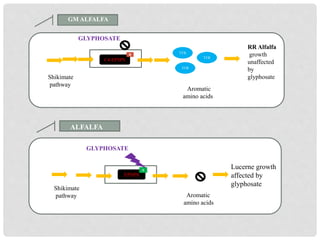
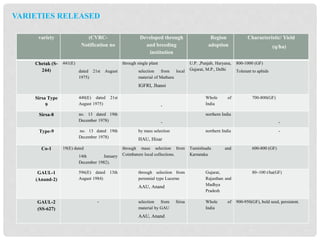
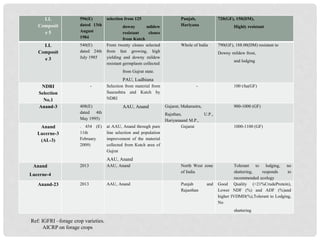
The document discusses lucerne (alfalfa) breeding, detailing its progress, breeding methods, and constraints faced. It highlights advancements in breeding techniques over the past century and the development of genetically modified varieties, such as Roundup Ready alfalfa, while addressing the challenges associated with cross-pollination and genetic diversity. The future prospects emphasize the need for improved hardiness, disease resistance, and biotechnological approaches to enhance yield and adaptability.

![SCIENTIFIC CLASSIFICATION :
Kingdom : Plantae
Subkingdom : Tracheobionta
Super division: Spermatophyta
Division : Magnoliophyta
Class : Magnoliopsida
Subclass : Rosidae
Order : Fabales
Family : Fabaceae
Genus : Medicago L.
Species : Medicago sativa L
Common name: alfalfa,rijka
Origin: south west asia
Chromosome no.: 2n=4x=32[auto tetraploid]
Wild relatives :Medicago falcata 2n=16 [diploid]
Medicago coerulea 2n=16 ,2n=32[diploid and tetraploid]](https://image.slidesharecdn.com/lucernebreedingprogressandconstraintsppt-210611064500/85/Lucerne-breeding-methods-progress-and-constraints-2-320.jpg)