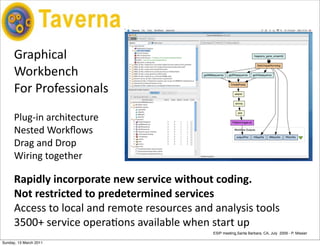
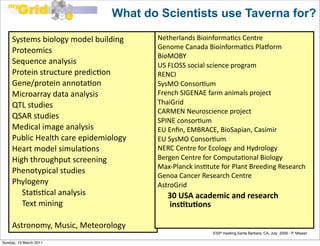
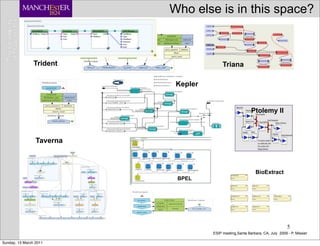
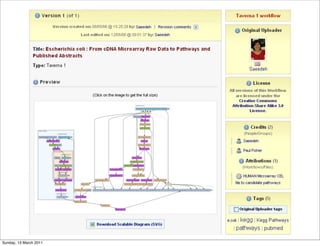
The document discusses the MyGrid project and its components, including Taverna, BioCatalogue, and MyExperiment, focusing on their roles in facilitating collaboration in e-science. It highlights Taverna's capabilities for scientists in various fields, such as systems biology and public health, and outlines the importance of data provenance for ensuring quality and reuse of scientific workflows. The goals of the project include improving workflow technology and addressing challenges related to data preservation and provenance.