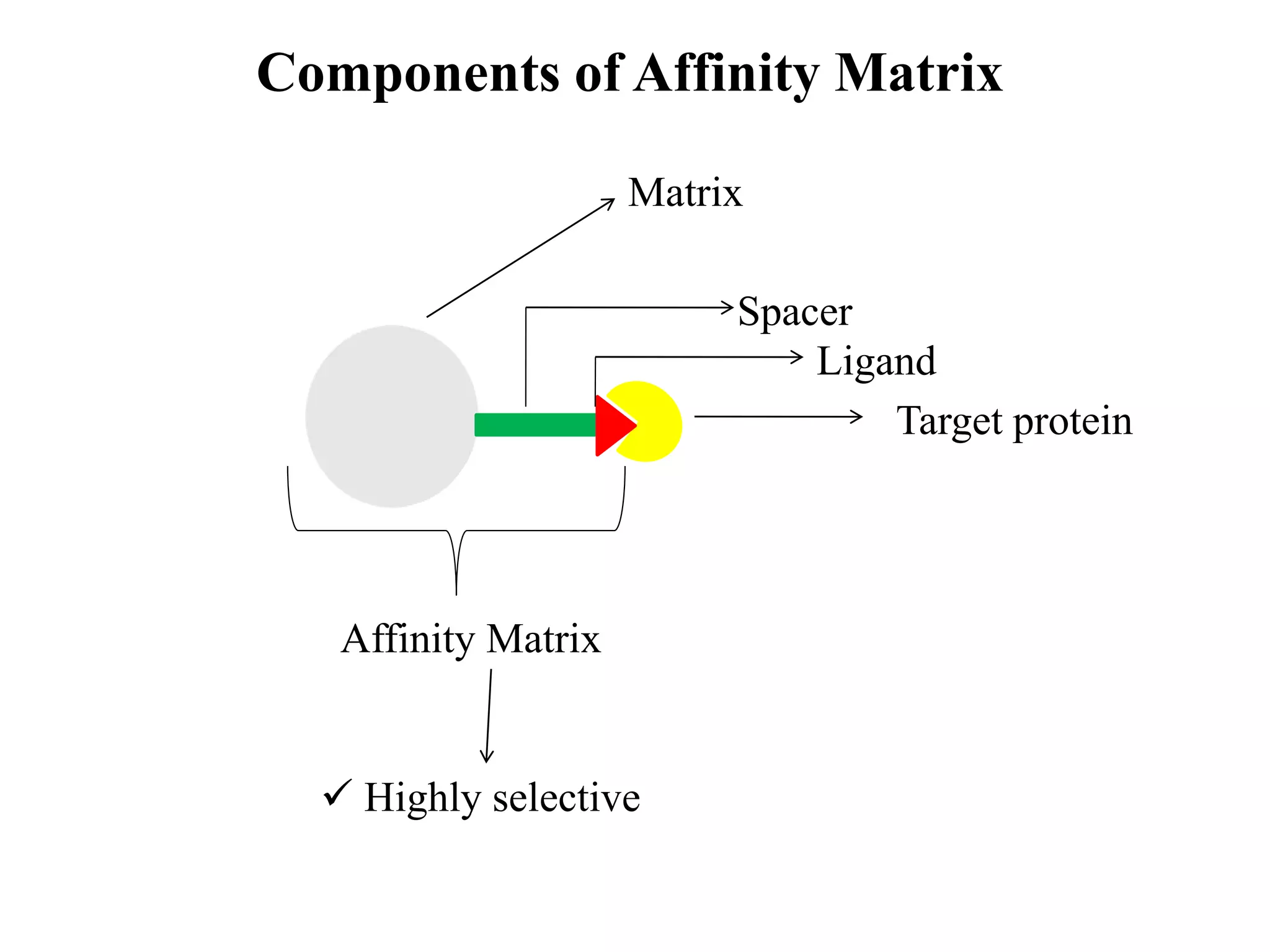
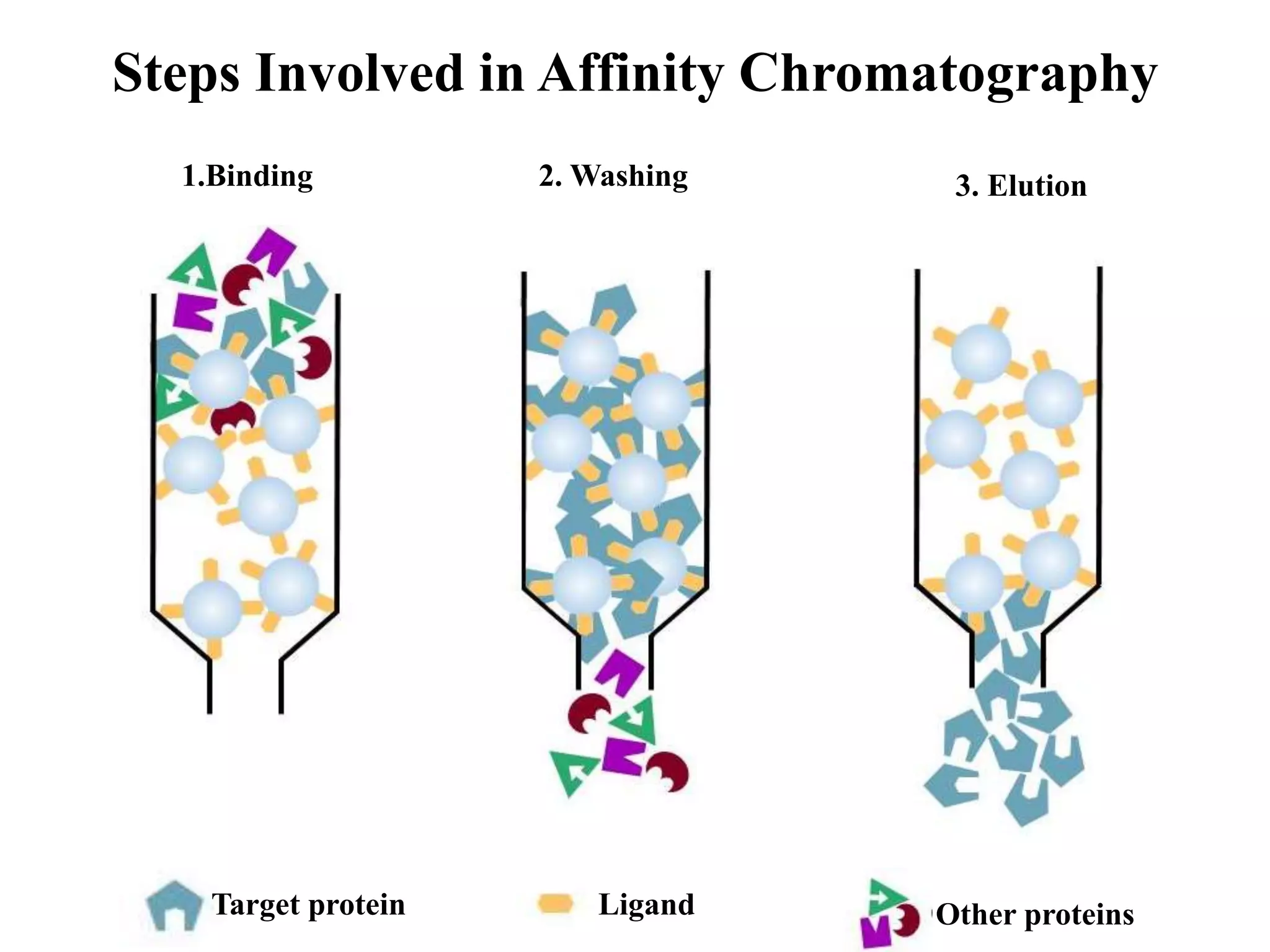
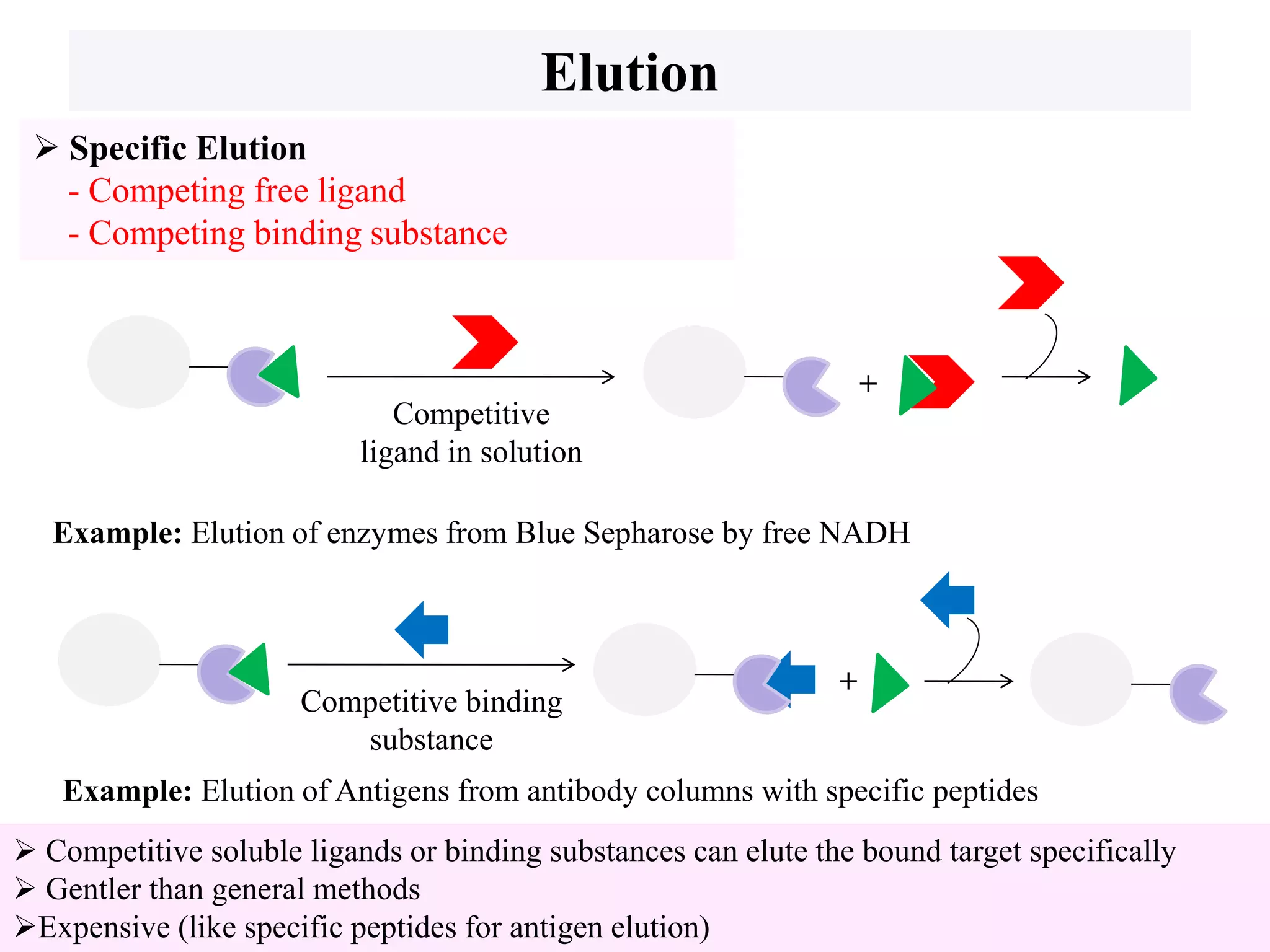
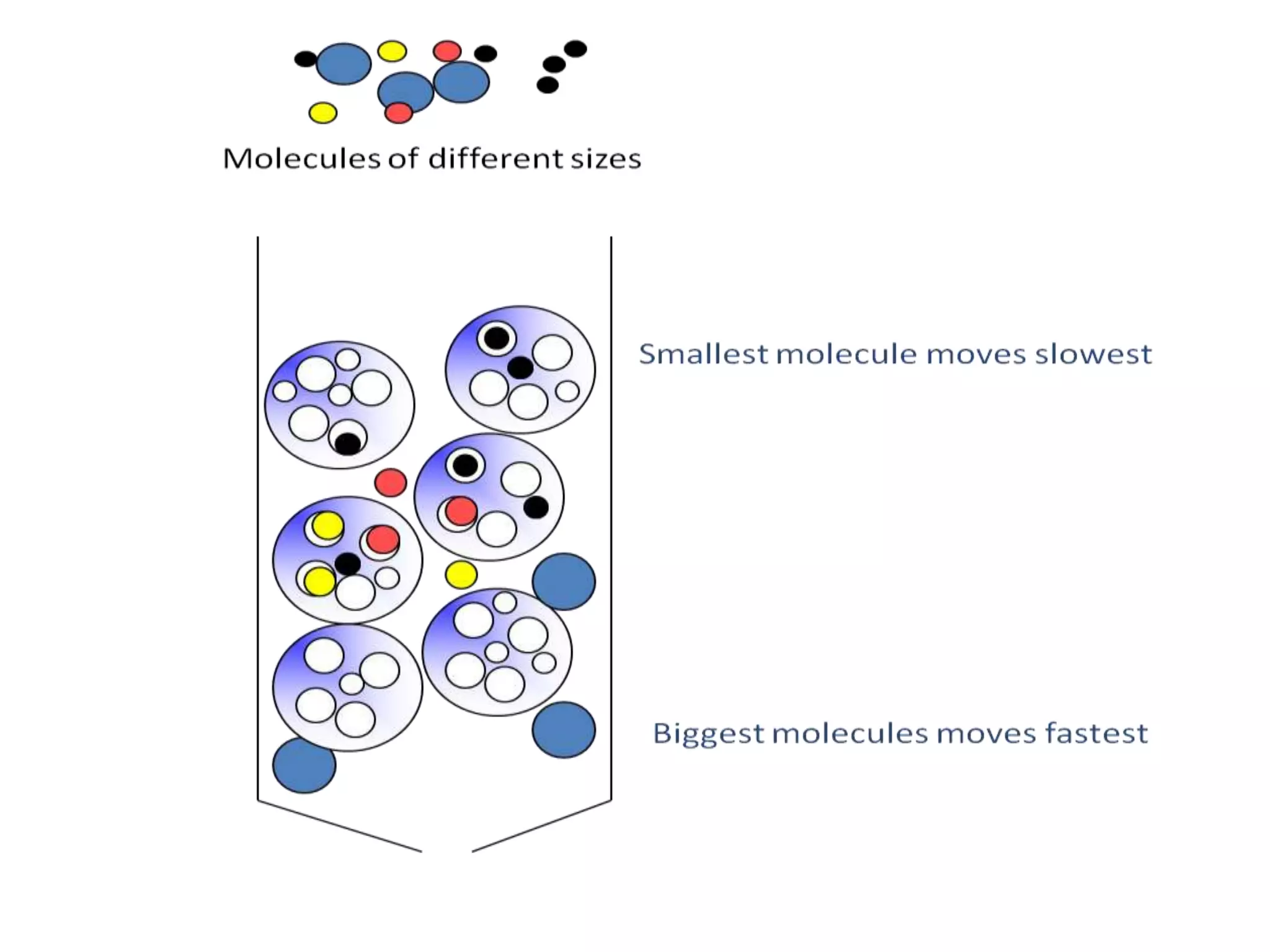
This document discusses affinity chromatography, a technique used to selectively separate or purify molecules from biochemical mixtures. It works by reversible and specific interactions between molecules like proteins, nucleic acids, or enzymes and a ligand immobilized on an inert support packed in a column. When a mixture passes through, the target molecule that binds strongly to the ligand is separated from others. The document describes the principle, components like matrix, ligand, and spacer, as well as steps involved in affinity chromatography including binding, washing, and specific elution methods. It also discusses applications like purification of substances from biological mixtures.