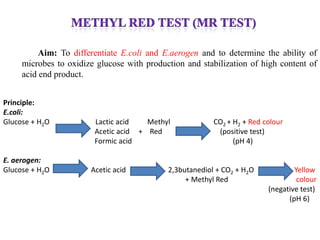
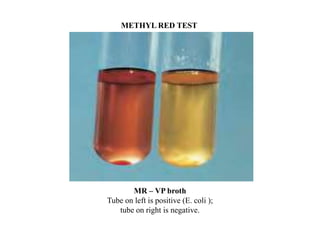
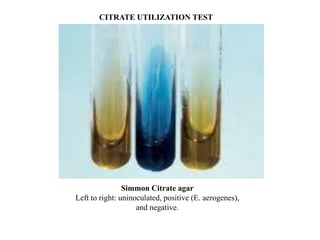
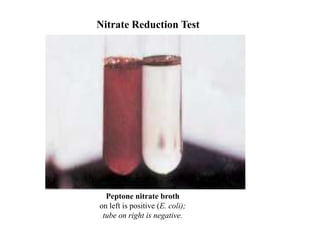
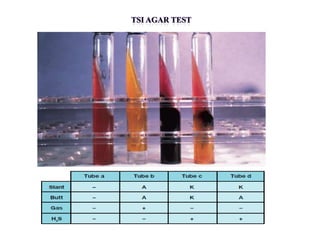
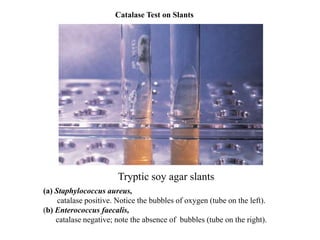
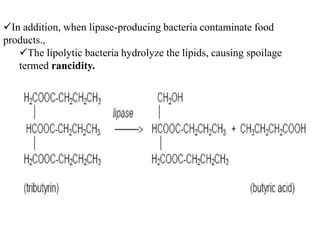
The document provides instructions for performing several microbiological tests, including sugar fermentation tests, indole production, methyl red, Voges-Proskauer, citrate utilization, nitrate reduction, urease, triple sugar iron, oxidase, catalase, amylase, and lipase. It explains the principles, expected results, and interpretations for each test. The tests can be used to differentiate bacterial species and determine their metabolic abilities.