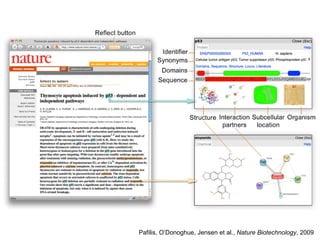
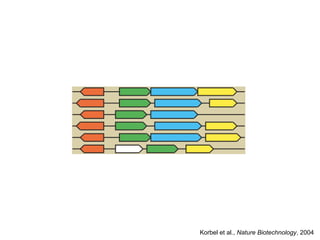
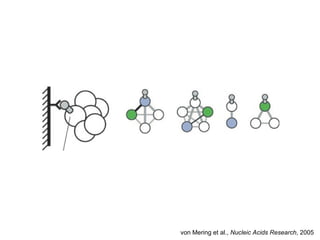
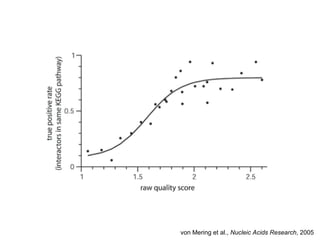
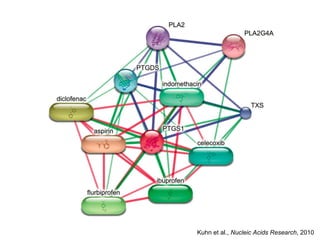
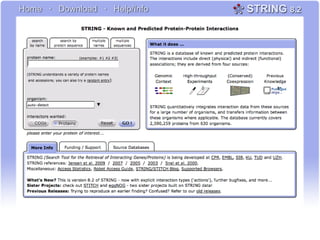
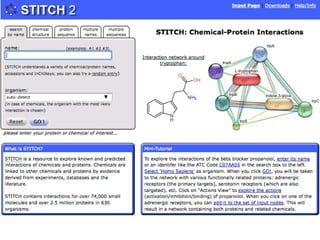
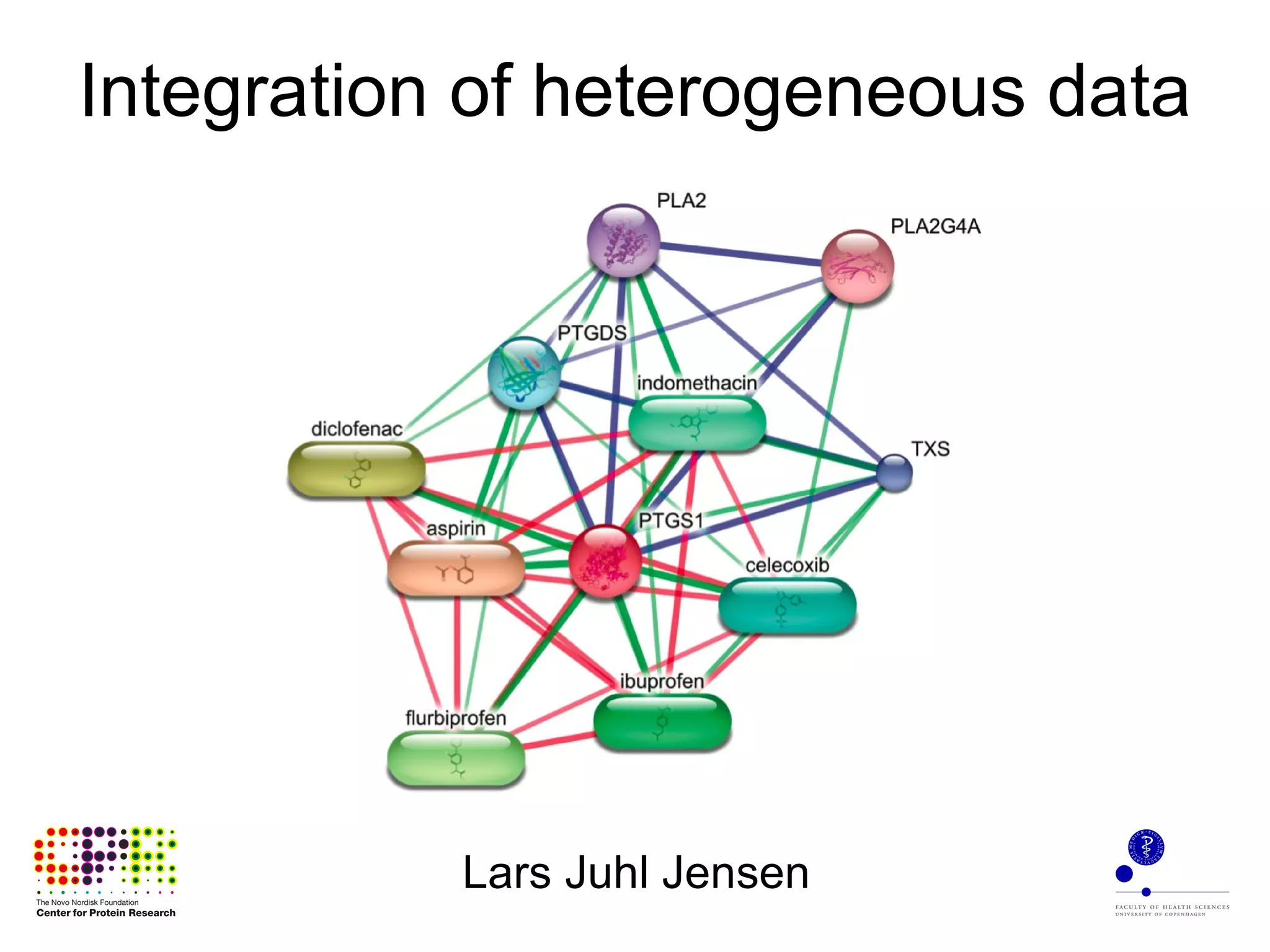
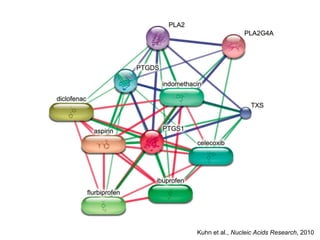
The document discusses the integration of heterogeneous biological data sources, including genomes, proteins, and small molecules, emphasizing their varying formats and identifiers. It highlights the challenges in data comparison and quality, as well as the importance of advanced methods in natural language processing for extracting relationships from literature. Various databases and methodologies for gene interactions, expression, and pathways are reviewed to enhance data accessibility for both humans and computational systems.









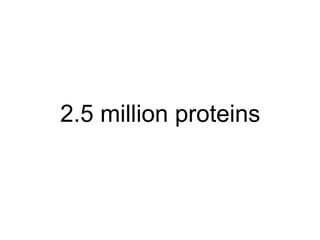



































































![Gene and protein names Cue words for entity recognition Verbs for relation extraction [ nxgene The GAL4 gene ] [ nxexpr T he expression of [ nxgene the cytochrome genes [ nxpg CYC1 and CYC7 ]]] is controlled by [ nxpg HAP1 ]](https://image.slidesharecdn.com/jen10talk5-100315121007-phpapp02/85/Integration-of-heterogeneous-data-90-320.jpg)