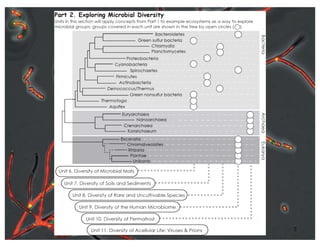
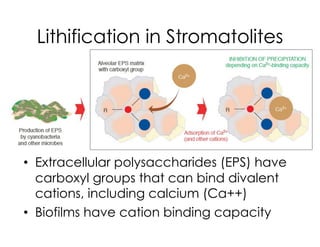
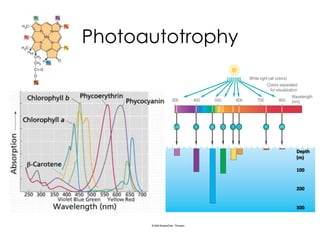
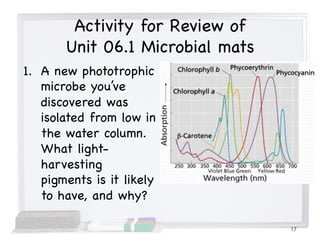
This document covers the diversity of microbial mats, including the definitions and characteristics of microbial mats, their layers, and functional guilds of microbes. It also contrasts the carbon fixation methods and energy acquisition strategies of three photosynthetic bacterial phyla and describes adaptations of two thermophilic bacterial phyla. The learning goals are aimed at understanding how these microbial communities interact and the implications for early life on Earth.