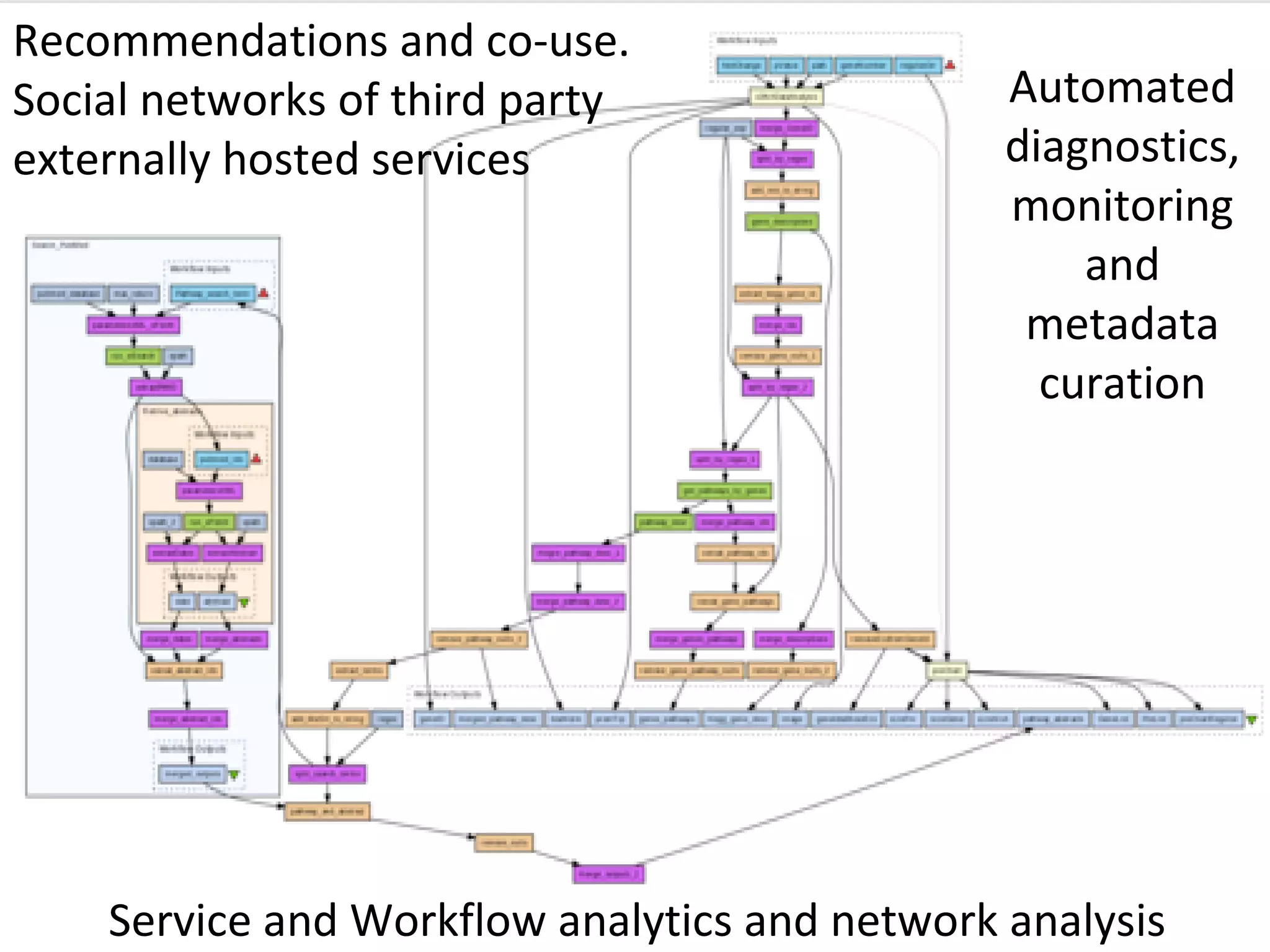
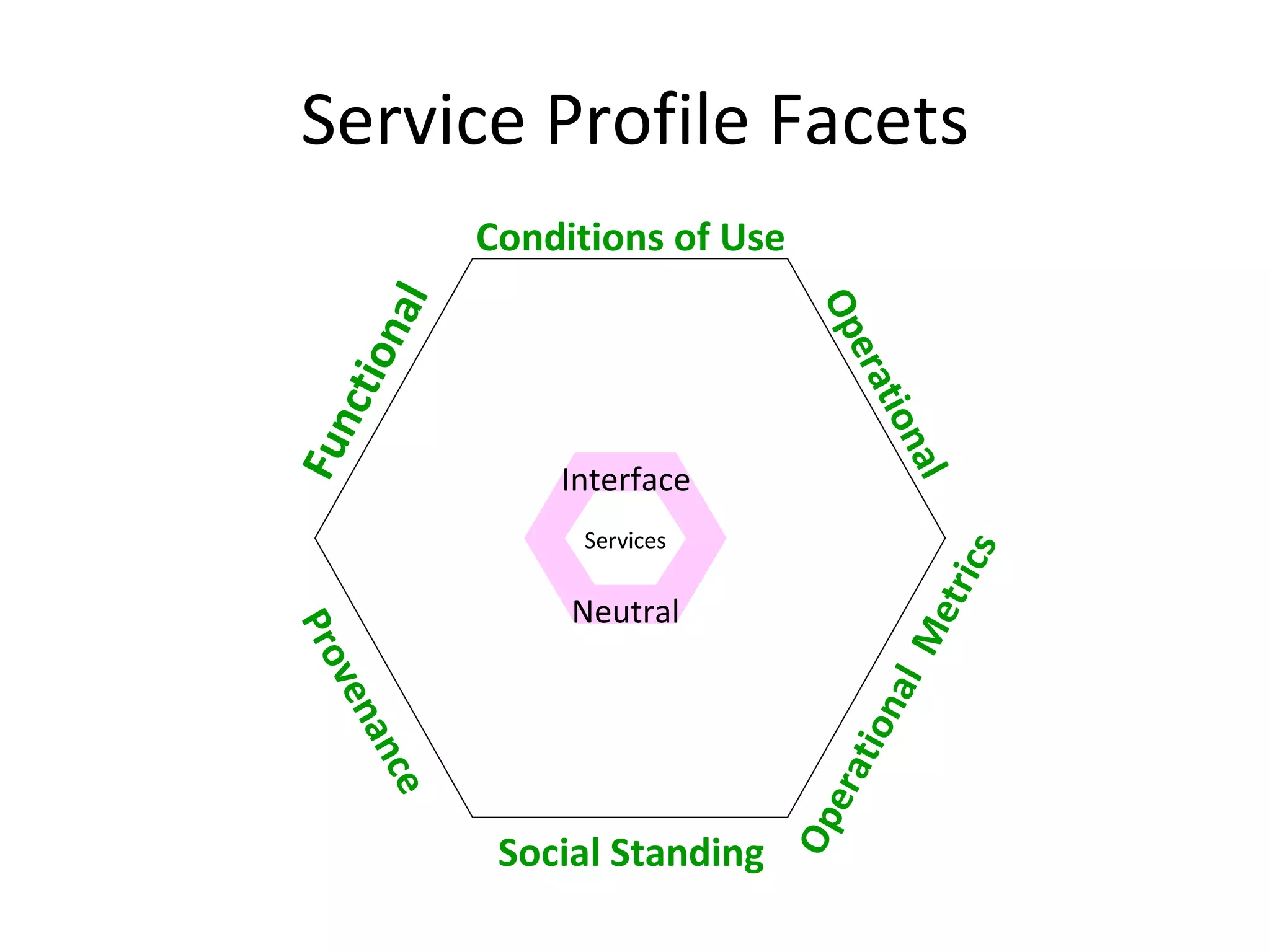
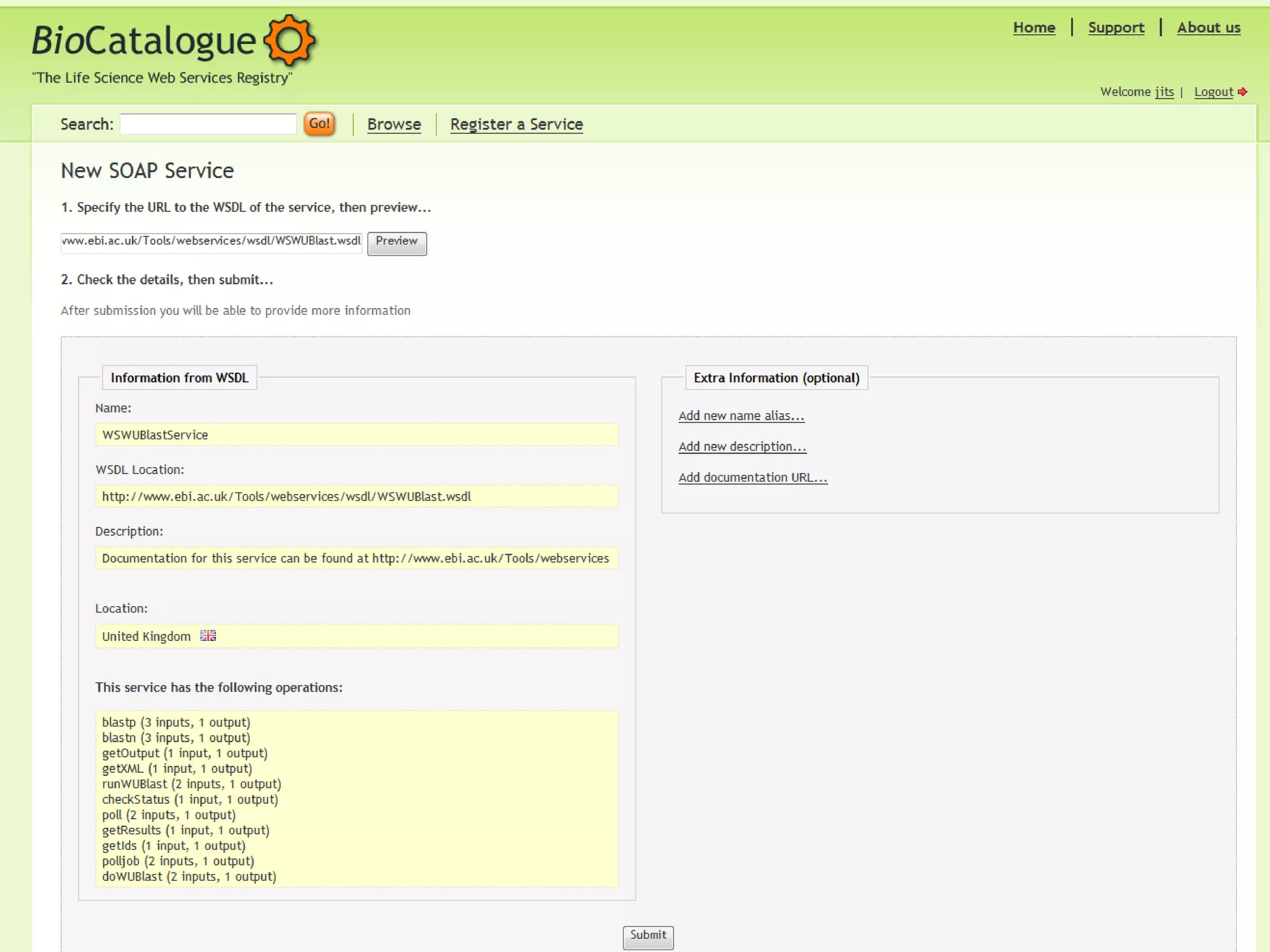
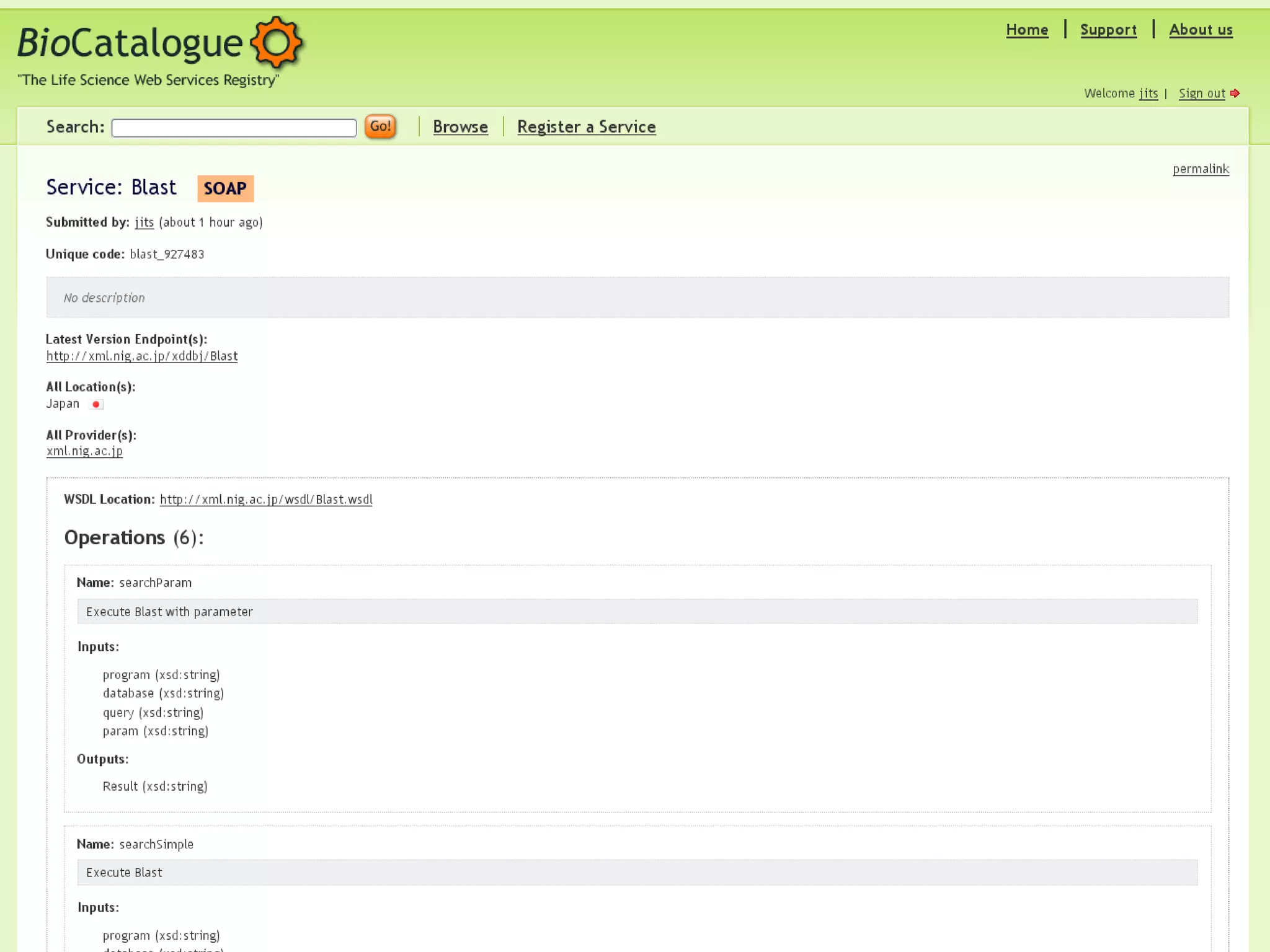
The document outlines the Biocatalogue project aimed at creating a registry for annotated biological web services with a pilot starting on June 1 for six months. It emphasizes sustainable digital curation processes, the importance of metadata, and the need for effective discovery and reuse of web services in bioinformatics. The project involves collaborations with major data centers and aims to enhance user interaction through social curation and improved accessibility to workflows and services.