This document discusses the use and future of ontologies in biology. It makes three key points:
1) Ontologies are necessary for representing biological knowledge but are not sufficient on their own.
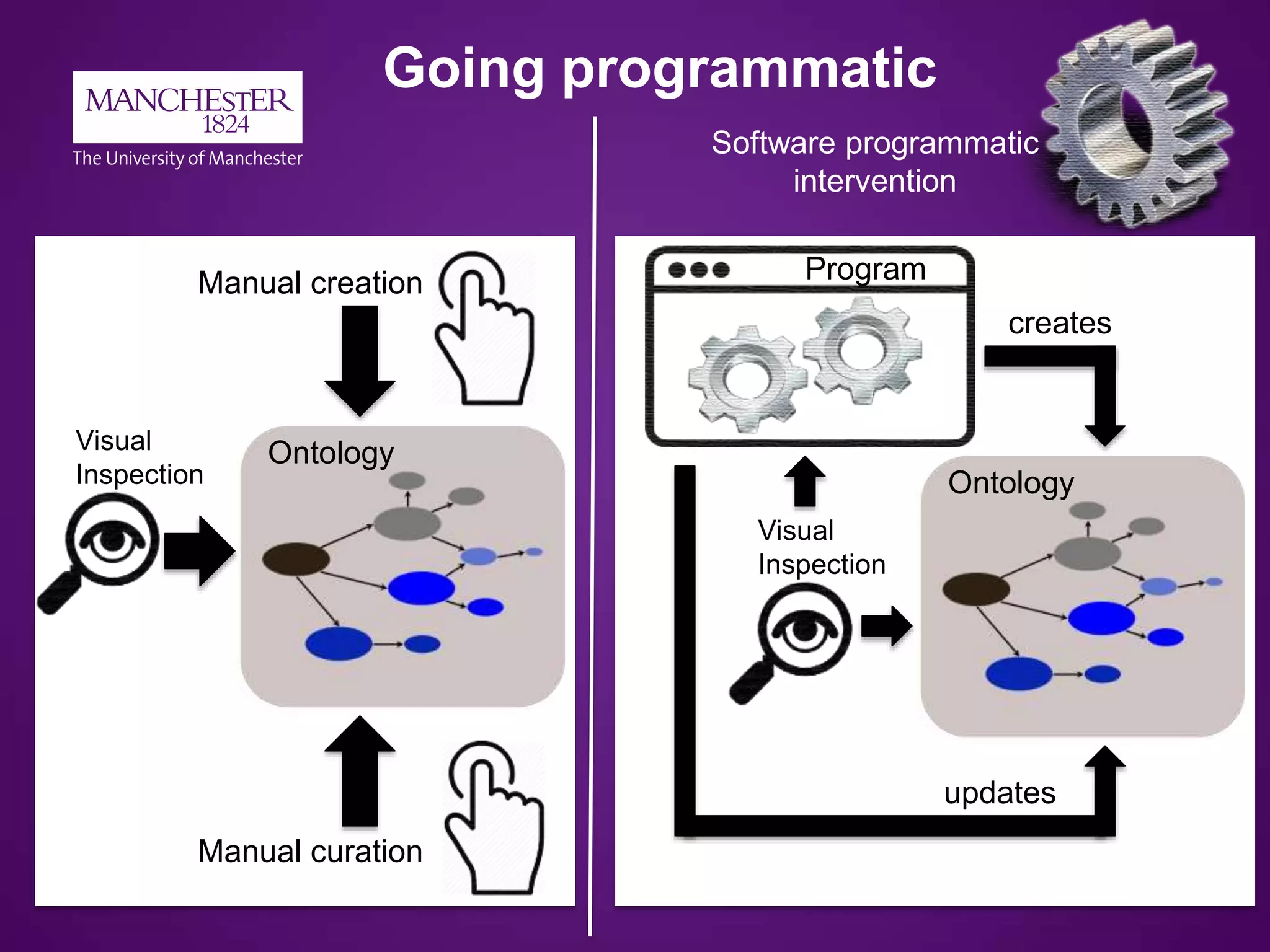
2) Ontology development needs to become more programmatic and automated to scale to the vast amount of biological data and knowledge.
3) Ontologies should be used to generate new conclusions and insights about biology, not just organize existing knowledge. More work is needed to enable reasoning over ontologies.


![Number of PubMed papers per year for
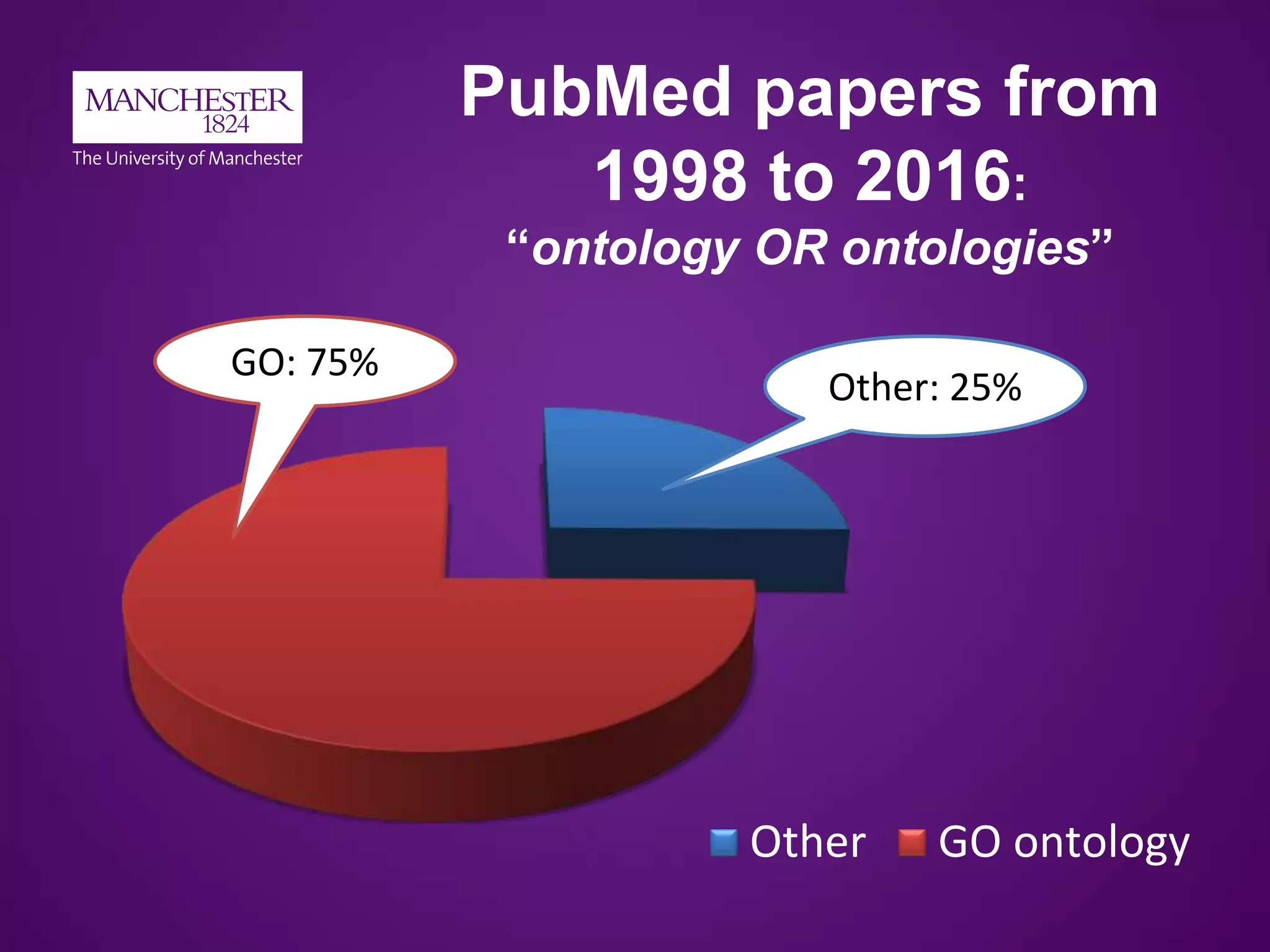
1998 to 2016 (without normalisation)
PubMed search:
(ontology[All Fields] OR ontologies[All Fields])
Number of PubMed papers per year for
1998 to 2016 (with normalisation)](https://image.slidesharecdn.com/bio-ontologies-rds-2017-170728193135/75/Ontologies-Necessary-but-not-sufficient-3-2048.jpg)
![Number of PubMed papers per year for
1998 to 2016 (without normalisation)
PubMed search:
("gene ontology"[MeSH Terms] OR ("gene"[All Fields] AND
"ontology"[All Fields]) OR "gene ontology"[All Fields])
Number of PubMed papers per year for
1998 to 2016 (with normalisation)](https://image.slidesharecdn.com/bio-ontologies-rds-2017-170728193135/75/Ontologies-Necessary-but-not-sufficient-4-2048.jpg)
![Word cloud for all the citations of
the 1998 search
Total 35
citations
Text from all
fields
No data
cleaning
Make word cloud via https://www.jasondavies.com/wordcloud/
PubMed search: (ontology[All Fields] OR ontologies[All Fields])
AND (”1998/01/01"[PDAT] : ”1998/12/31"[PDAT])](https://image.slidesharecdn.com/bio-ontologies-rds-2017-170728193135/75/Ontologies-Necessary-but-not-sufficient-6-2048.jpg)
![Word cloud for all the citations of
the 2005 search
First 200
citations of a
total of 516
citations for
for 2005
Text from all
fields
No data
cleaning
Make word cloud via https://www.jasondavies.com/wordcloud/
PubMed search: (ontology[All Fields] OR ontologies[All Fields])
AND ("2005/01/01"[PDAT] : "2005/12/31"[PDAT])](https://image.slidesharecdn.com/bio-ontologies-rds-2017-170728193135/75/Ontologies-Necessary-but-not-sufficient-7-2048.jpg)
![Word cloud for the first 200
citations of the 2016 search
First 200
citations of a
total of 2647
for 2016
Text from all
fields
No data
cleaning
Make word cloud via https://www.jasondavies.com/wordcloud/
PubMed search: (ontology[All Fields] OR ontologies[All Fields])
AND ("2016/01/01"[PDAT] : "2016/12/31"[PDAT])](https://image.slidesharecdn.com/bio-ontologies-rds-2017-170728193135/75/Ontologies-Necessary-but-not-sufficient-8-2048.jpg)