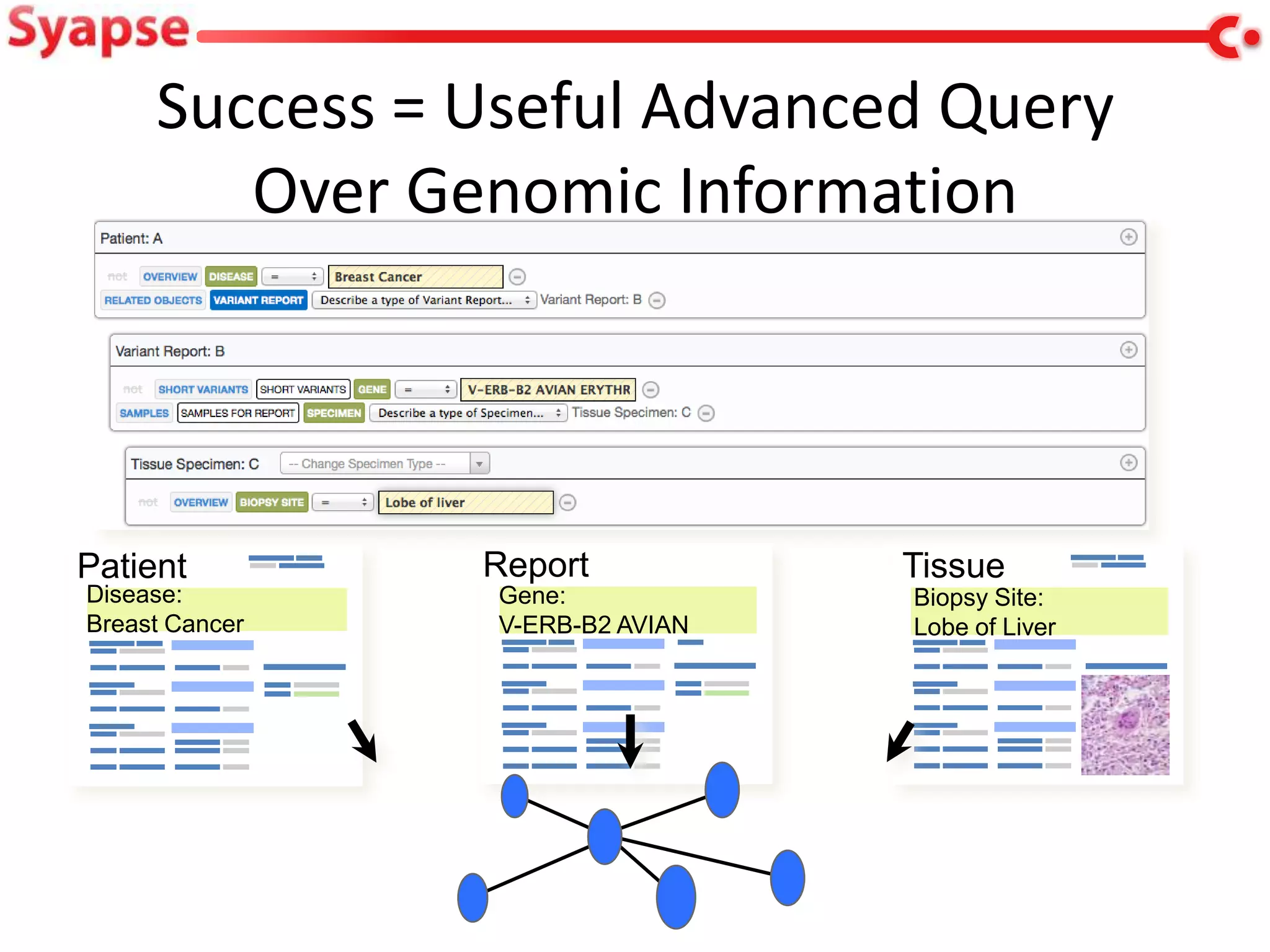
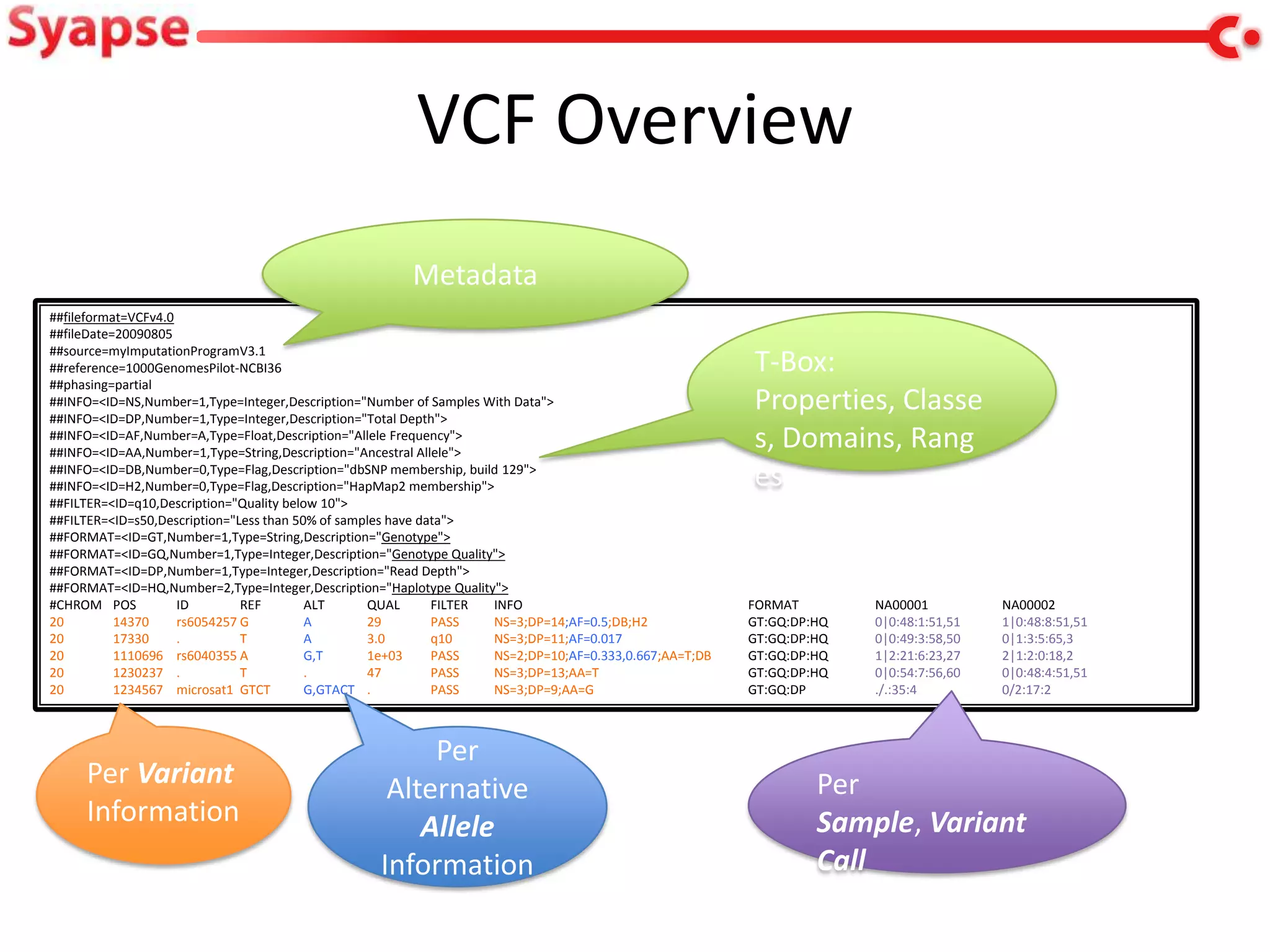
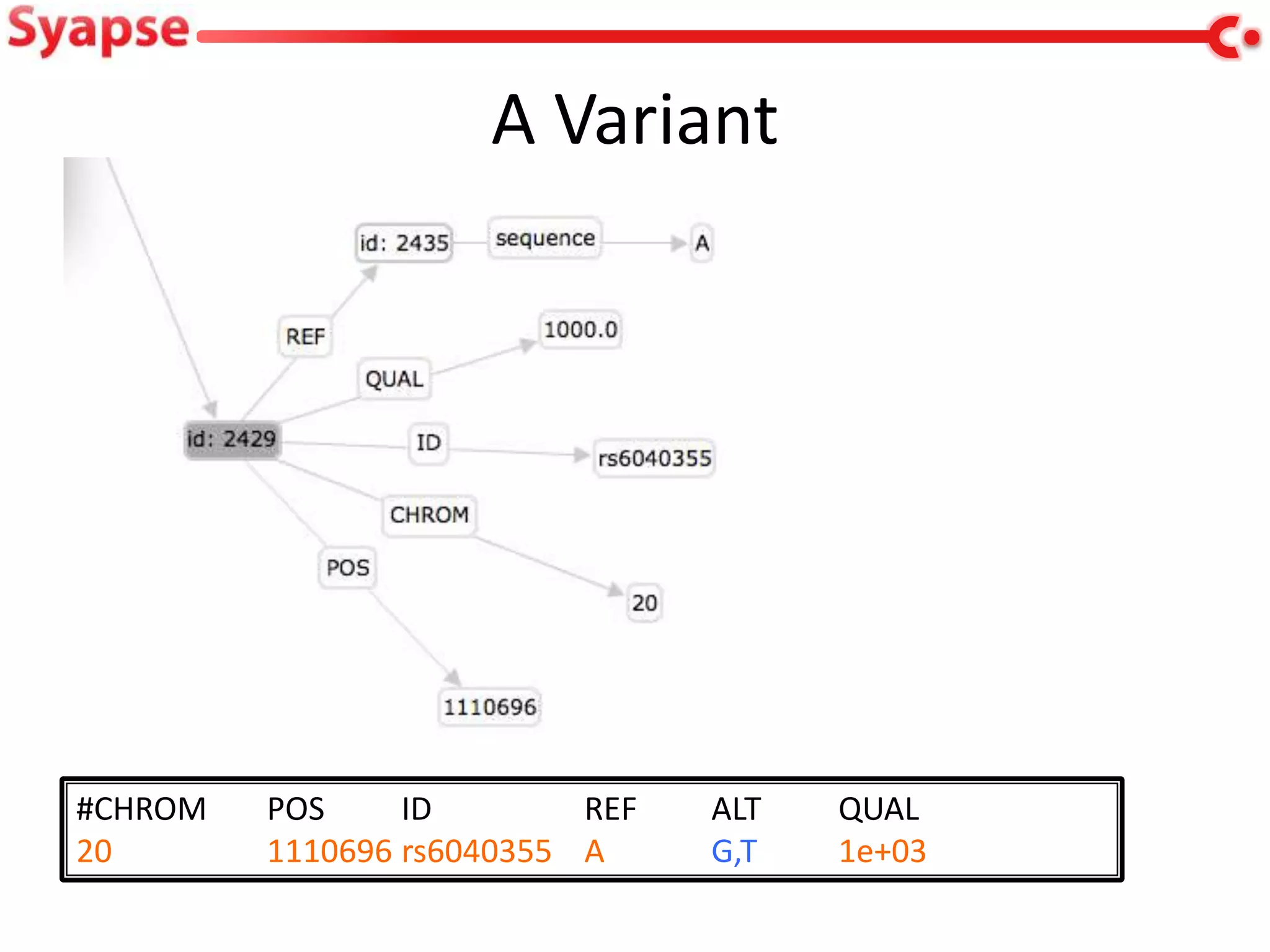
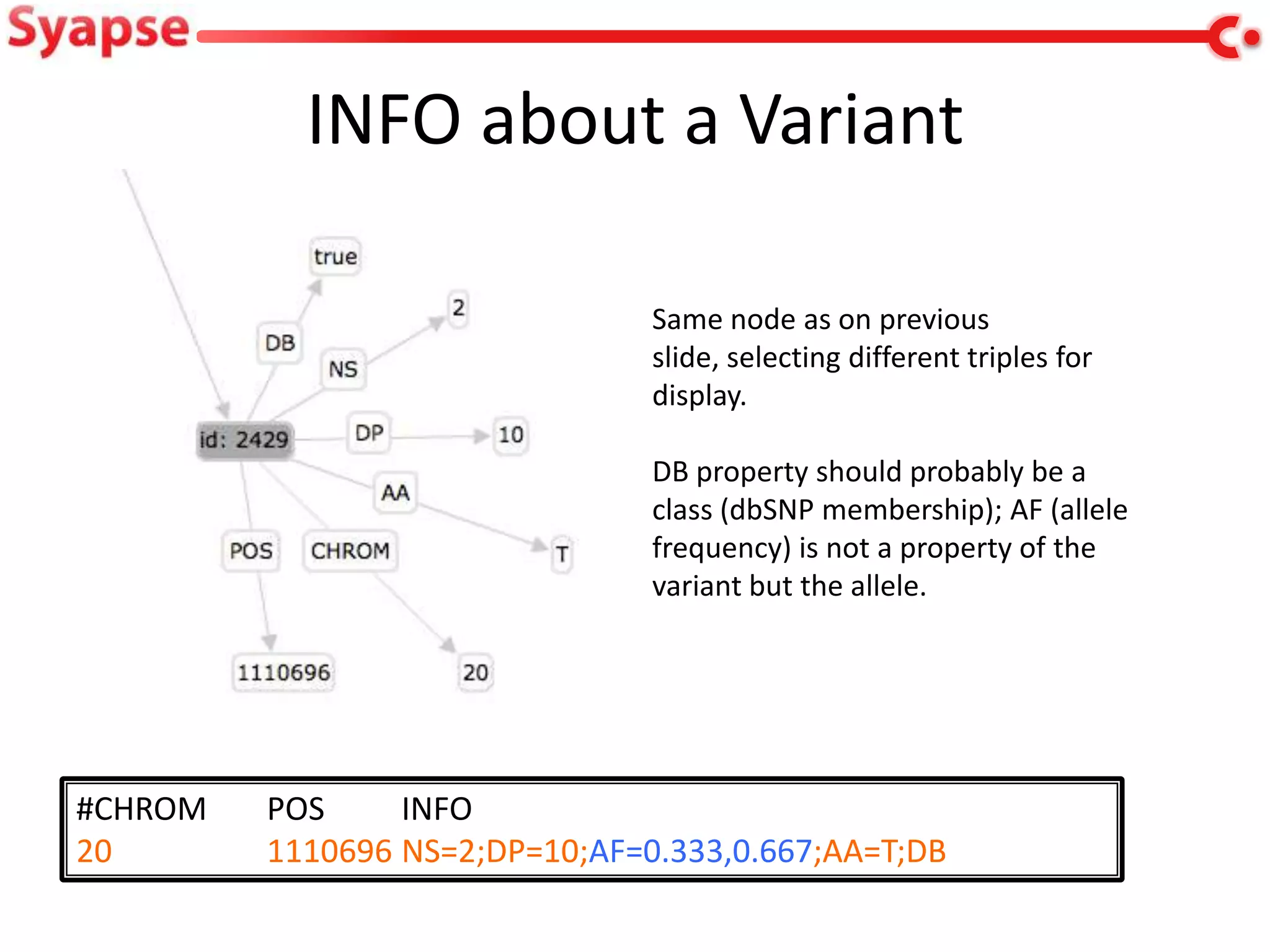
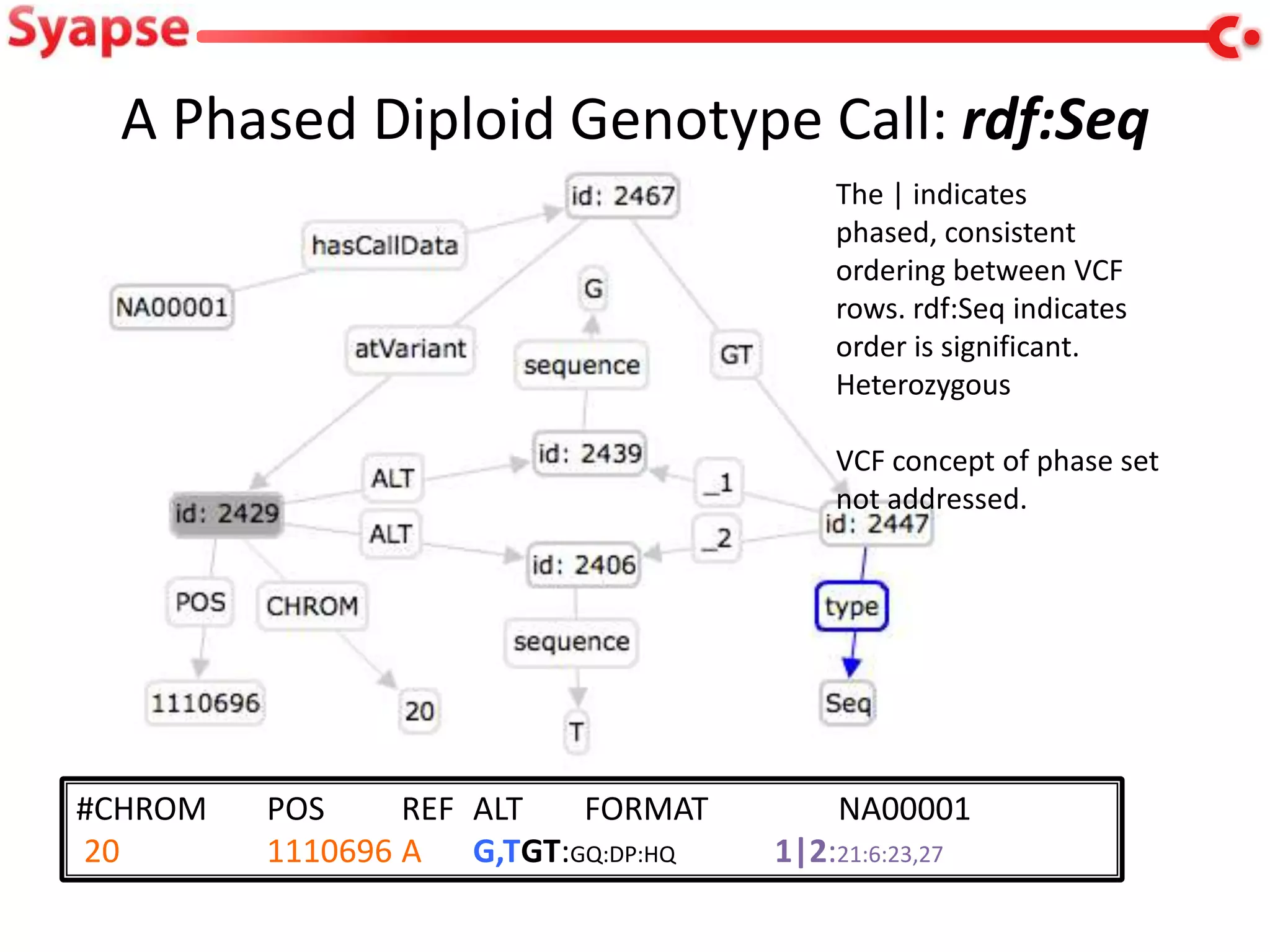
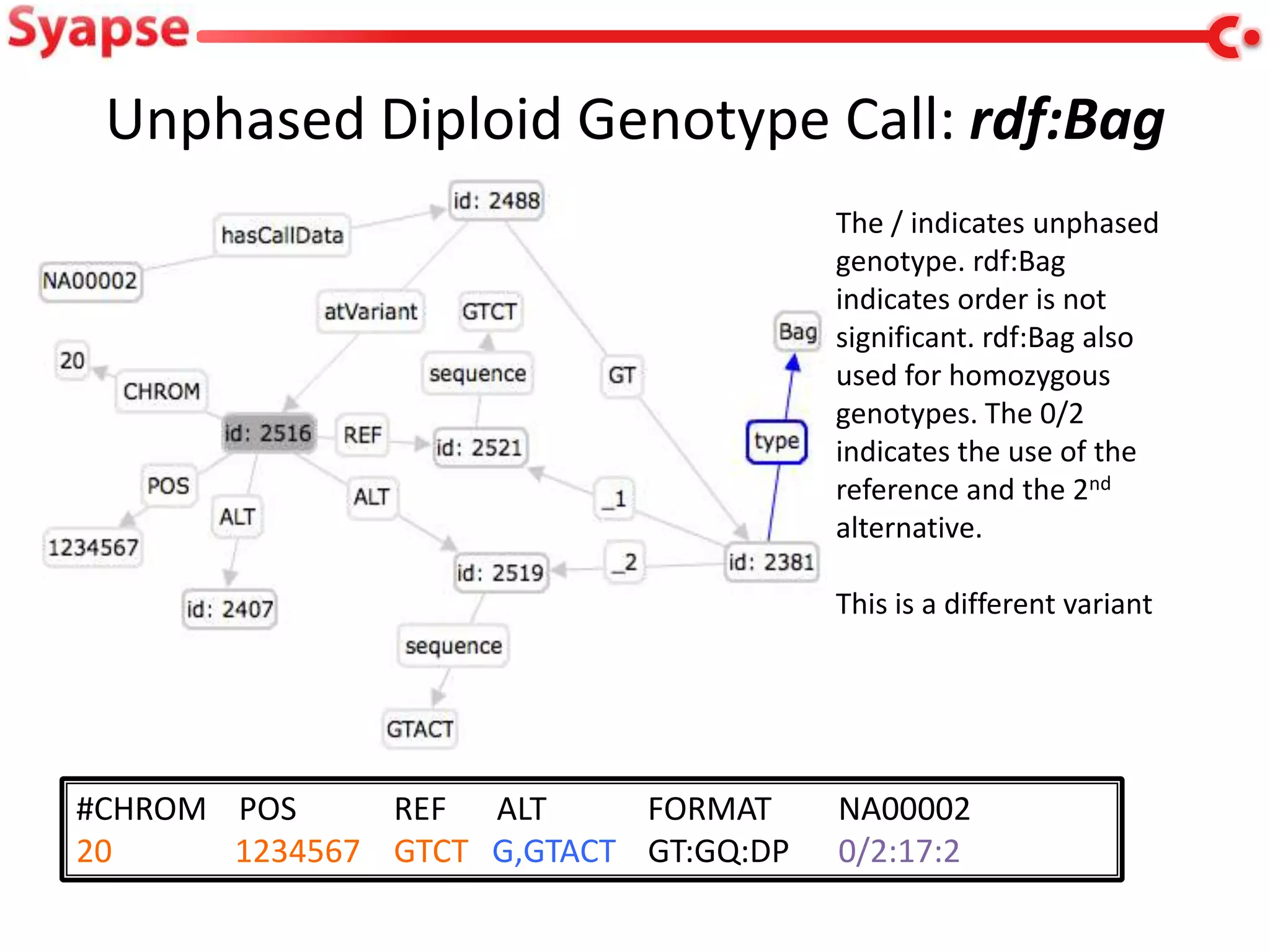
This document explores mapping variant call format (VCF) genomic data into resource description framework (RDF) triples. It presents examples of transforming VCF metadata, variant-level information, allele information, and genotype calls into RDF. It discusses challenges around materializing versus virtual transformations and questions whether the transformed data would be useful for querying to answer interesting biological questions.


![Background
• Syapse Discovery provides an end-to-end
solution for […] laboratories deploying next-
generation sequencing-based diagnostics, […]
configurable semantic data structure enables
users to bring omics data together with
traditional medical information […]
• This work: initial exploration of VCF
import, what useful information is in a VCF
etc.](https://image.slidesharecdn.com/w3c-jjc-lifesci-130401125200-phpapp01/75/VCF-and-RDF-3-2048.jpg)