The document discusses genome-wide association studies (GWAS) and their significance in identifying genetic associations with diseases and traits. It highlights the consistency of associations between certain gene polymorphisms and various conditions, alongside practical considerations for study design, such as sample size and population stratification. Additionally, it mentions technological advancements in genotyping and the implications of p-values in statistical analyses of genetic data.













































![Sources / References / Reading
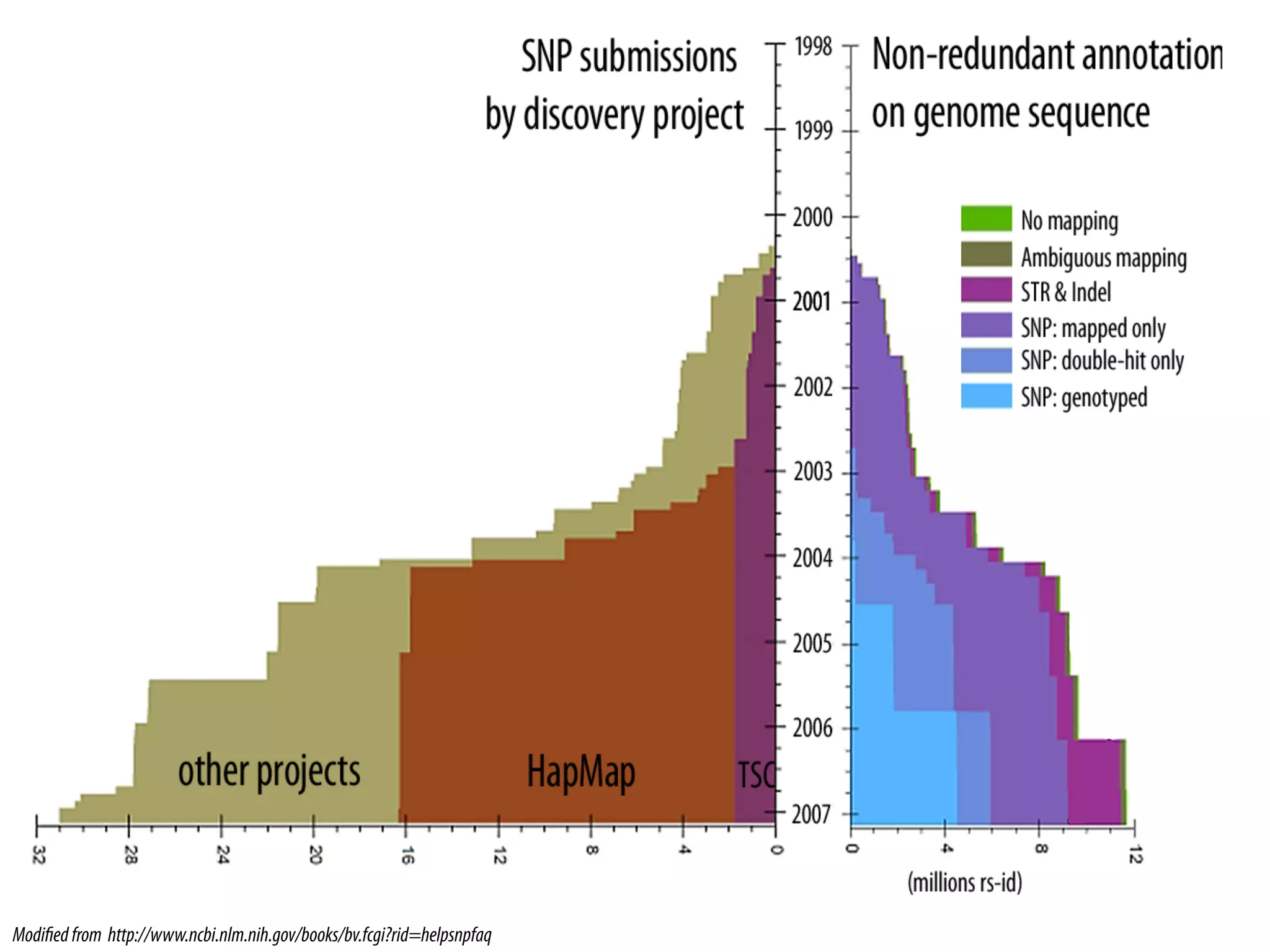
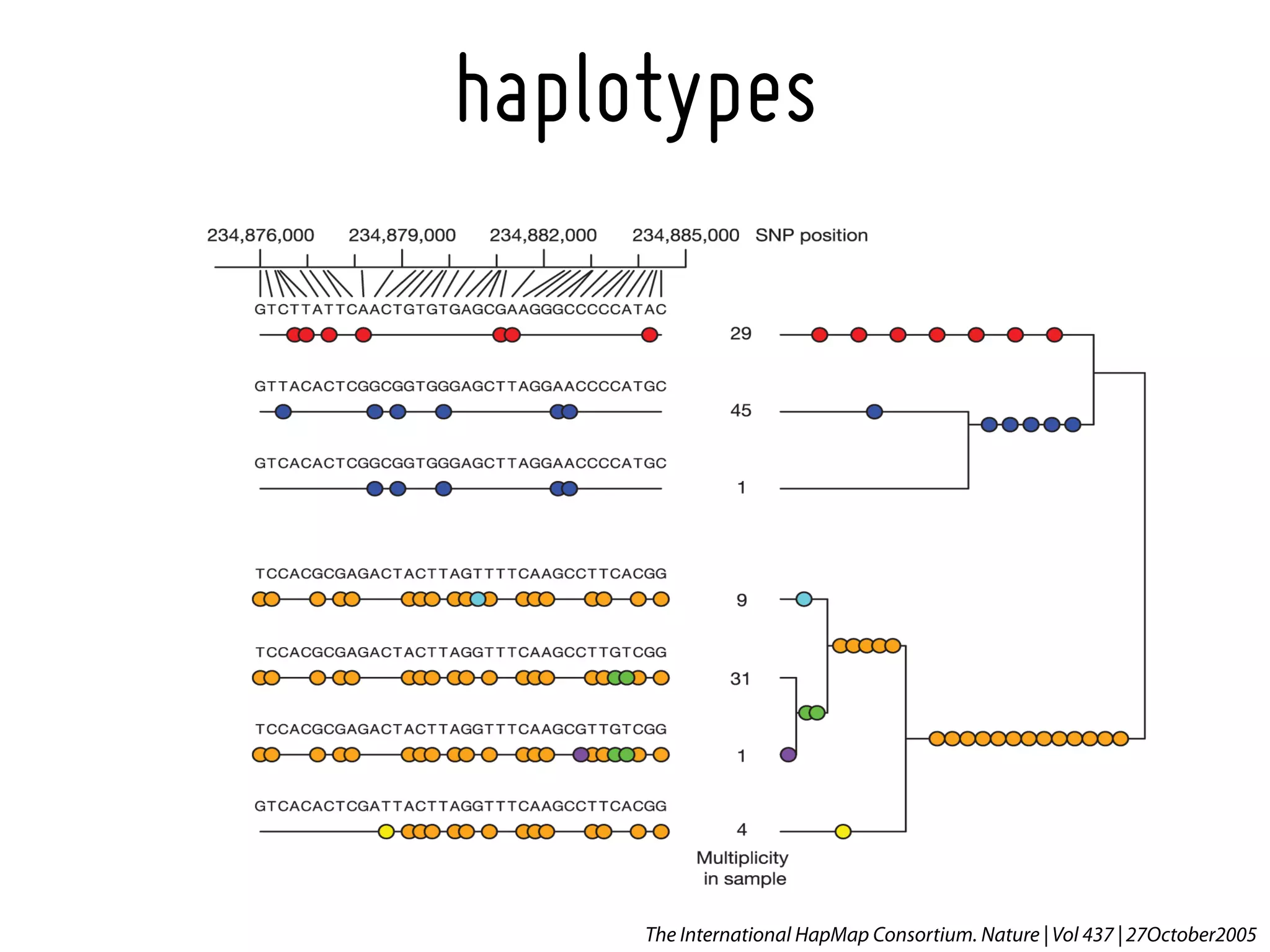
1. The International HapMap Consortium.* A haplotype map of the human genome. Nature, 2005. 437(7063): p. 1299-320.[16255080].
2. The Type 1 Diabetes Genetics Consortium.* Genome-wide association study and meta-analysis find that over 40 loci affect risk of type 1 diabetes. Nature Genetics, 2009
May 10 [19430480]
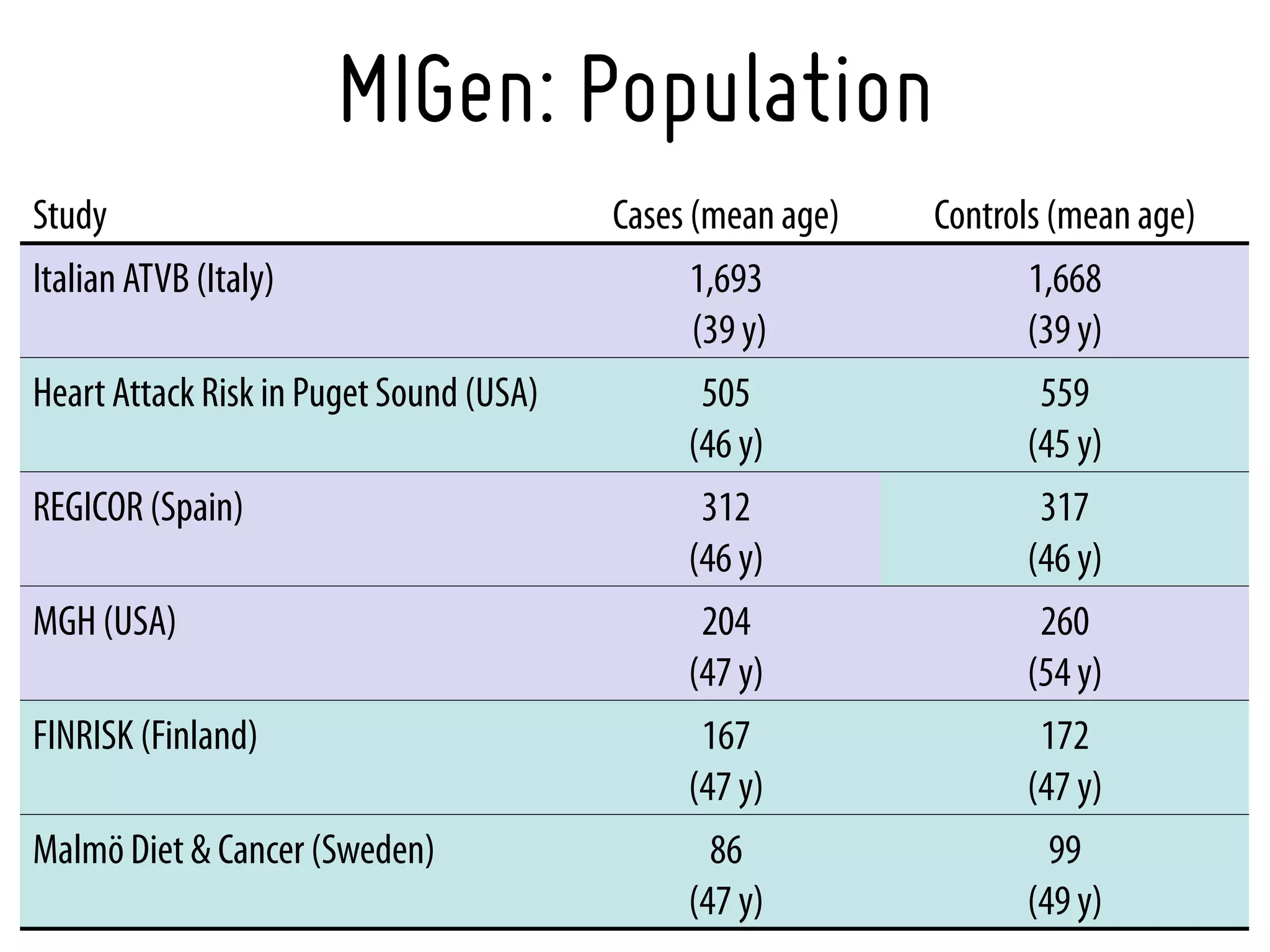
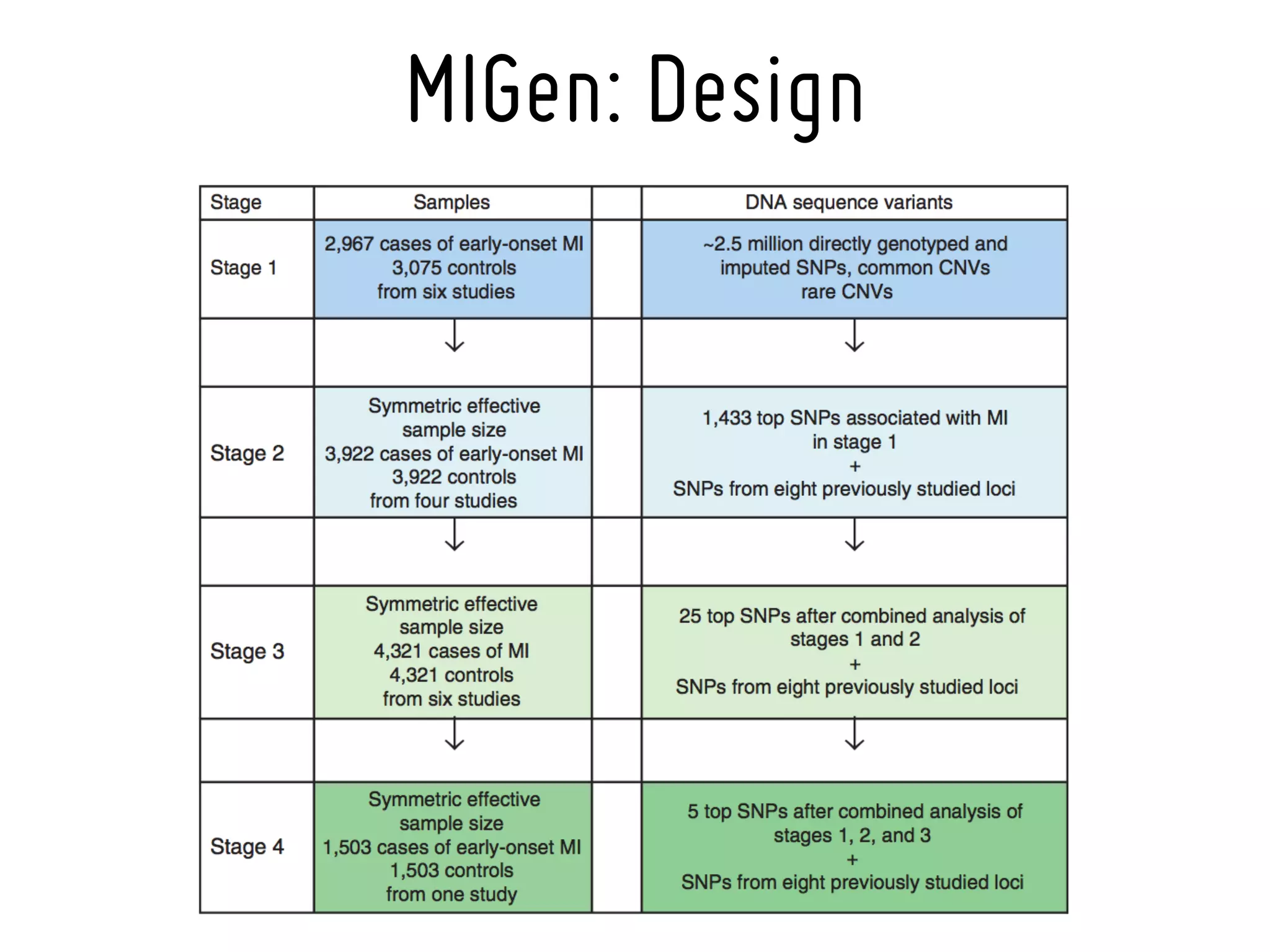
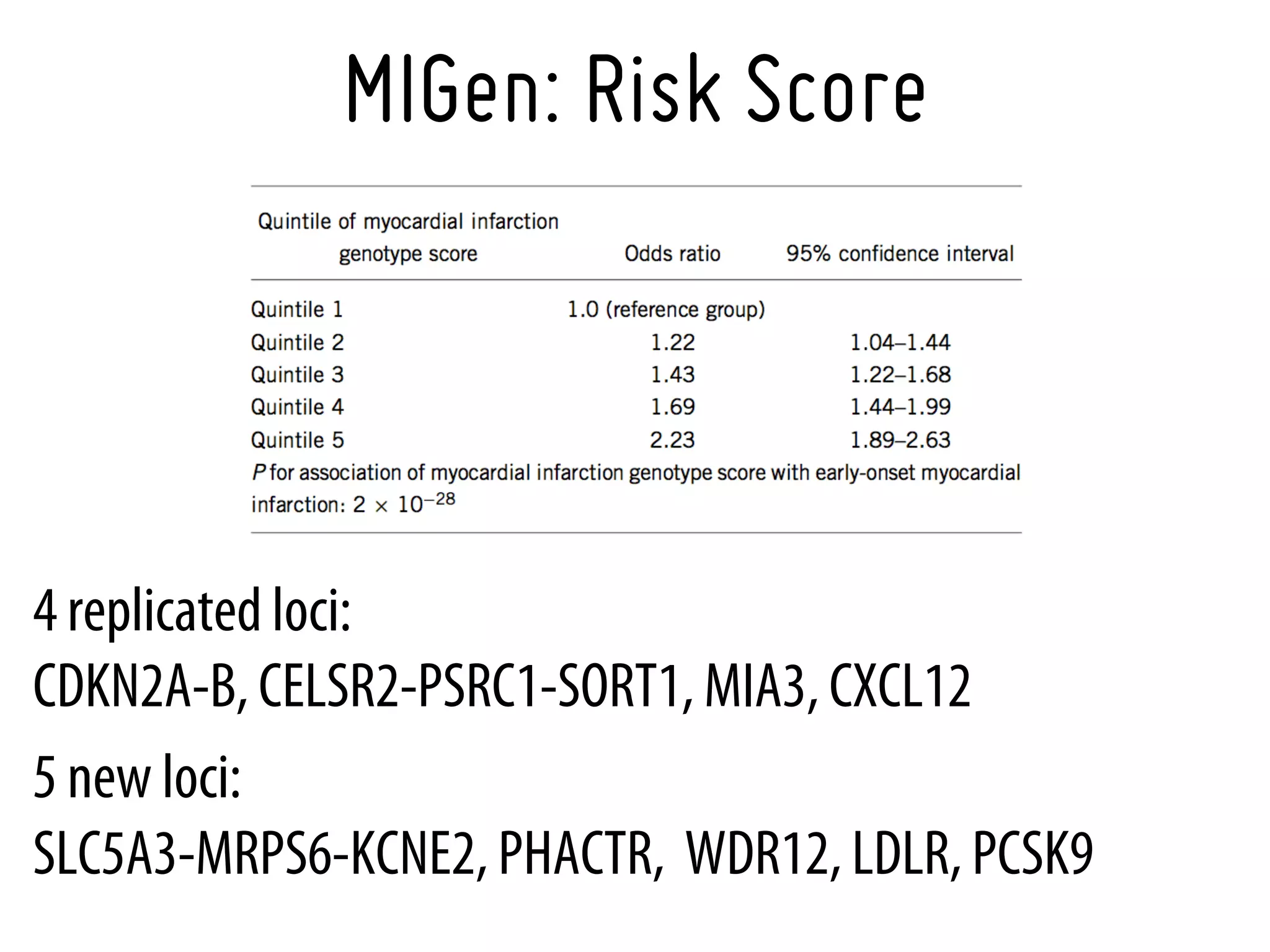
3. Myocardial Infarction Genetics Consortium.* Genome-wide association of early-onset myocardial infarction with single nucleotide polymorphisms and copy number
variants. Nat Genet. 2009 Mar;41(3):334-41 [19198609]
4. The Wellcome Trust Case Control Constortium.* Genome-wide association study of 14,000 cases of seven common diseases and 3,000 shared controls. Nature, 2007. 447
(7145): p. 661-78.[17554300].
5. Balding, D.J., A tutorial on statistical methods for population association studies. Nat Rev Genet, 2006. 7(10): p. 781-91.[16983374].
6. Christensen, K. and J.C. Murray, What genome-wide association studies can do for medicine. N Engl J Med, 2007. 356(11): p. 1094-7.[17360987].
7. Frazer, K.A., et al., A second generation human haplotype map of over 3.1 million SNPs. Nature, 2007. 449(7164): p. 851-61.[17943122].
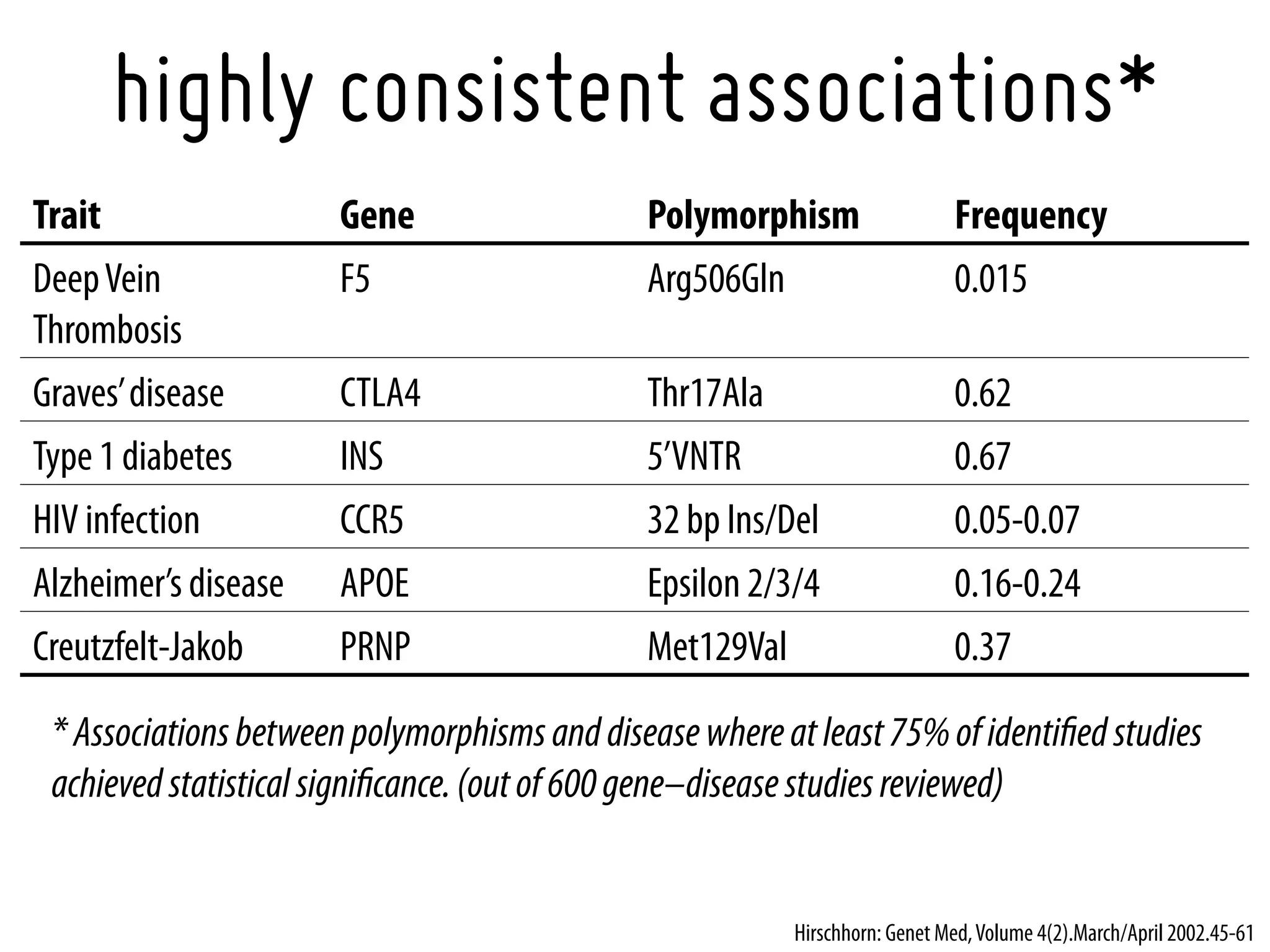
8. Hirschhorn, J.N., et al., A comprehensive review of genetic association studies. Genet Med, 2002. 4(2): p. 45-61.[11882781].
9. Hoover, R. The evolution of epidemiologic research: from cottage industry to "big" science. Epidemiology. 2007 Jan;18(1):13-7. [17179754]
10. Hunter, D.J. and P. Kraft, Drinking from the fire hose--statistical issues in genomewide association studies. N Engl J Med, 2007. 357(5): p. 436-9.[17634446].
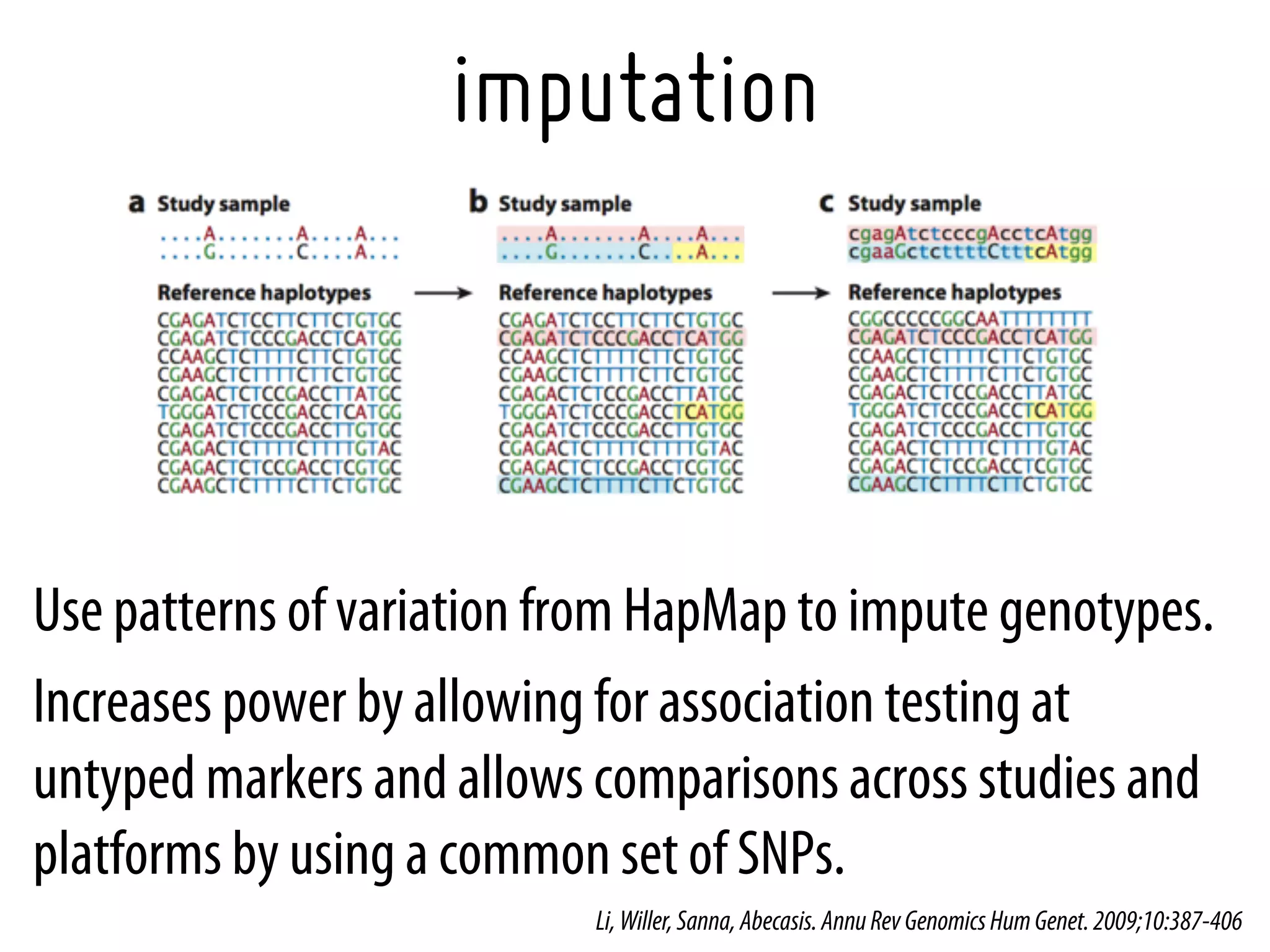
11. Li Y, Willer C, Sanna S, Abecasis G., Genotype imputation. Annu Rev Genomics Hum Genet. 2009;10:387-406. [19715440]
12. Johnson AD and O’Donnell CJ: Open access database of GWA results, BMC Medical Genetics 2009: 10:6
13. Manolio, T.A., et al., Genetics of ultrasonographic carotid atherosclerosis. Arterioscler Thromb Vasc Biol, 2004. 24(9): p. 1567-77.[15256397].
14. Manolio, T.A., L.D. Brooks, and F.S. Collins, A HapMap harvest of insights into the genetics of common disease. J Clin Invest, 2008. 118(5): p. 1590-605.[18451988].
15. Manolio TA, Collins FS, Cox NJ, et al. Finding the missing heritability of complex diseases. Nature. 2009 Oct 8;461(7265):747-53. [19812666]
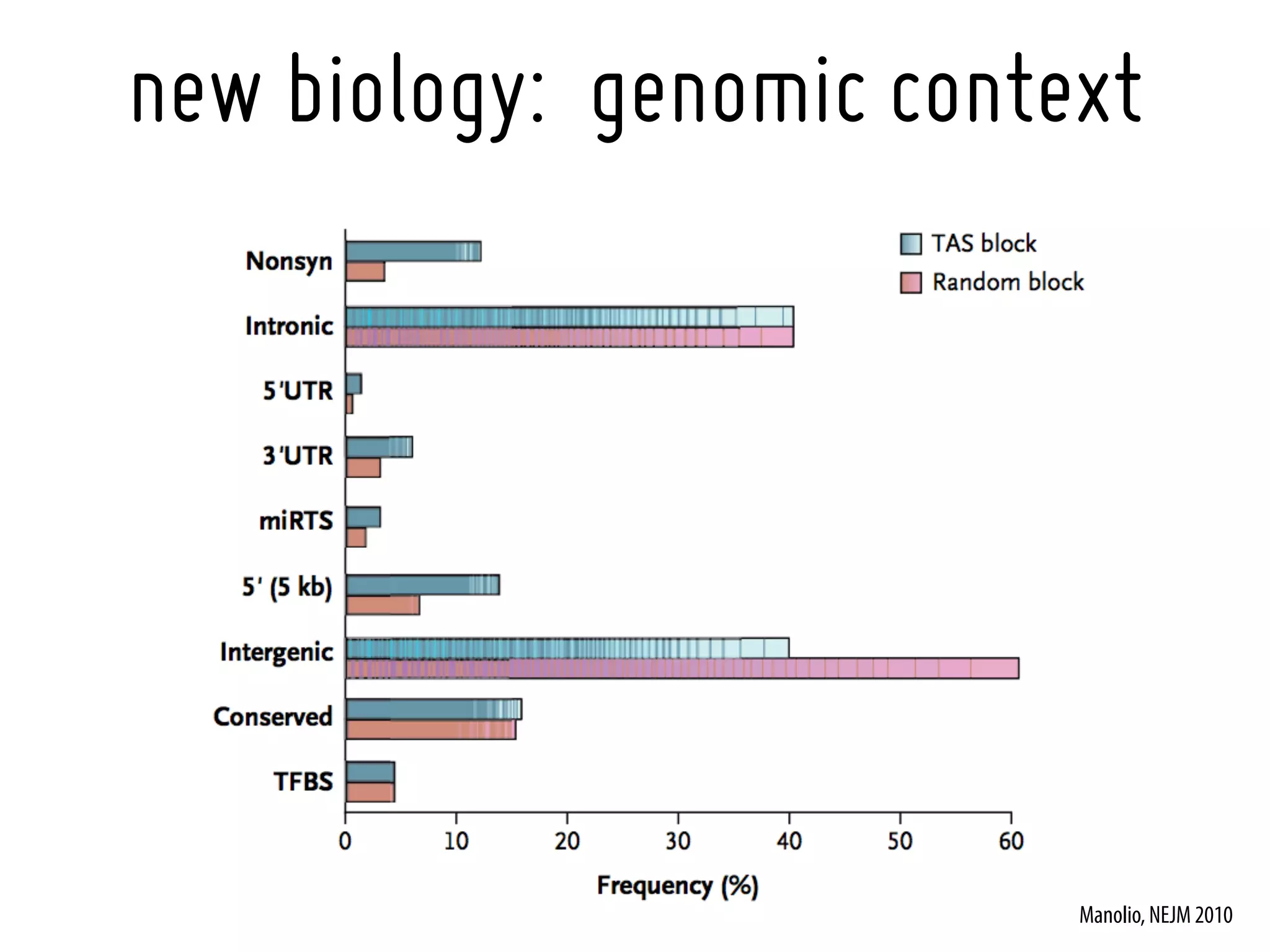
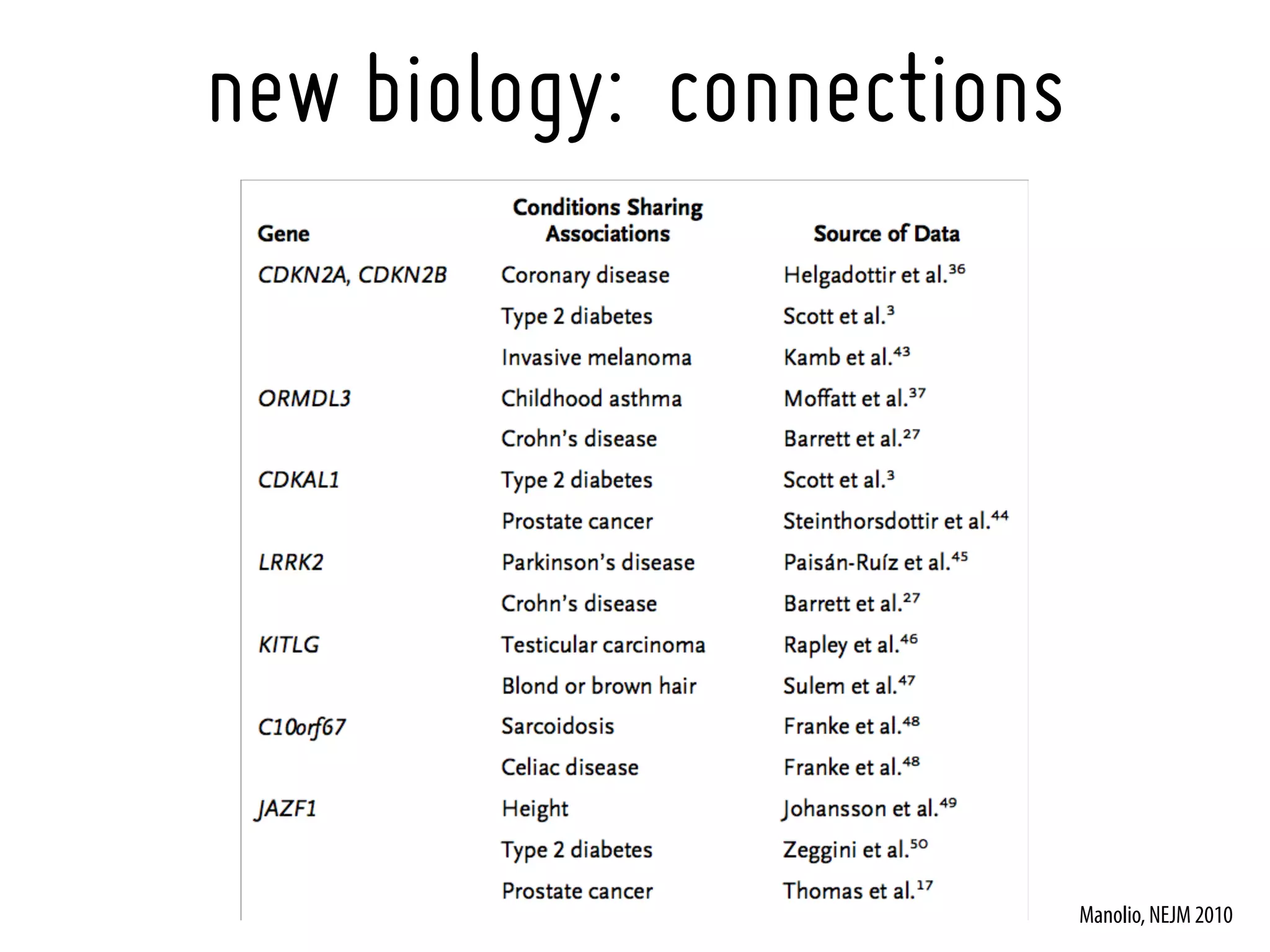
16. Manolio, TA. Genomewide association studies and assessment of the risk of disease. N Engl J Med. 2010 Jul 8;363(2):166-76. [20647212]
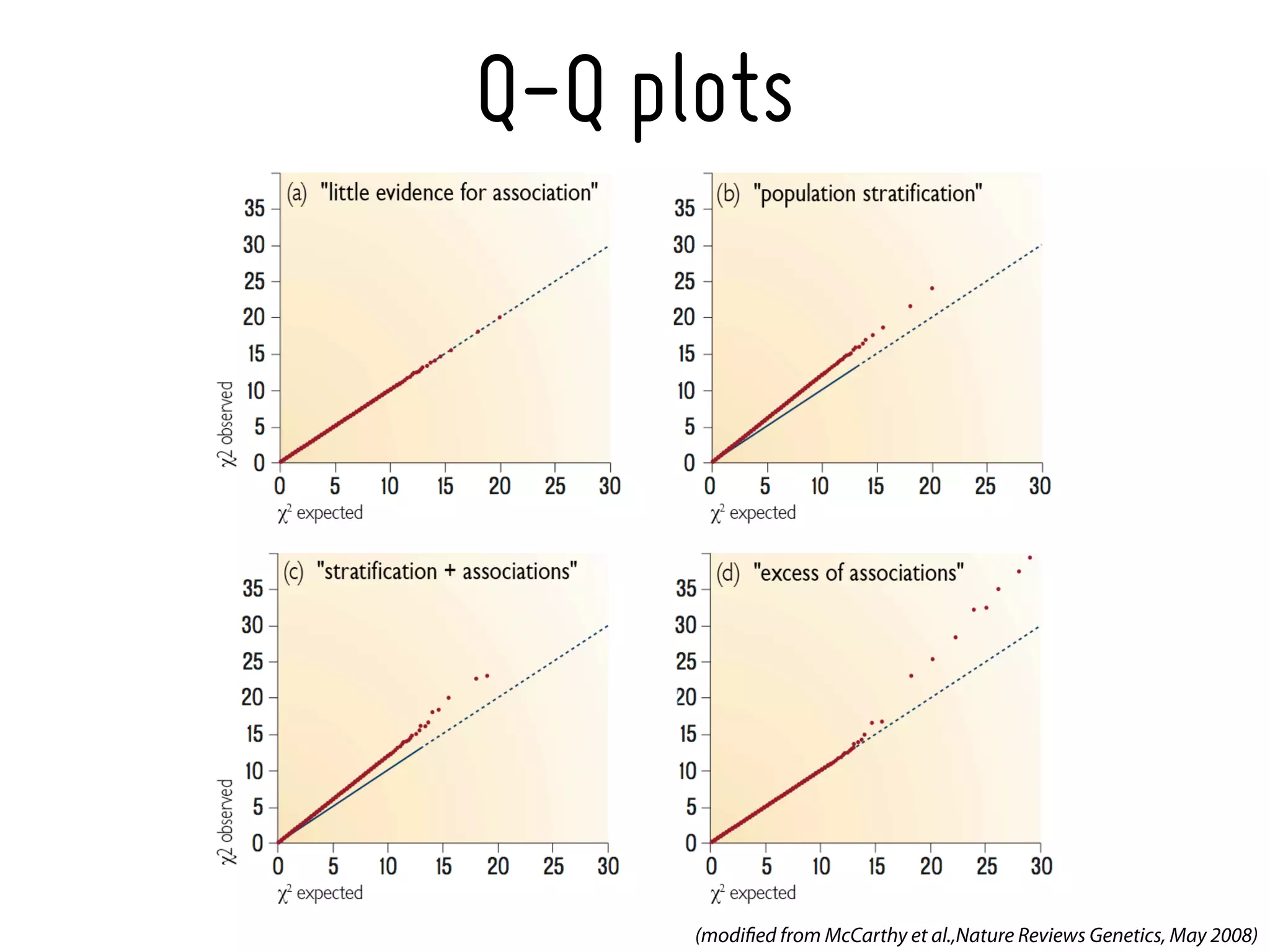
17. McCarthy, M.I., et al., Genome-wide association studies for complex traits: consensus, uncertainty and challenges. Nat Rev Genet, 2008. 9(5): p. 356-69.[18398418].
18. Pearson, T.A. and T.A. Manolio, How to interpret a genome-wide association study. JAMA, 2008. 299(11): p. 1335-44.[18349094].
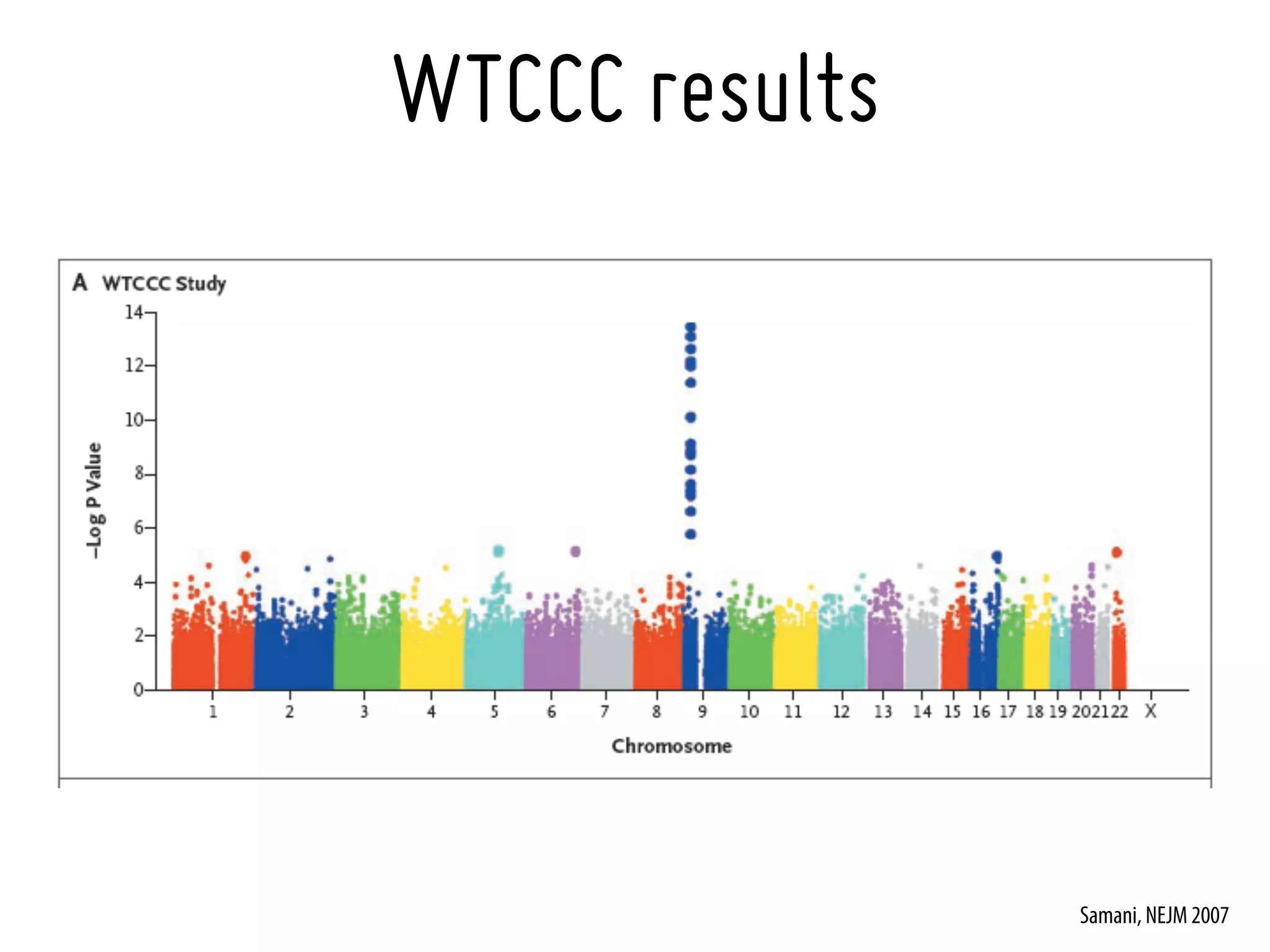
19. Samani NJ, Erdmann J, Hall AS, et al. Genomewide association analysis of coronary artery disease. N Engl J Med. 2007 Aug 2;357(5):443-53. [17634449]
20. Teslovich TM, Musunuru K, Smith AV, et al. Biological, clinical and population relevance of 95 loci for blood lipids. Nature. 2010 Aug 5;466(7307):707-13. [20686565]
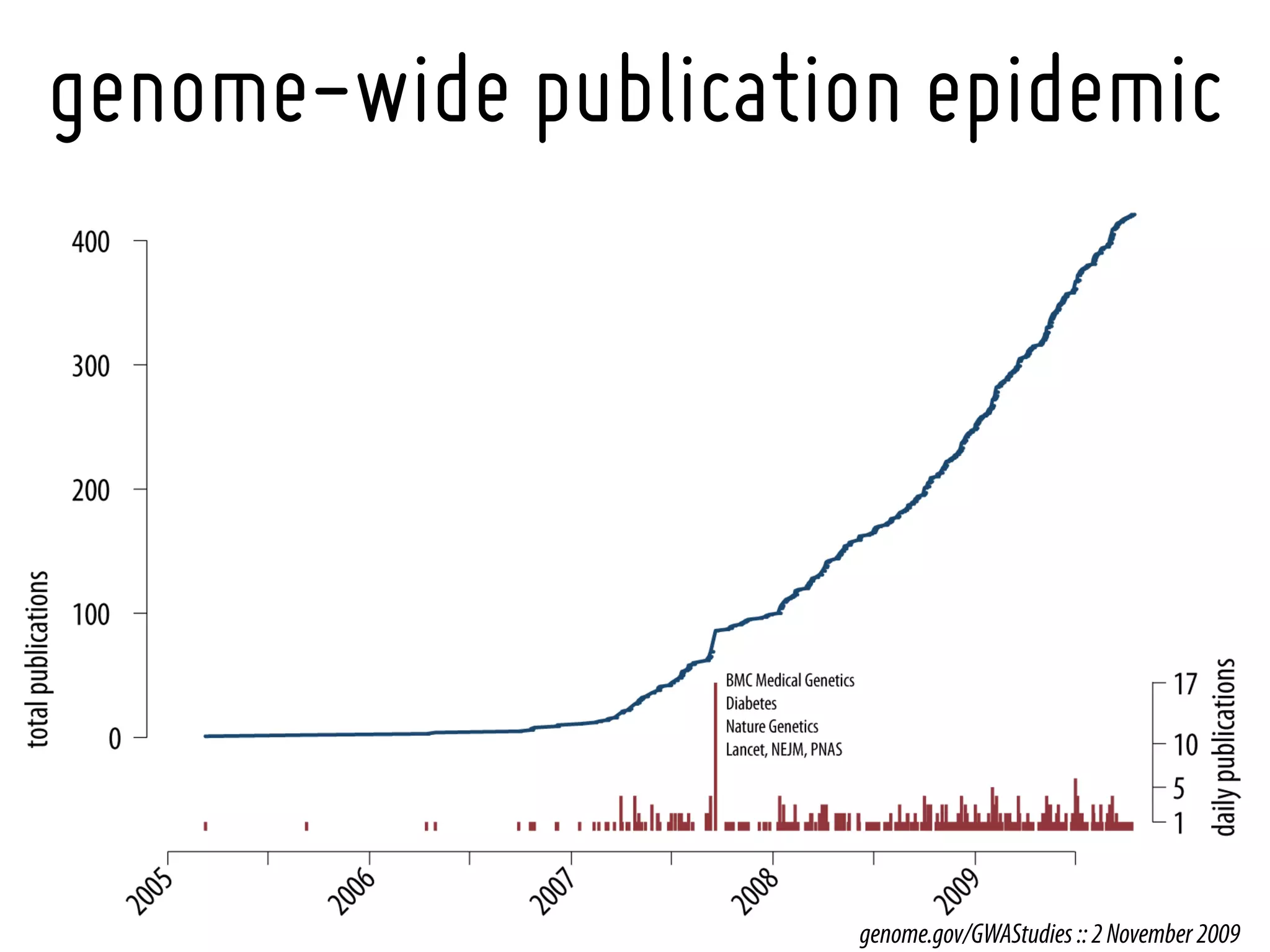
21. NHGRI catalog of published GWA studies (http://genome.gov/GWASstudies)](https://image.slidesharecdn.com/epi519gwas-talk-1224190848163468-9/75/Epi519-Gwas-Talk-64-2048.jpg)
