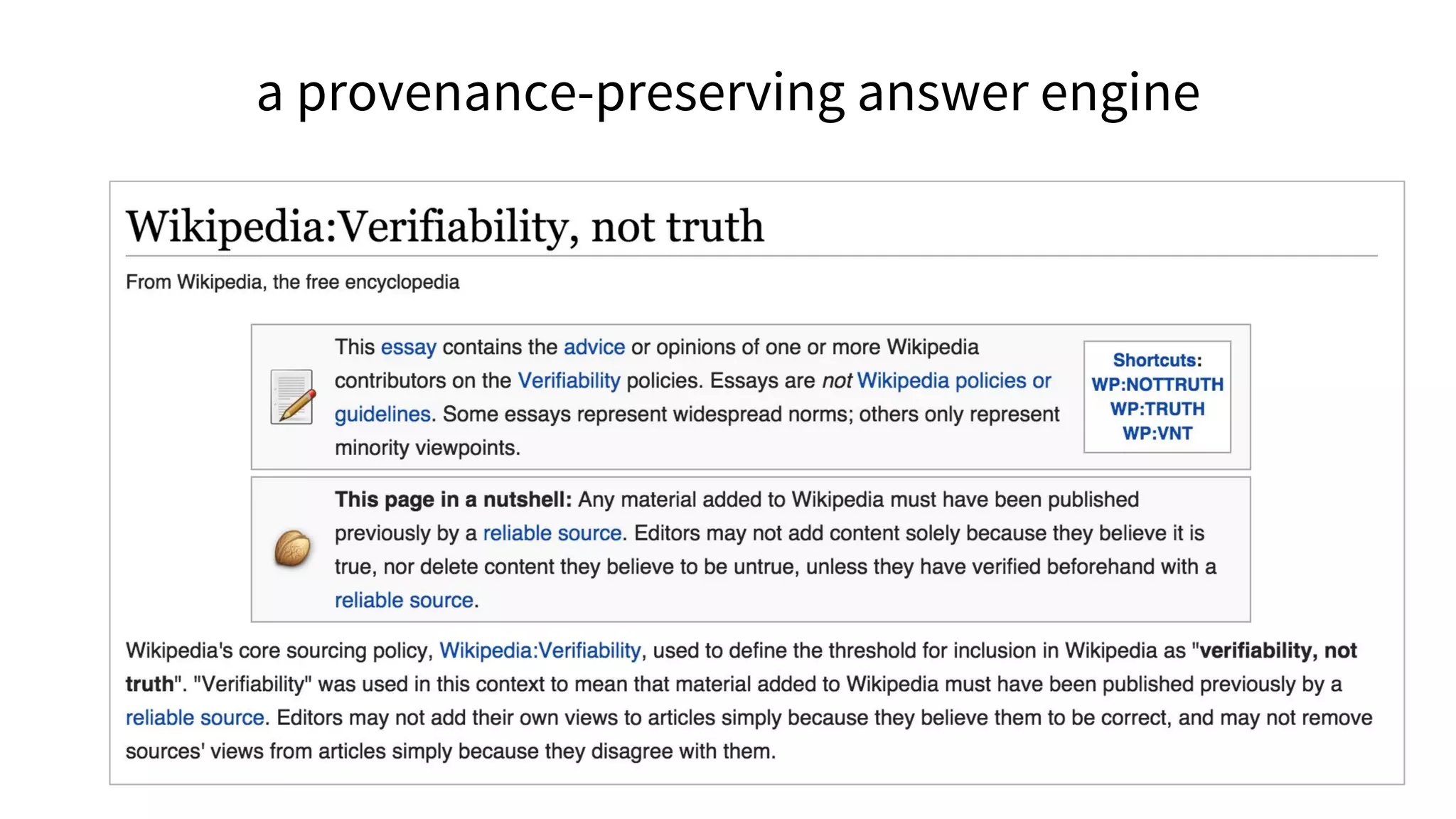
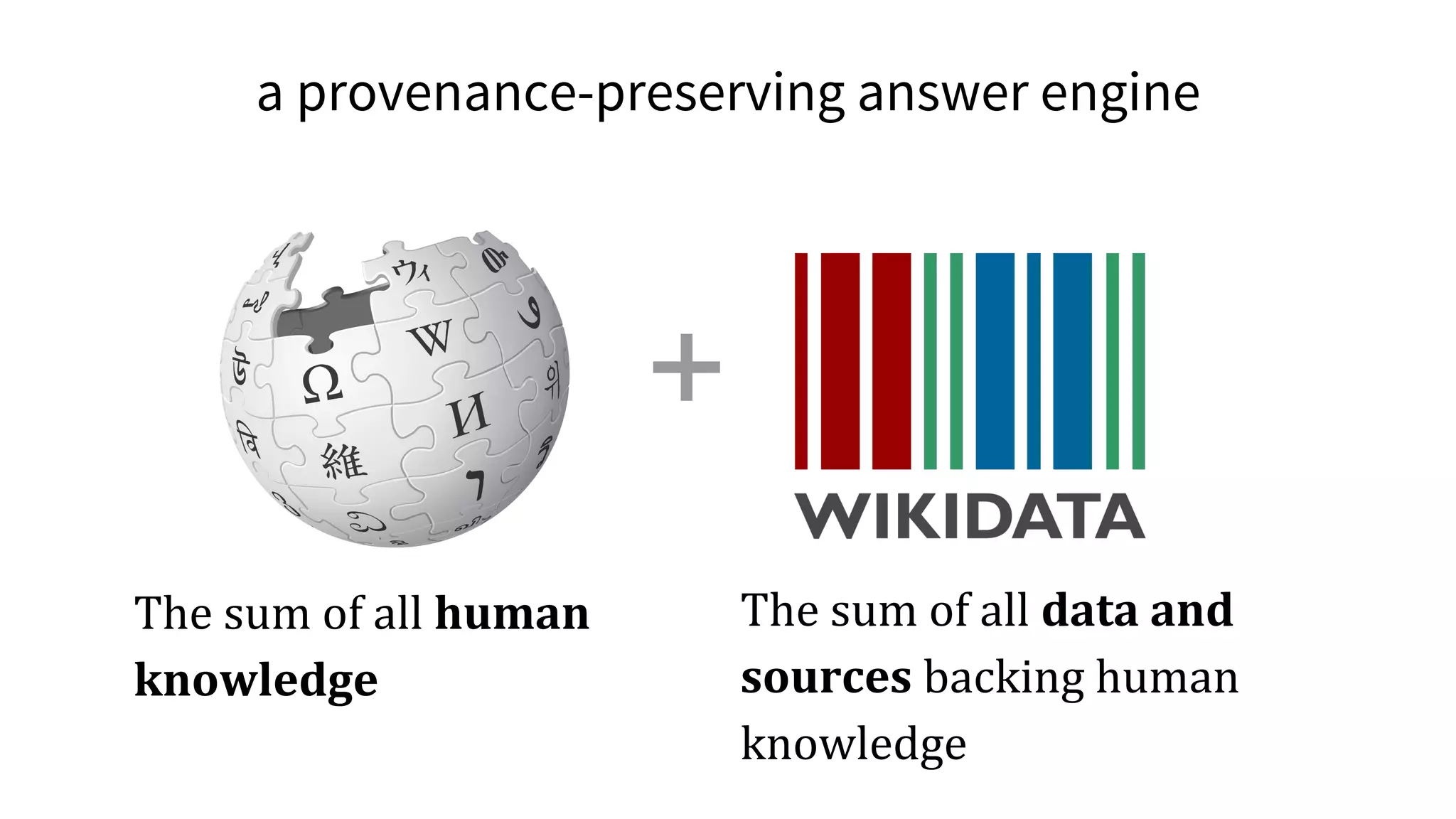

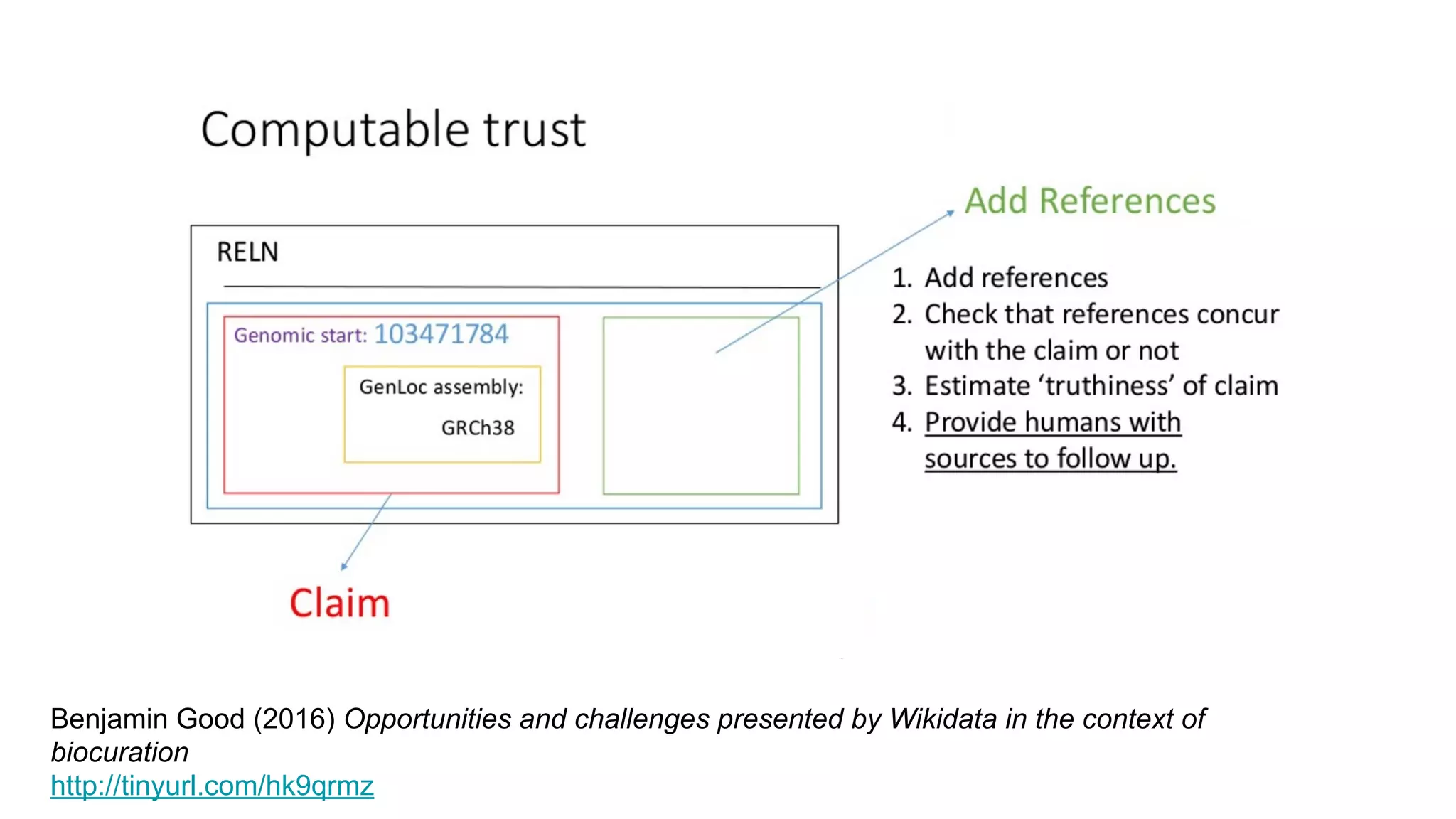
![The molecular origins of insulin go at least as far back as
the simplest unicellular [[eukaryotes]].<ref
name='LeRoith'>{{cite journal | vauthors = LeRoith D, Shiloach
J, Heffron R, Rubinovitz C, Tanenbaum R, Roth J | title =
Insulin-related material in microbes: similarities and
differences from mammalian insulins | journal = Can. J.
Biochem. Cell Biol. | volume = 63 | issue = 8 | pages = 839–49
| year = 1985 | pmid = 3933801 | doi = 10.1139/o85-106
}}</ref> Apart from animals, insulin-like proteins are also
known to exist in Fungi and Protista kingdoms.
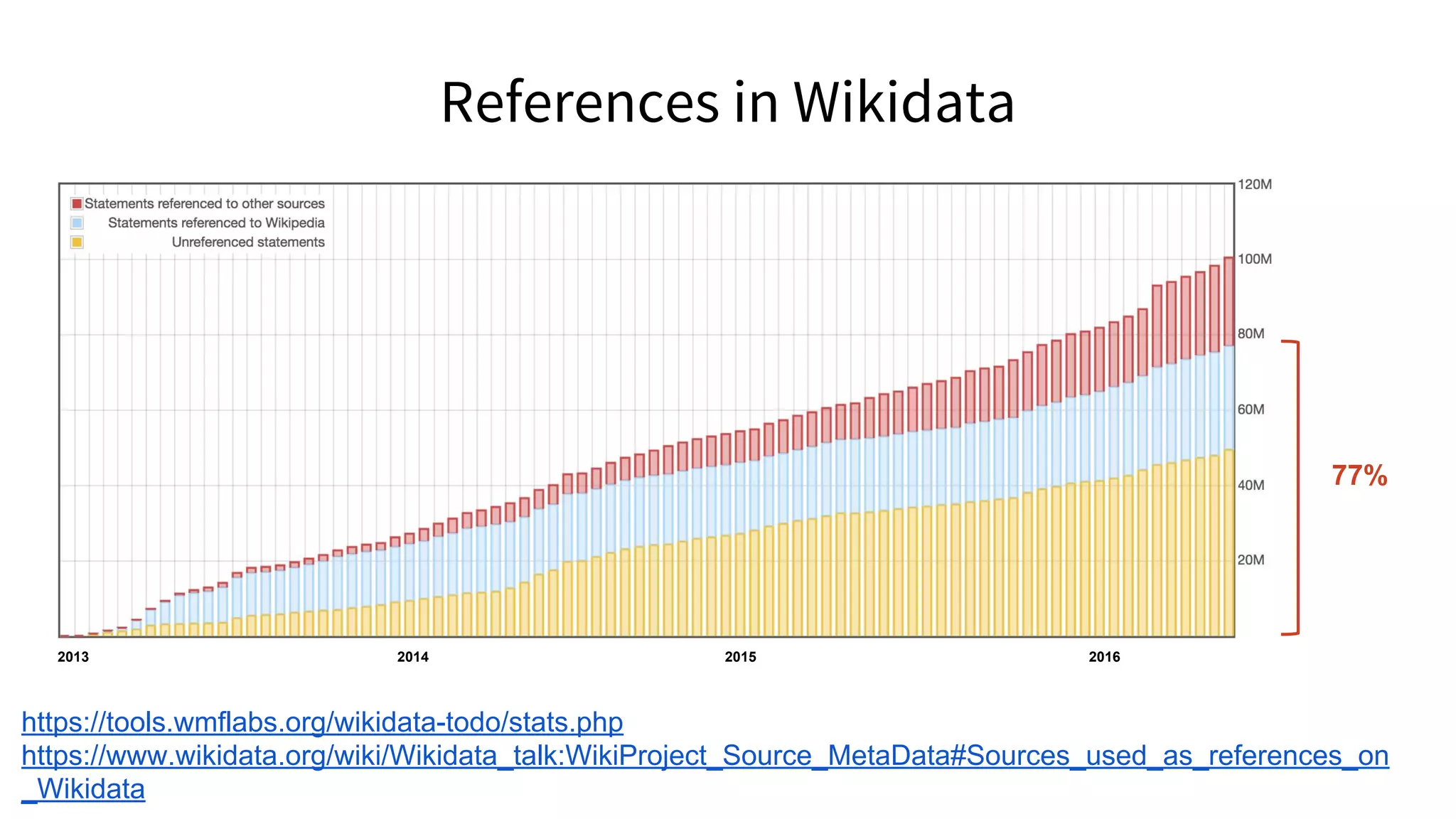
References in Wikipedia](https://image.slidesharecdn.com/vivo2016keynoteverifiablelinkedopenknowledge-160819162835/75/Verifiable-linked-open-knowledge-that-anyone-can-edit-43-2048.jpg)









![Thank you
Acknowledgments
Daniel Mietchen, Jonathan Dugan, Lydia Pintscher, Cameron Neylon, James Hare, James Heilman,
Magnus Manske, the Gene Wiki team (especially Andra Waagmeester and Benjamin Good), the
University of Chicago Knowledge Lab, all WikiCite 2016 participants and Wikidata Source Metadata
project contributors.
Additional image credits
Printing press, M. Wirth https://thenounproject.com/term/printing/11880/ [CC BY]
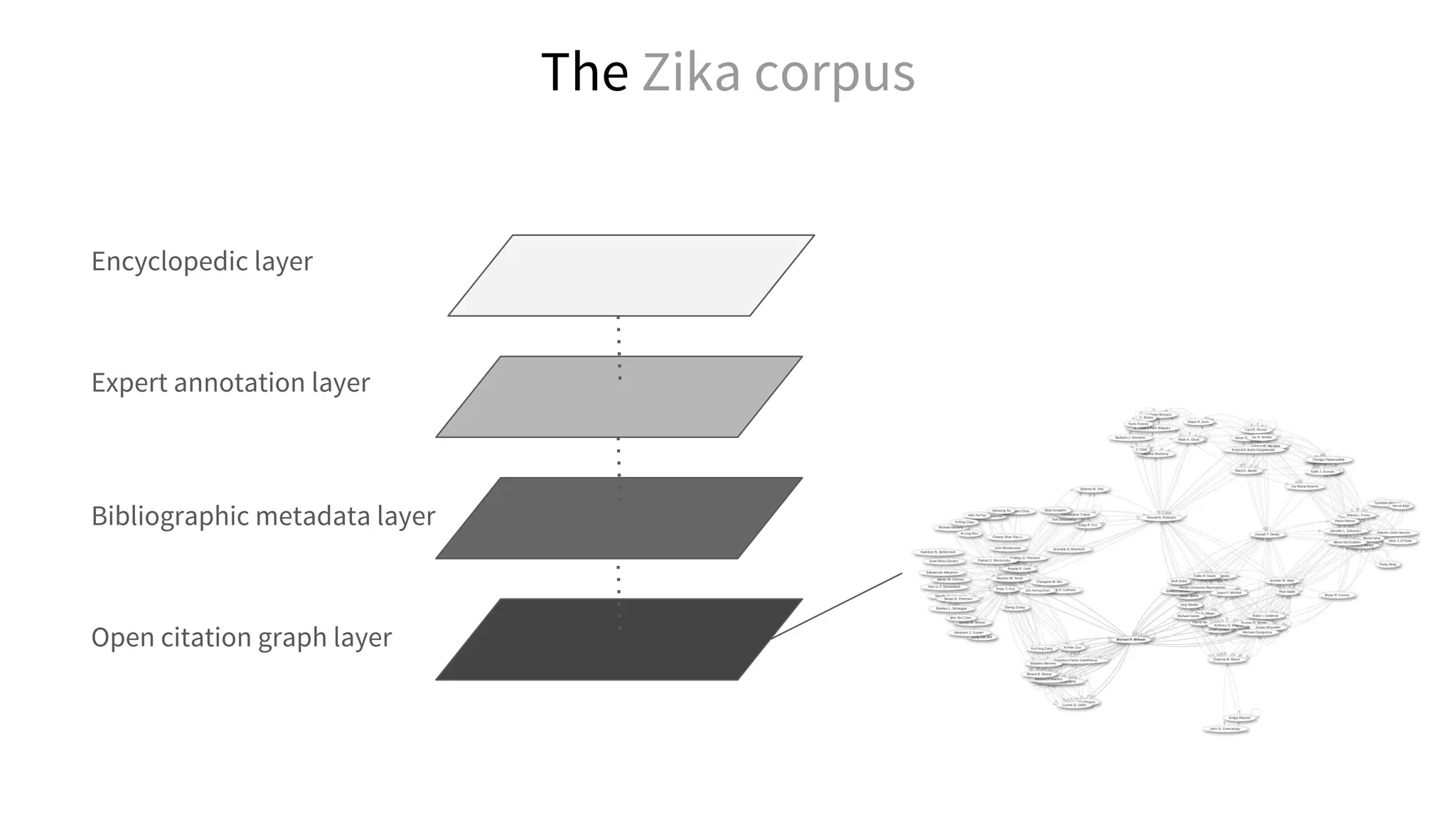
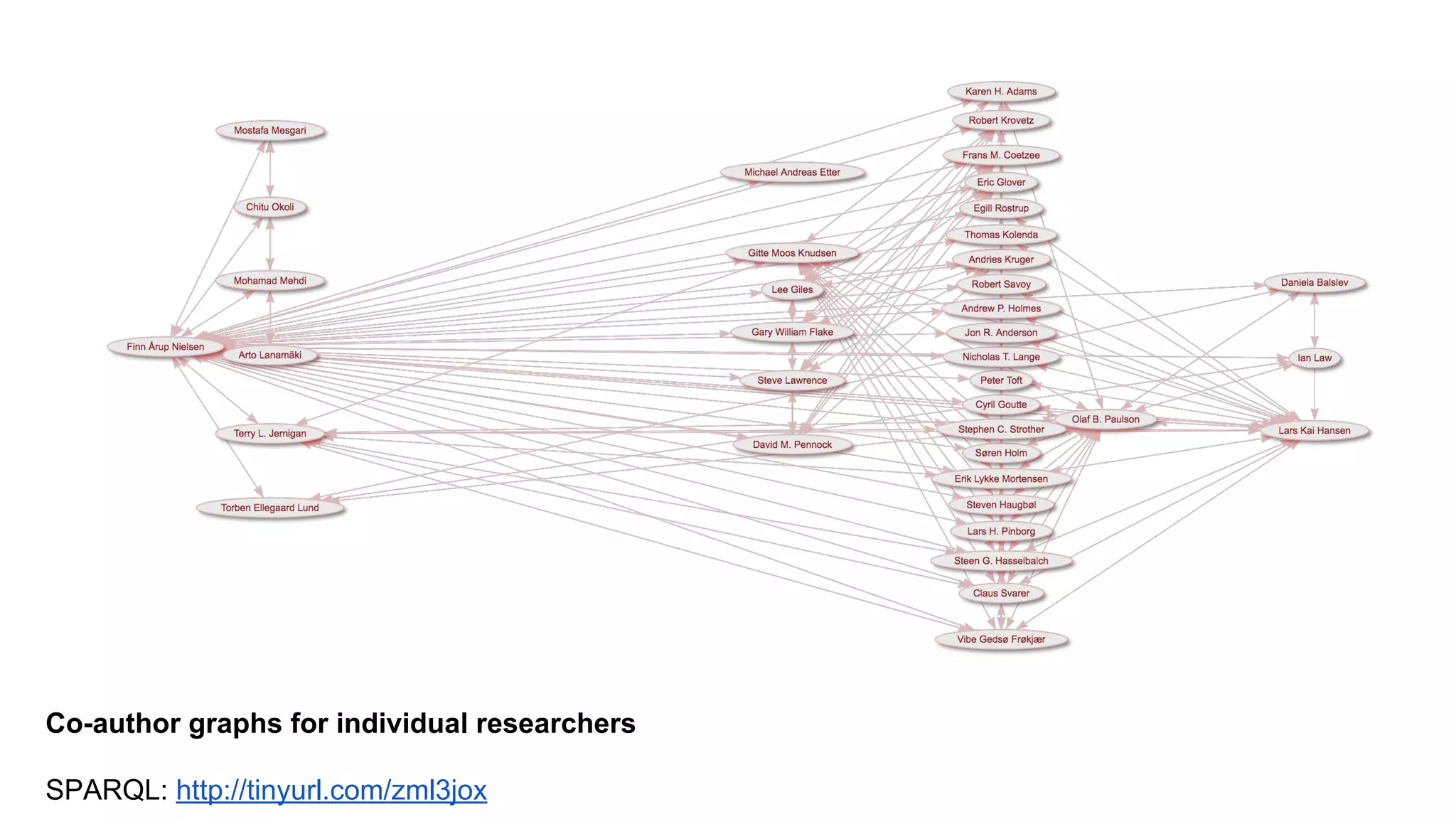
Cocitation network for openfMRI papers, F. Å. Nielsen https://twitter.com/fnielsen/status/752860630932156416
dario@wikimedia.org • @readermeter • @Wikidata • @WikiCite • @WikiResearch](https://image.slidesharecdn.com/vivo2016keynoteverifiablelinkedopenknowledge-160819162835/75/Verifiable-linked-open-knowledge-that-anyone-can-edit-64-2048.jpg)