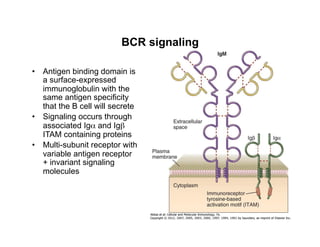
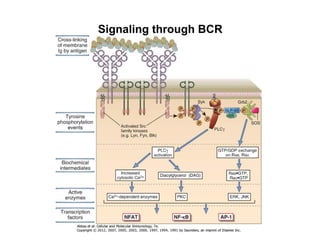
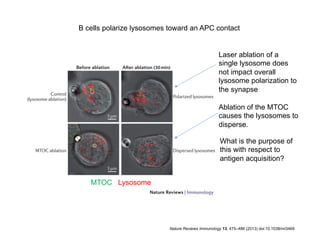
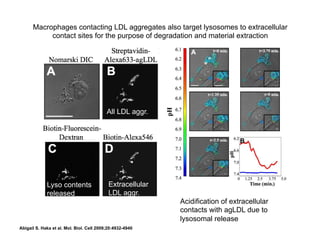
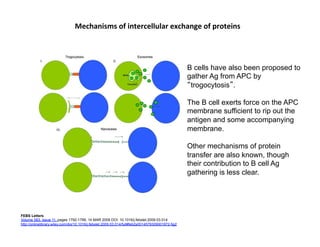
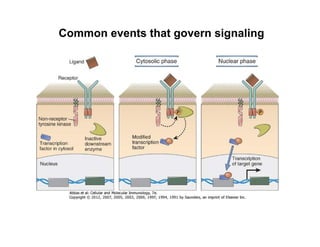
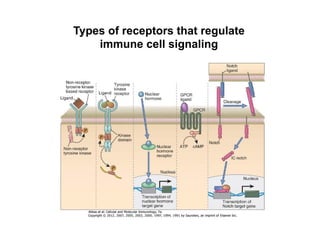
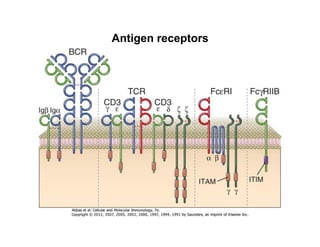
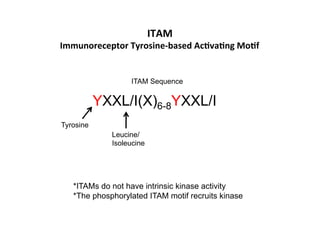
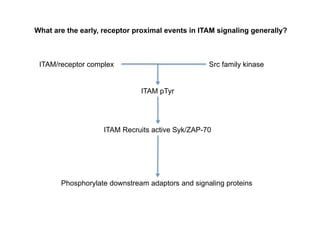
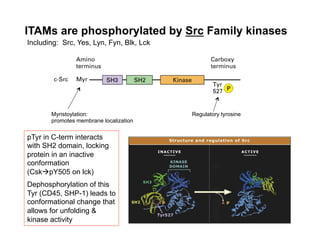
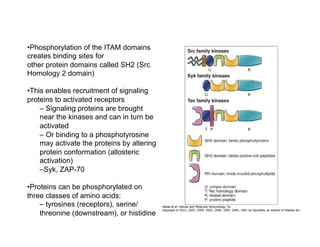
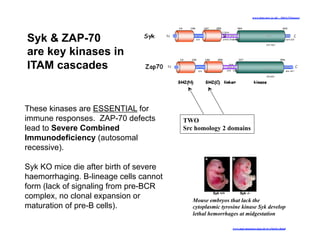
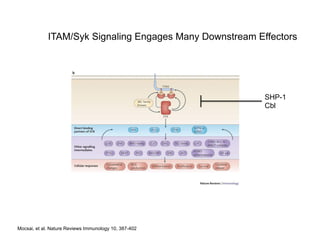
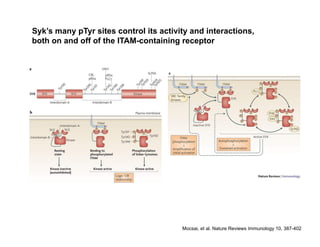
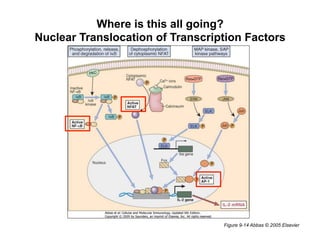
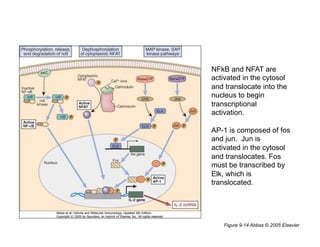
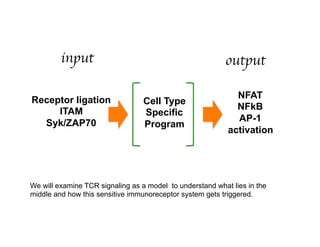
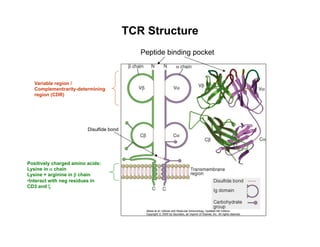
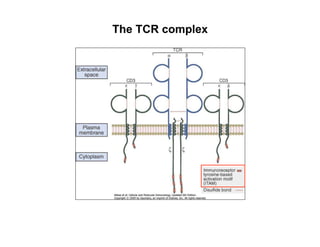
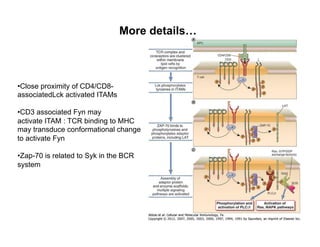
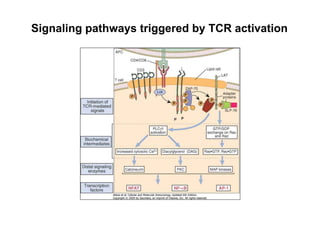
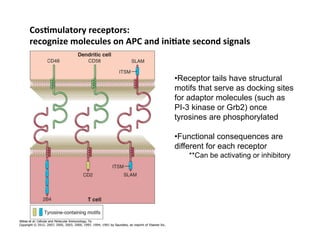
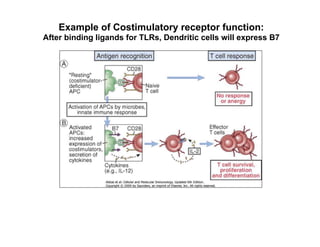
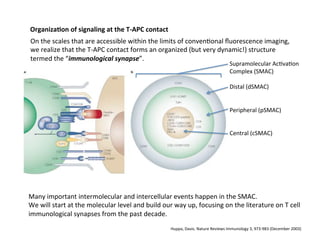
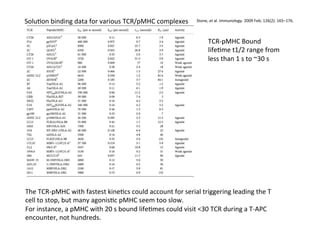
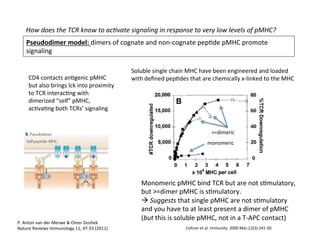
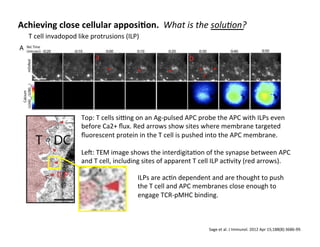
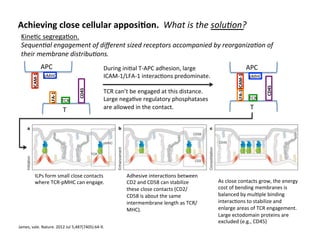
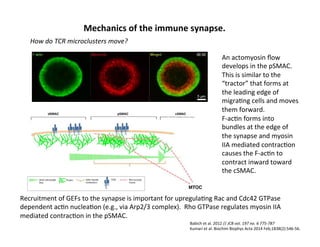
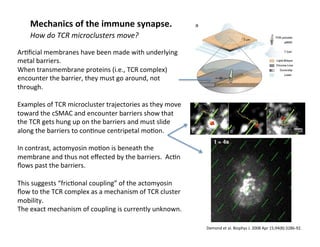
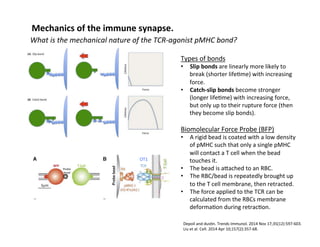
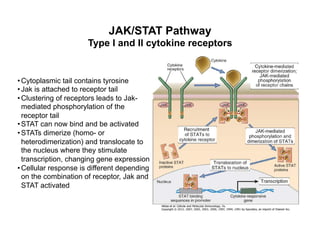
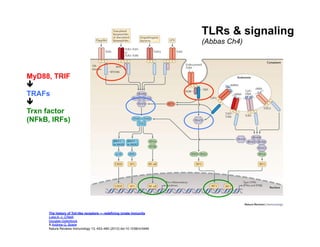
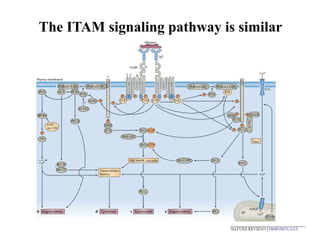
The document provides an overview of antigen receptor signaling and the immunological synapse between T cells and antigen presenting cells (APCs). It discusses how T cell receptor (TCR) signaling is initiated upon antigen binding and transduced through invariant CD3 and ζ proteins containing immunoreceptor tyrosine-based activation motifs (ITAMs). ITAMs are phosphorylated by Src family kinases, recruiting Syk/ZAP-70 kinases and initiating downstream signaling cascades involving transcription factors like NFAT, NF-κB, and AP-1. The interaction between T cells and APCs results in the formation of the immunological synapse, containing central, peripheral, and distal supramolecular activation complexes where receptor clustering
























![Why
does
it
all
ma]er?
Mechanis&c
Underpinning
for
T
cell
Sensi&vity
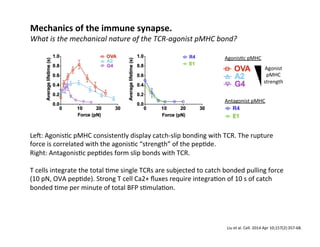
These
studies
help
us
to
understand
the
physical
parameters
that
define
whether
a
pMHC
will
be
s&mulatory
or
not.
They
also
help
us
to
understand
the
exquisite
sensi&vity
of
T
cells
for
low
densi&es
of
agonist
pMHC.
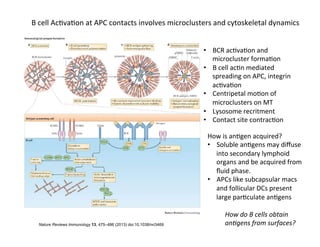
Sensi&vity
is
important
because
professional
APCs
like
DCs
might
not
express
large
densi&es
of
a
par&cular
cognate
pMHC
for
a
T
cell.
Also,
a
scanning
T
cell/APC
interac&on
in
the
lymph
node
is
rather
short
lived
(~minutes),
so
the
T
cell
needs
to
be
able
to
signal
and
stop
migra&ng
if
even
a
small
amount
of
cognate
pMHC
is
found.
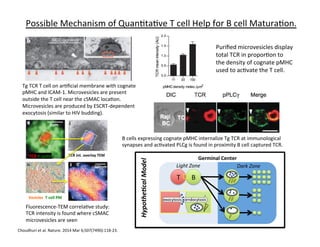
Possible
Mechanism
of
Quan&ta&ve
T
cell
Help
for
B
cell
Matura&on.
Review
re:
role
of
IS
in
T
cell
help
for
B
cells-‐-‐Dus&n.
Mol
Cell.
2014
Apr
24;54(2):255-‐62.
B
T
Ag
Y
Y
AnAgen
gathering
Efficiency
α
BCR
affinity
AnAgen
presentaAon
#
pMHC
α
BCR
affinity
TCR
signals,
acAvaAon
#
CD40L
α
#
pMHC
seen
T
B
T
cell
help
#
CD40L
α
B
cell
prolif
A
2014
ar&cle
suggested
that
exocytosis
of
TCR
in
the
cSMAC
may
contribute
to
long-‐term
regula&on
of
B
cell
responses
to
T
cell
help
las&ng
beyond
the
&me
frame
of
direct
T-‐B
interac&ons.](https://image.slidesharecdn.com/biom514cellsignalingneumann2017fullpghandout-170818220924/85/Immunoreceptor-Signaling-Lecture-Slides-72-320.jpg)





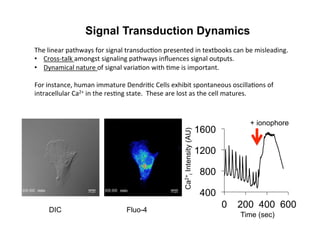
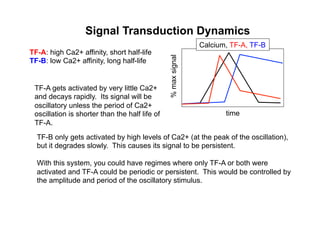
![Signal Transduction Dynamics
Dolmetsch, et al. Nature 392, 933-936(30 April 1998)
• T cells also exhibit Ca2+ oscillations and spikes during activation
• Ca2+ clamp method can reproduce arbitrary oscillation amplitudes
and periods (left)
At low levels of Ca2+, oscillatory [Ca2+]i
increases the number of cells expressing
an NFAT reporter.](https://image.slidesharecdn.com/biom514cellsignalingneumann2017fullpghandout-170818220924/85/Immunoreceptor-Signaling-Lecture-Slides-91-320.jpg)