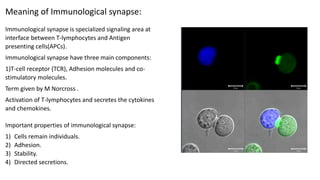
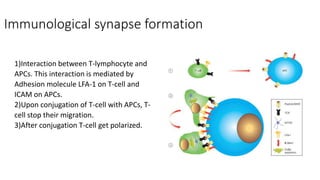
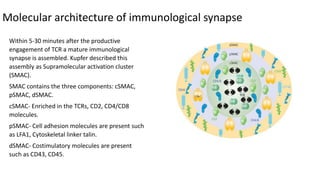
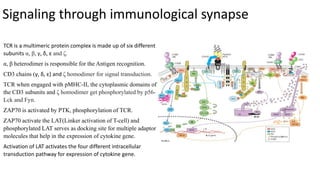
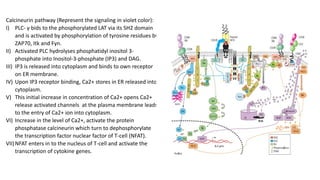
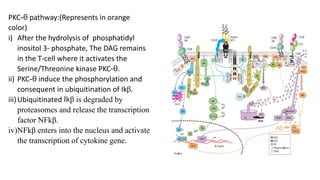
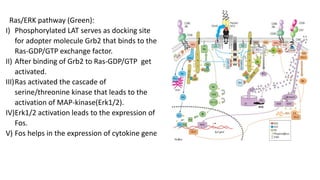
The immunological synapse is a specialized signaling structure formed at the interface between T lymphocytes and antigen presenting cells. It consists of three main components: T cell receptors, adhesion molecules, and co-stimulatory molecules. Formation of the mature immunological synapse involves molecular redistribution through diffusion and cytoskeletal movement over 5-30 minutes, resulting in a structure with central, peripheral, and distal supramolecular activation clusters that facilitate T cell signaling and activation. This signaling activates transcription factors that induce cytokine gene expression and T cell effector functions.