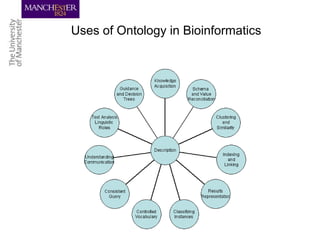
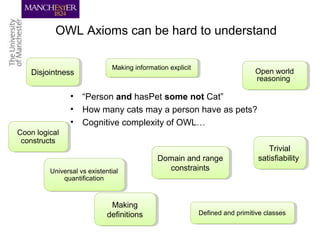
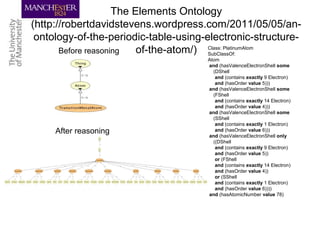
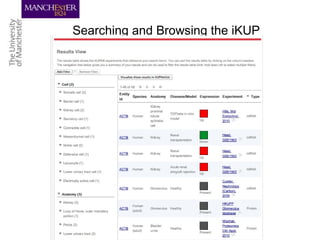
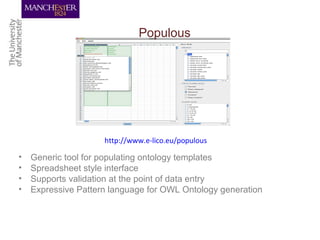
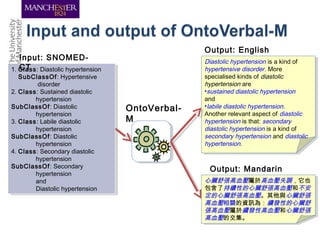
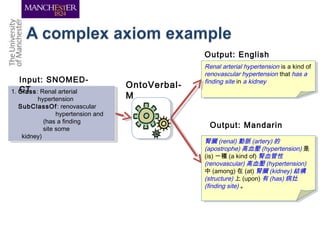
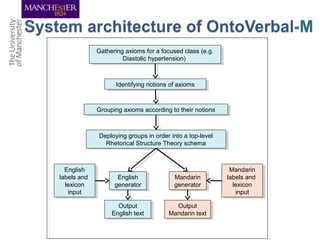
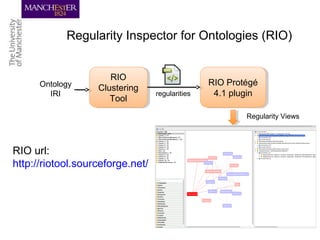
This document discusses techniques for managing and understanding large biomedical ontologies authored in OWL. It notes that OWL ontologies can be difficult to author and comprehend due to their complexity. However, several approaches are making ontologies more manageable, such as generating natural language descriptions, abstracting ontologies to find patterns of axioms, and hiding the OWL syntax from end users. Tools like Protege plugins can help apply these techniques to improve ontology development and application.













![Examples of regularities in SNOMED-CT
• Regularities expressed as axioms with variables holding similar
entities:
– E.g. ?Chronic_finding = [Chronic pyonephrosis (disorder),
Chronic pneumothorax (disorder)]
1. Syntactic Regularity describing ‘Chronic findings’:
?Chronic_finding EquivalentTo ?Disorder and (RoleGroup some
(‘Clinical course (attribute)’ some ‘Chronic (qualifier value)’))
?Chronic_finding EquivalentTo ?Disorder and (RoleGroup some
(‘Clinical course (attribute)’ some ‘Chronic (qualifier value)’))
2. Syntactic Regularity describing ‘Acute findings’:
?Acute_finding EquivalentTo ?Disorder and (RoleGroup some
(‘Clinical course (attribute)’ some
'Sudden onset AND/OR short duration (qualifier value)'))
?Acute_finding EquivalentTo ?Disorder and (RoleGroup some
(‘Clinical course (attribute)’ some
'Sudden onset AND/OR short duration (qualifier value)'))](https://image.slidesharecdn.com/wj5zoeqktzgo2fqyykp0-signature-f84076d802018ccc31f808b4d674dd5edfbd24357d894cb2420d389f15b69e02-poli-140716122622-phpapp01/85/Working-with-big-biomedical-ontologies-25-320.jpg)


