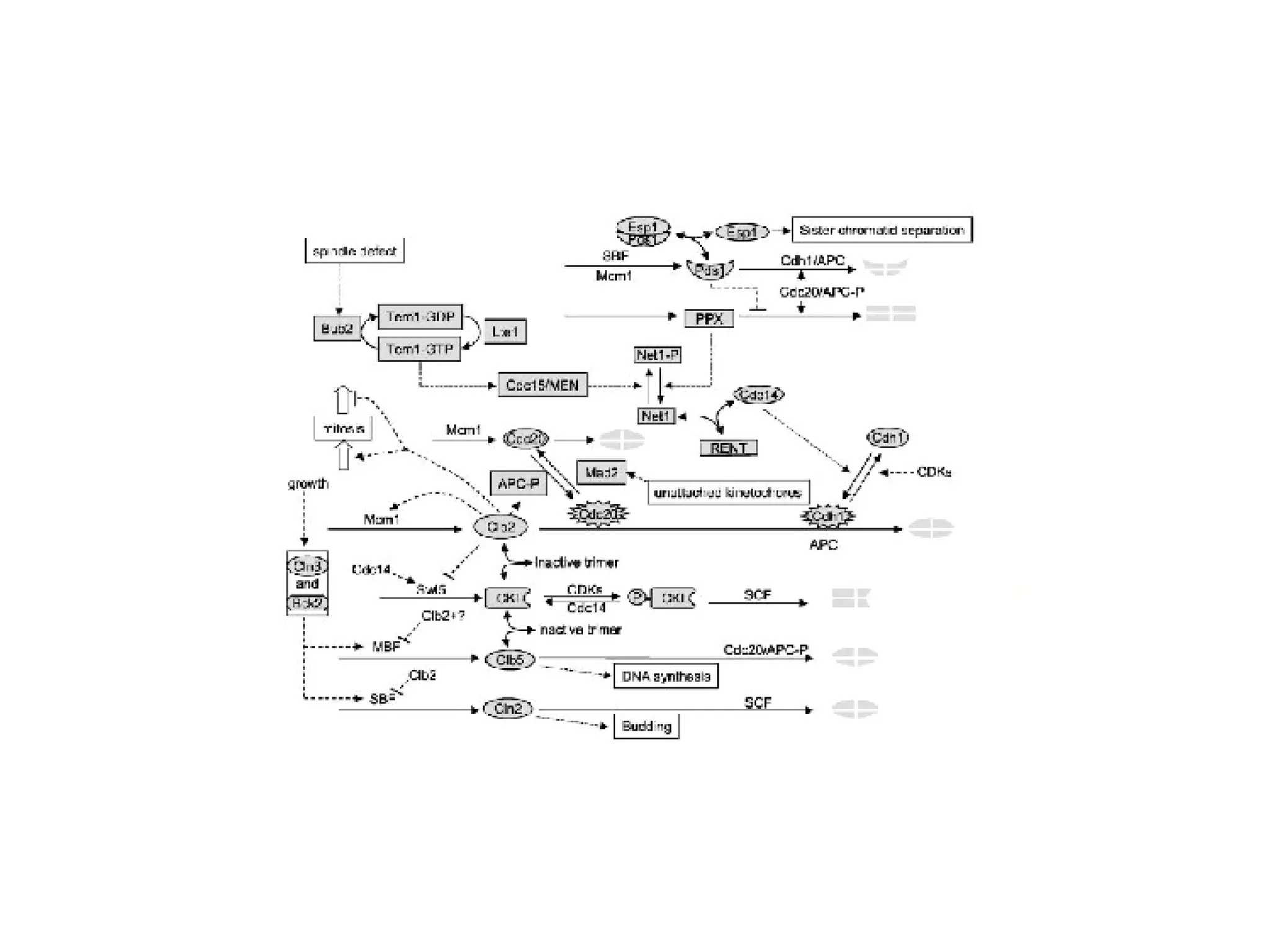
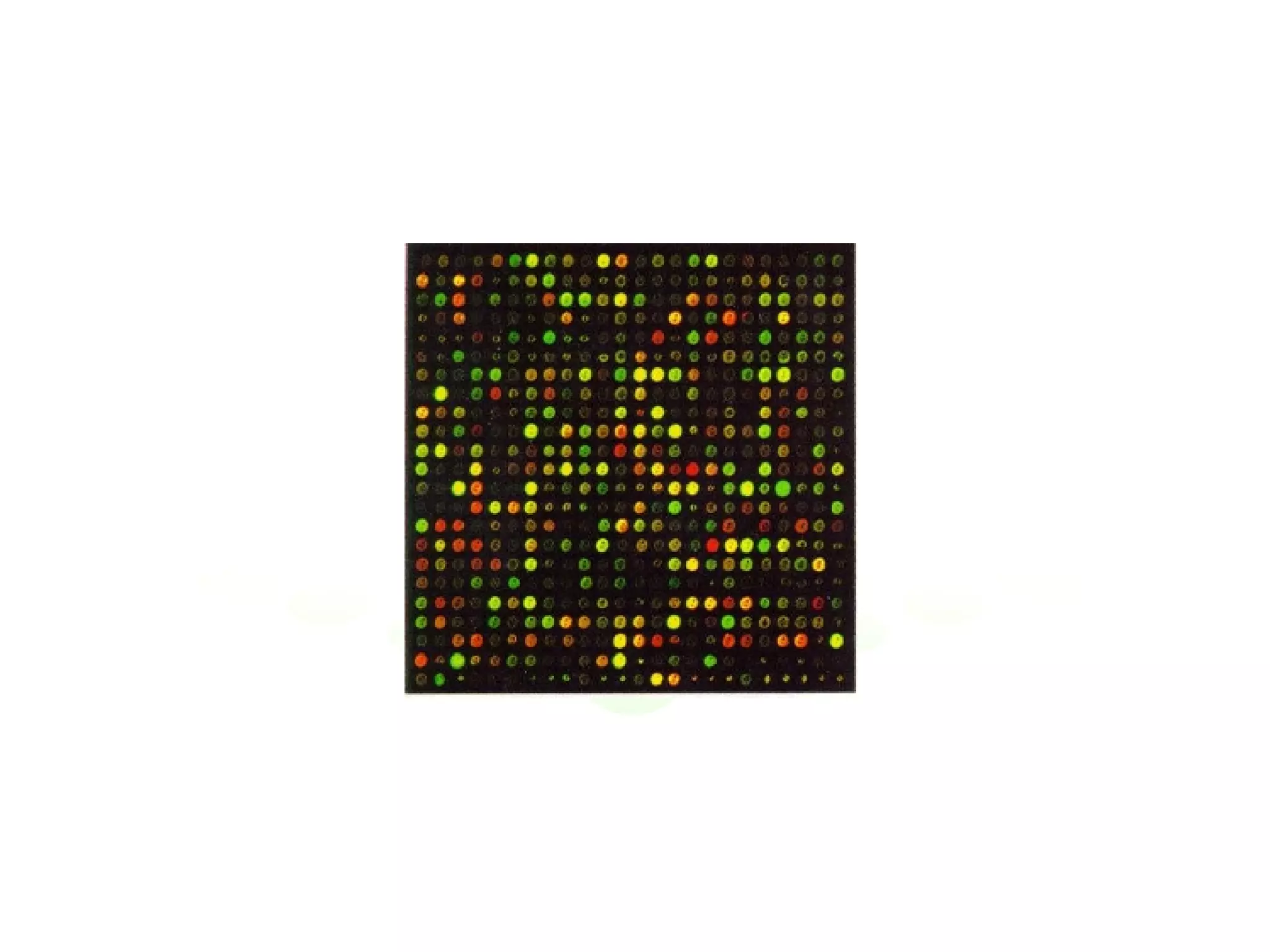
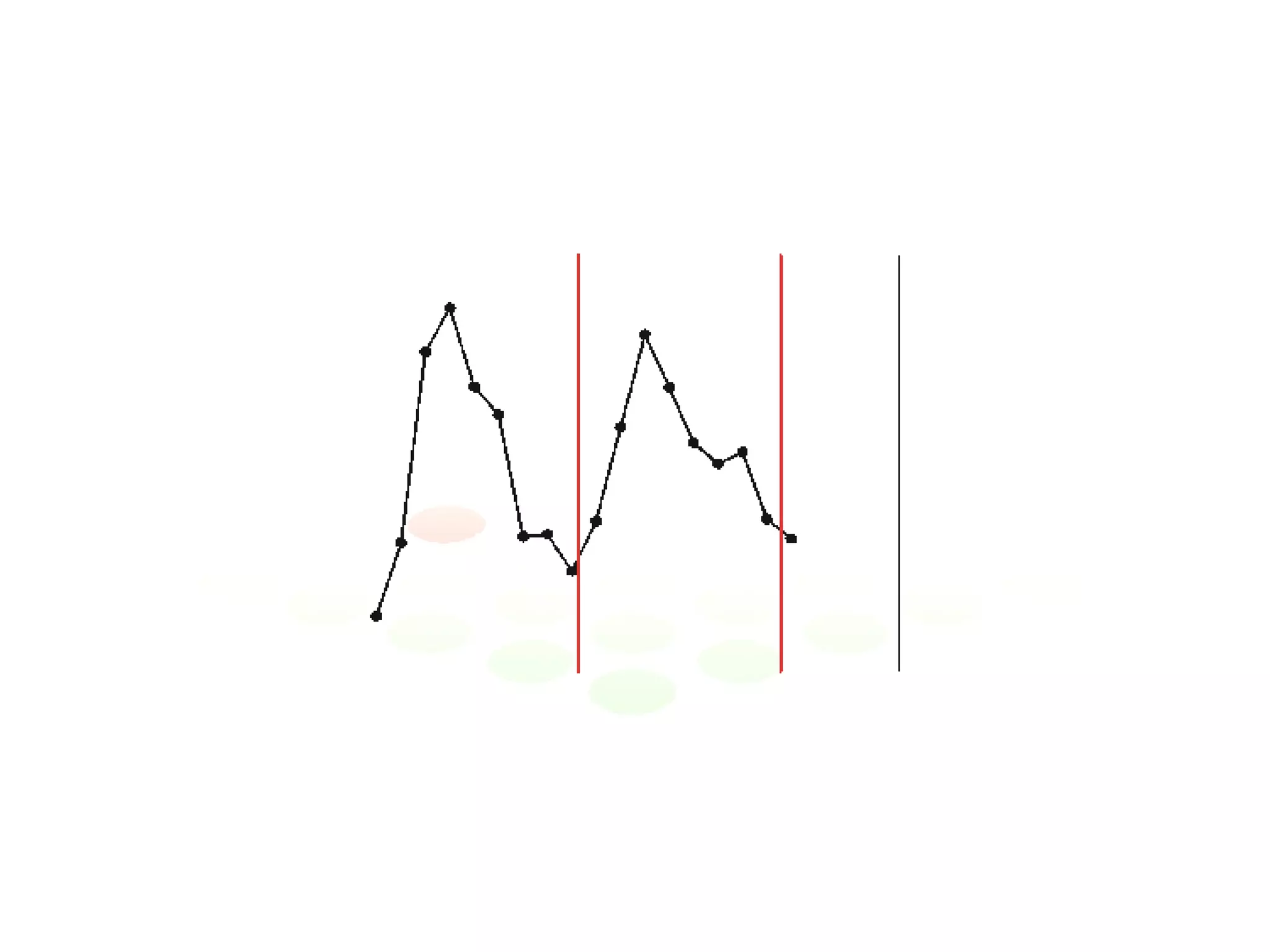
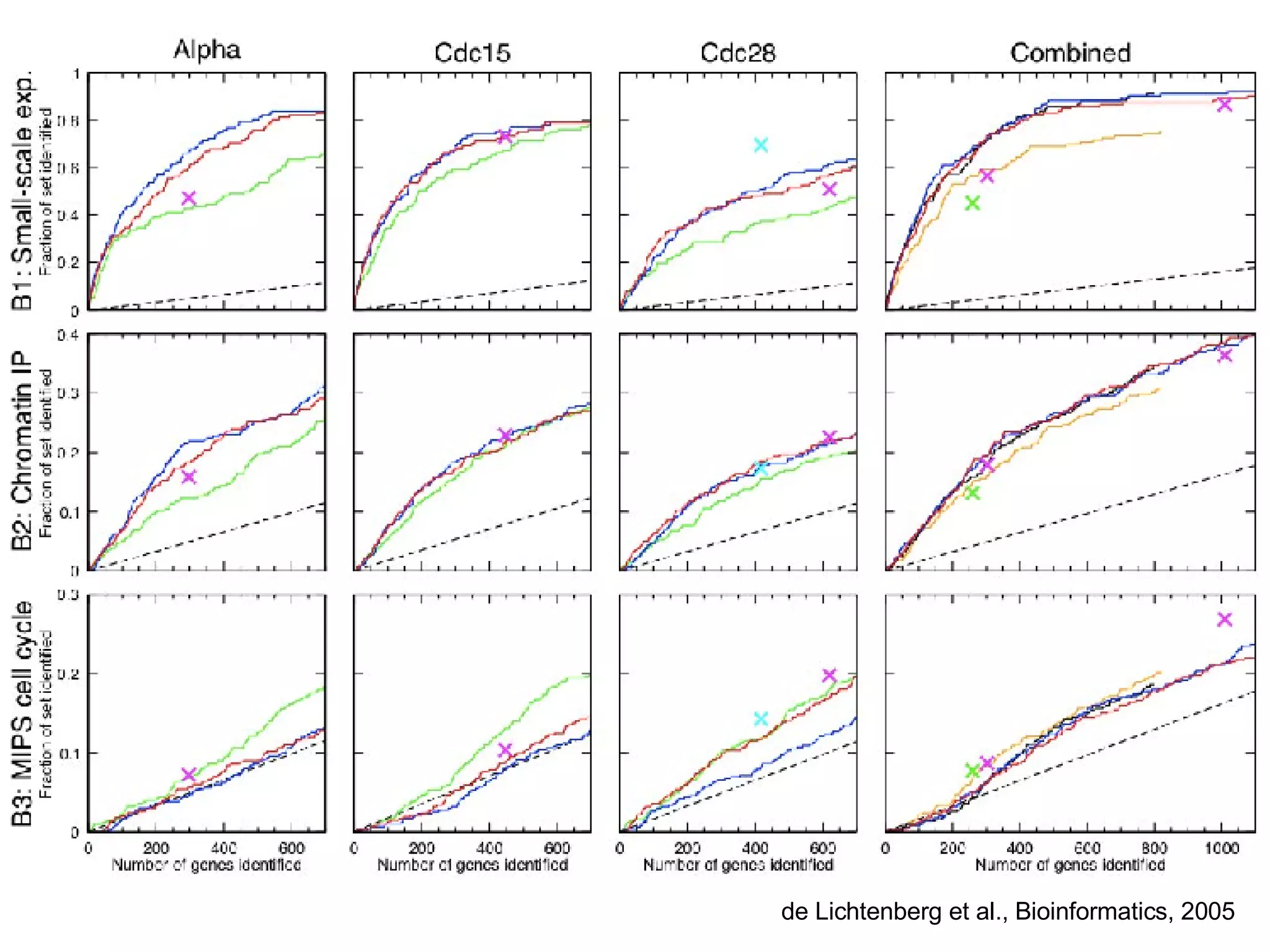
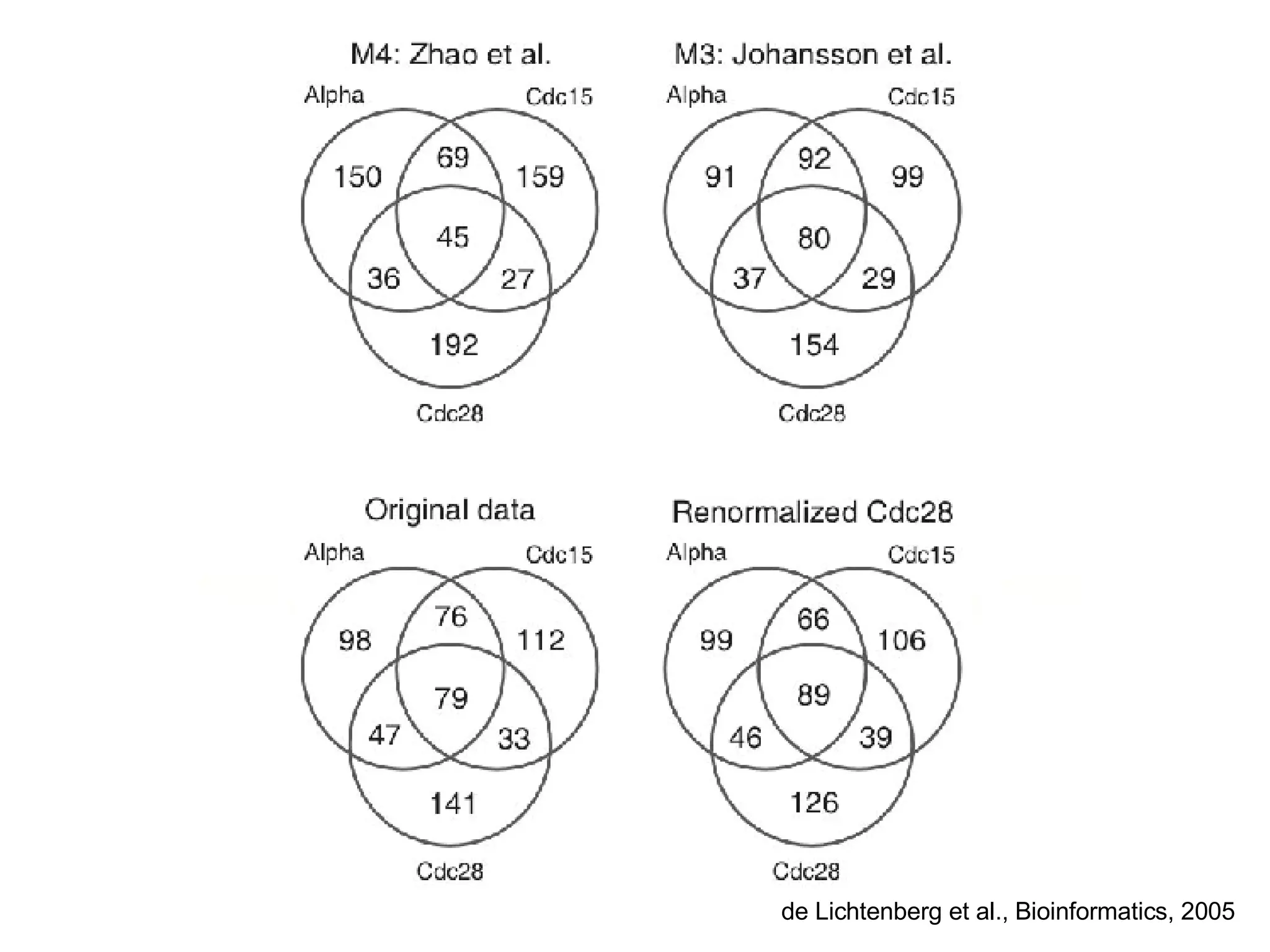
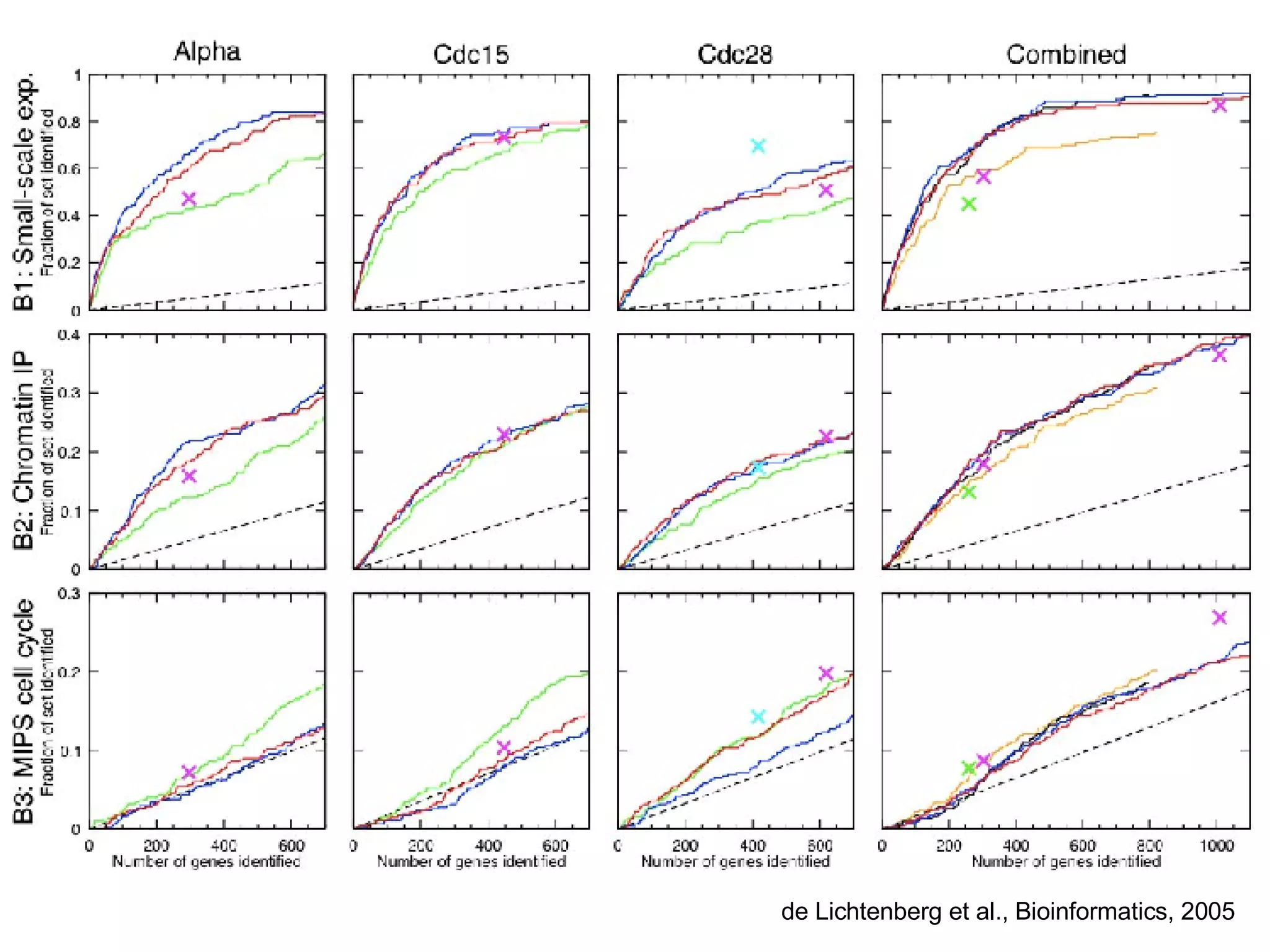
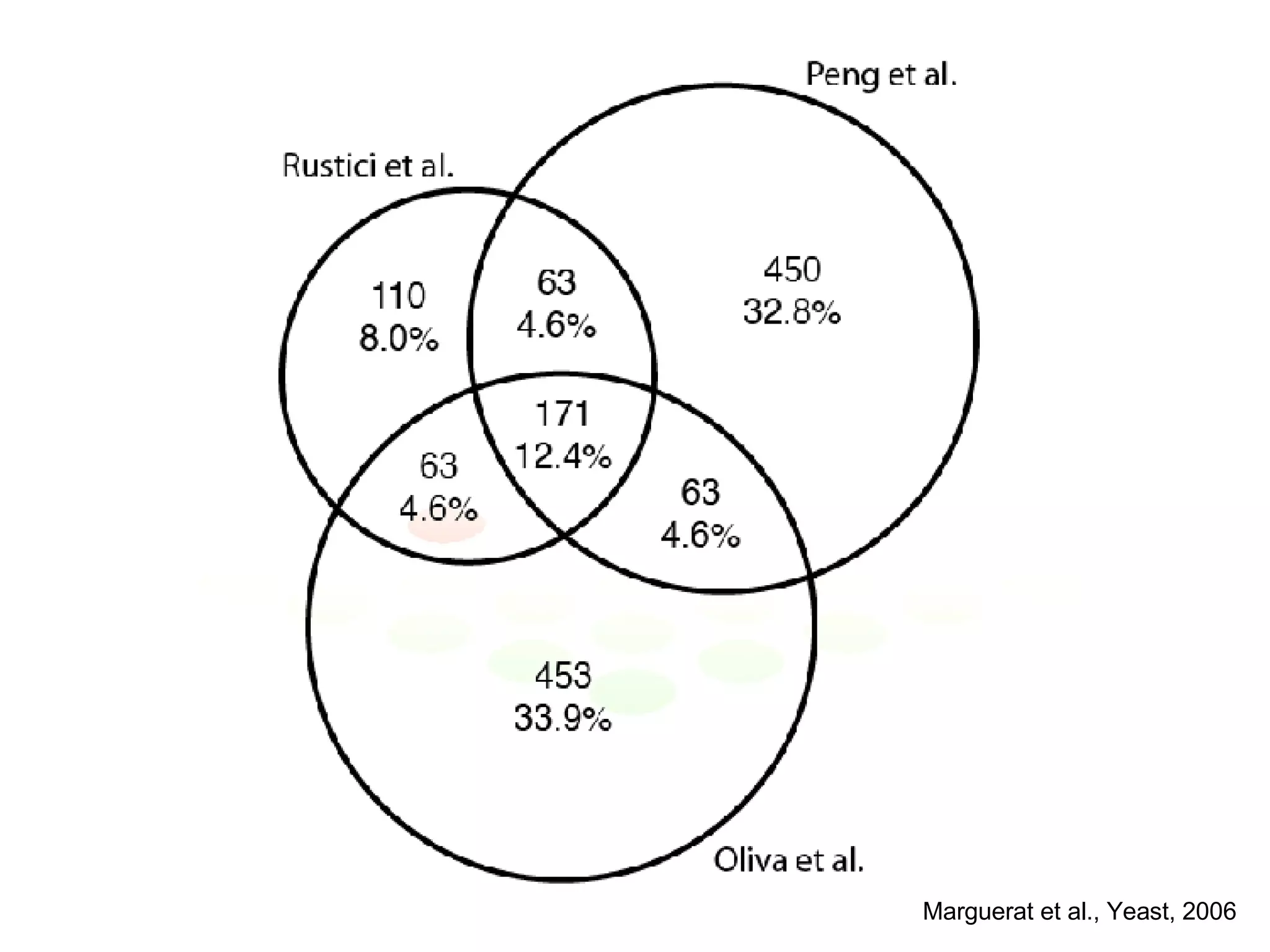
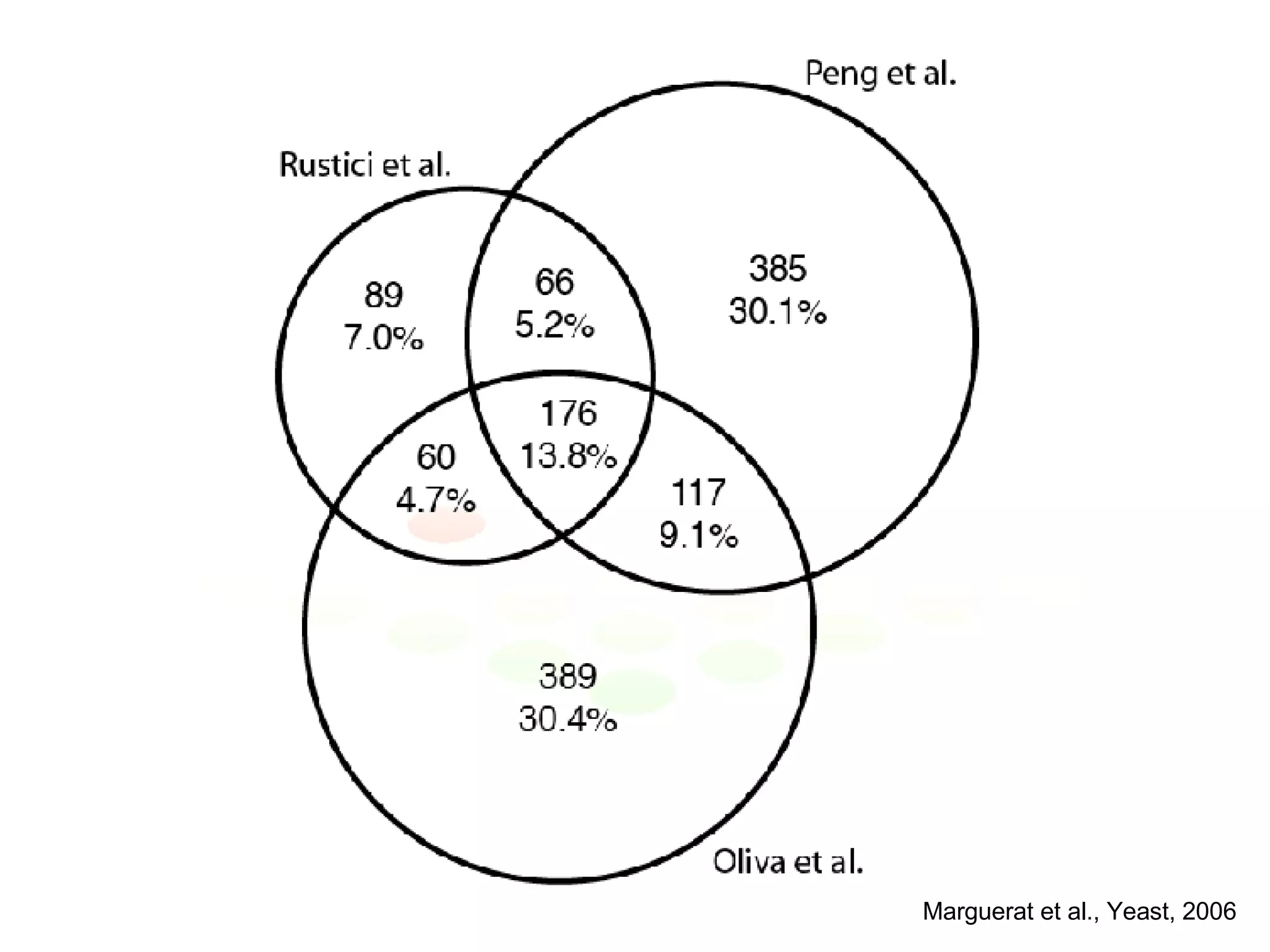
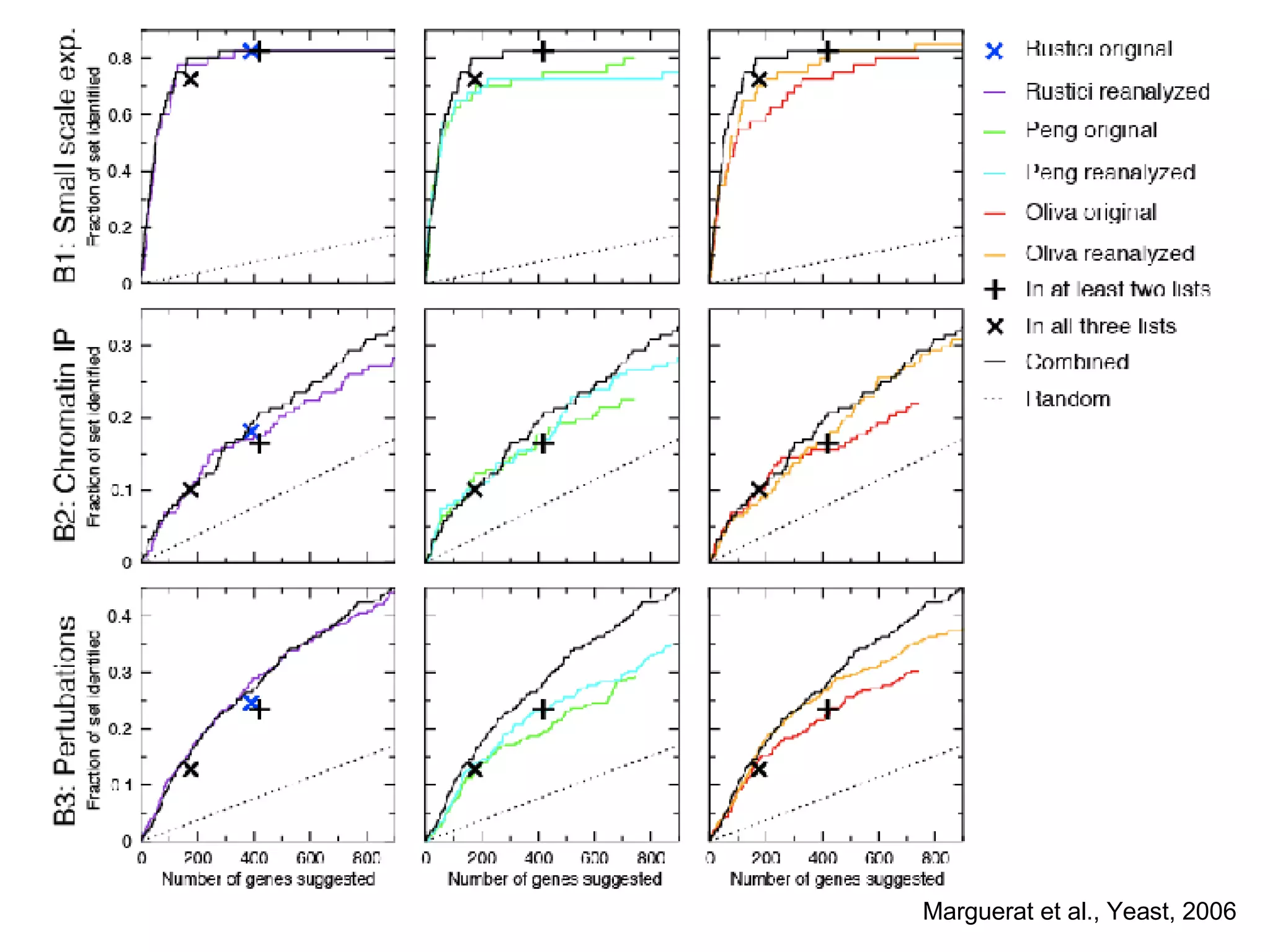
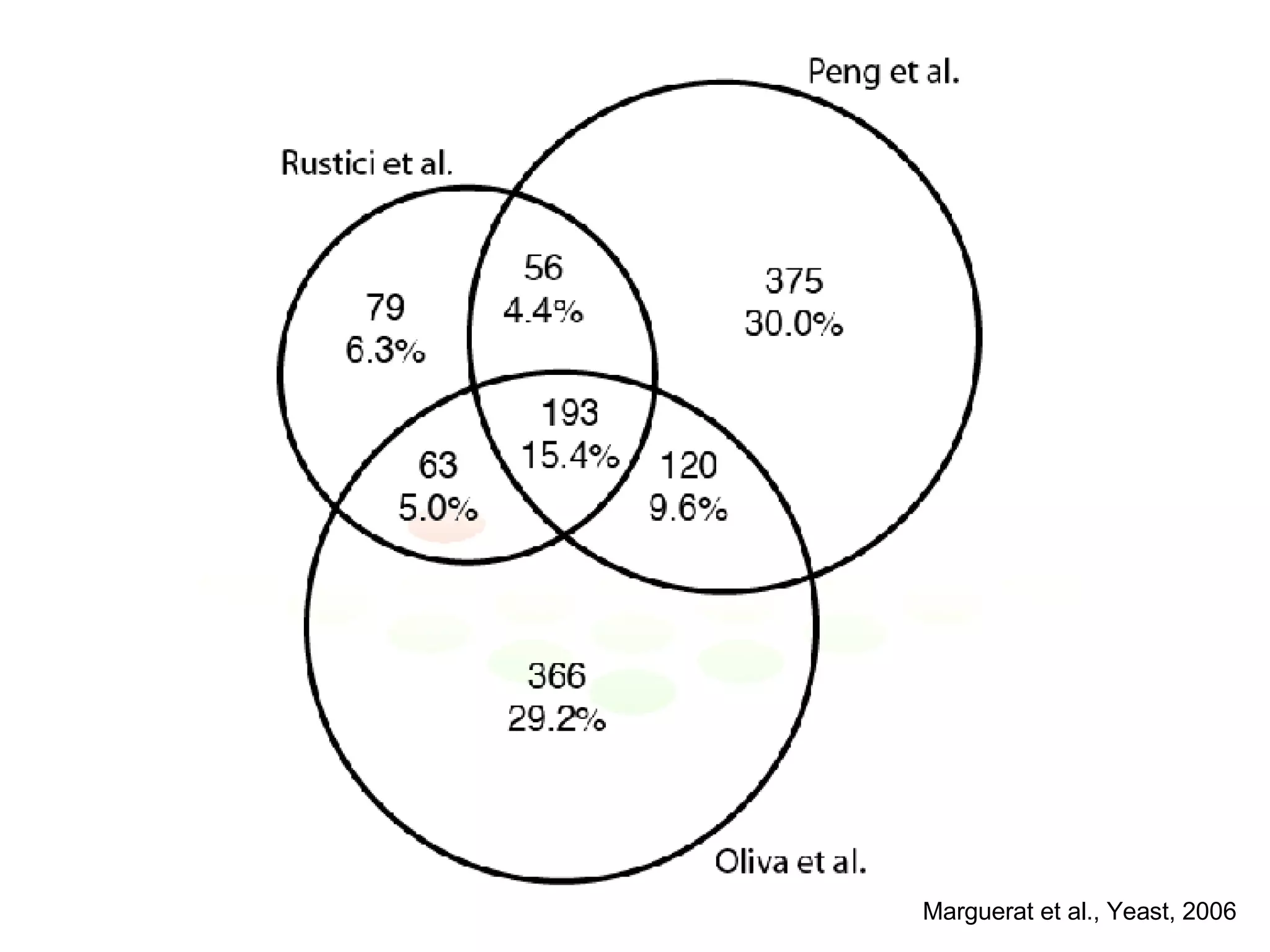
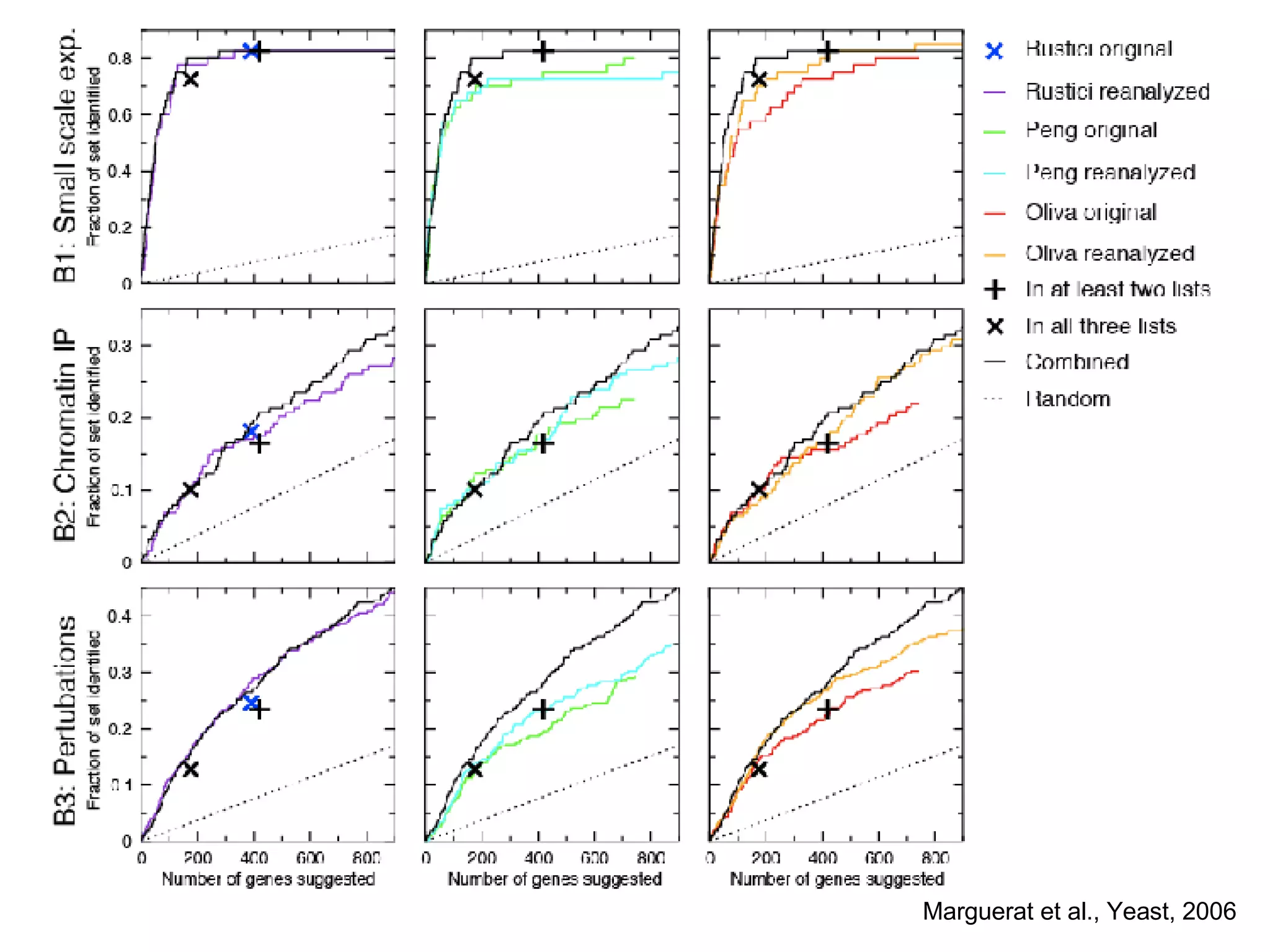
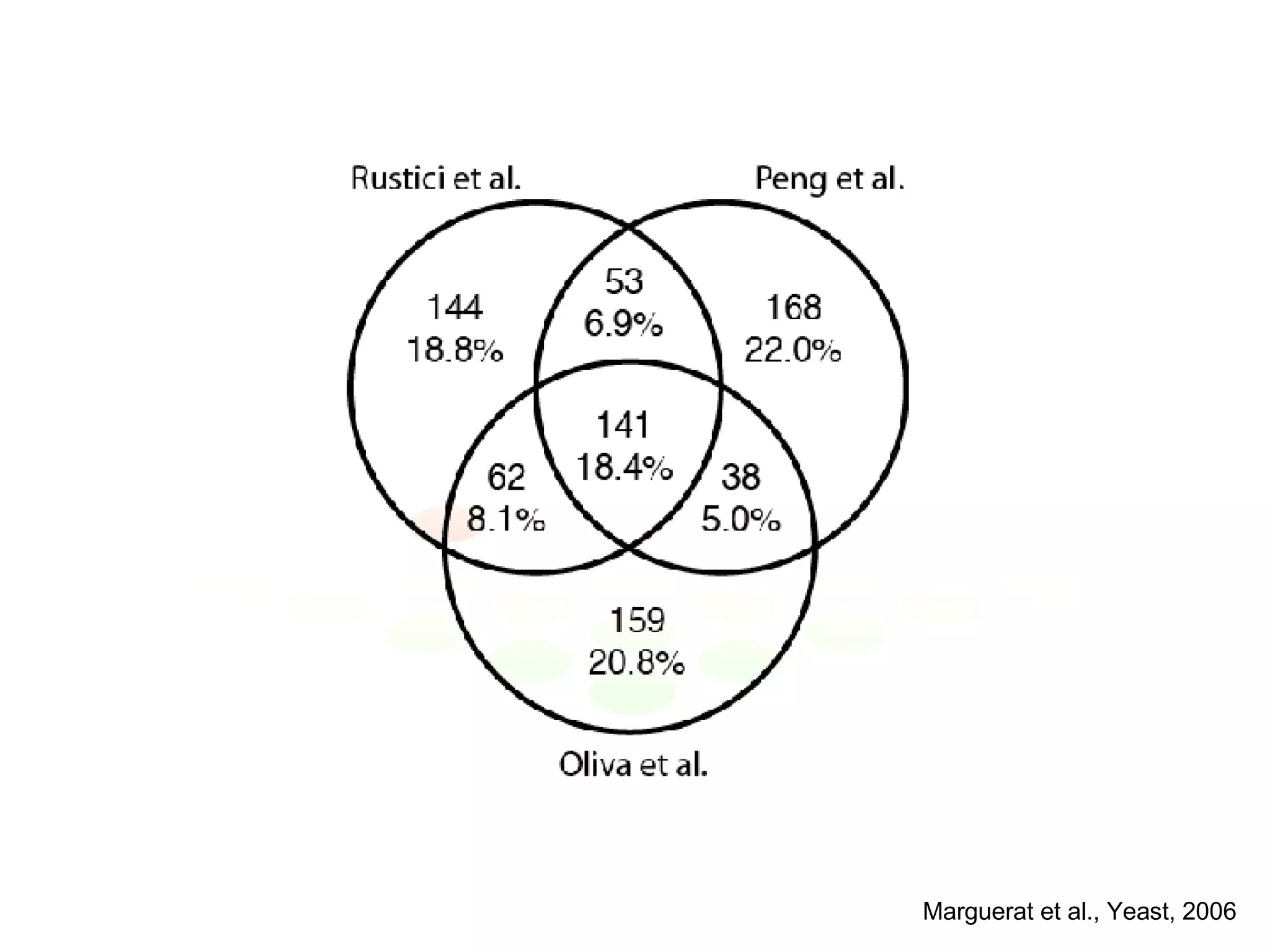
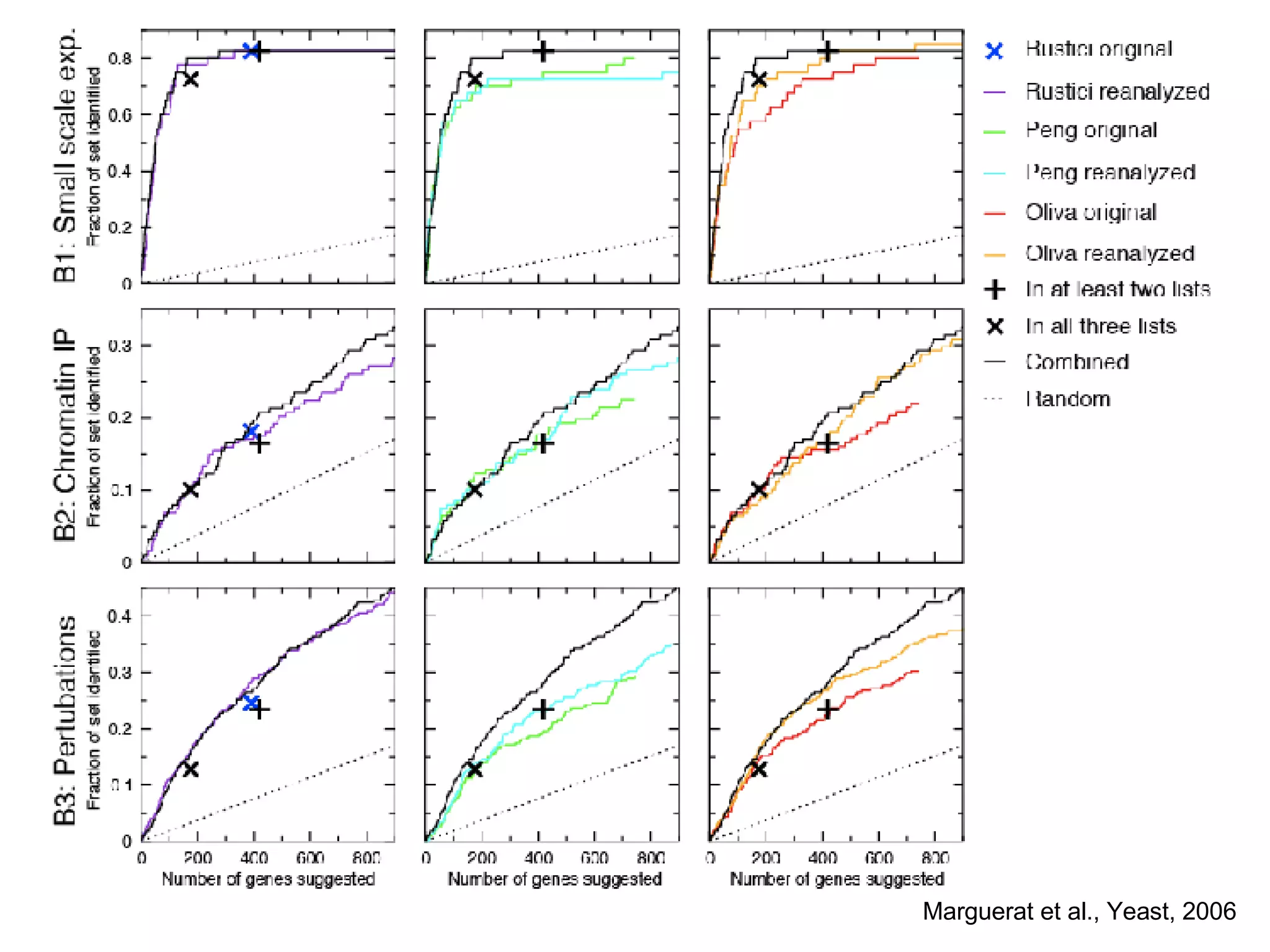
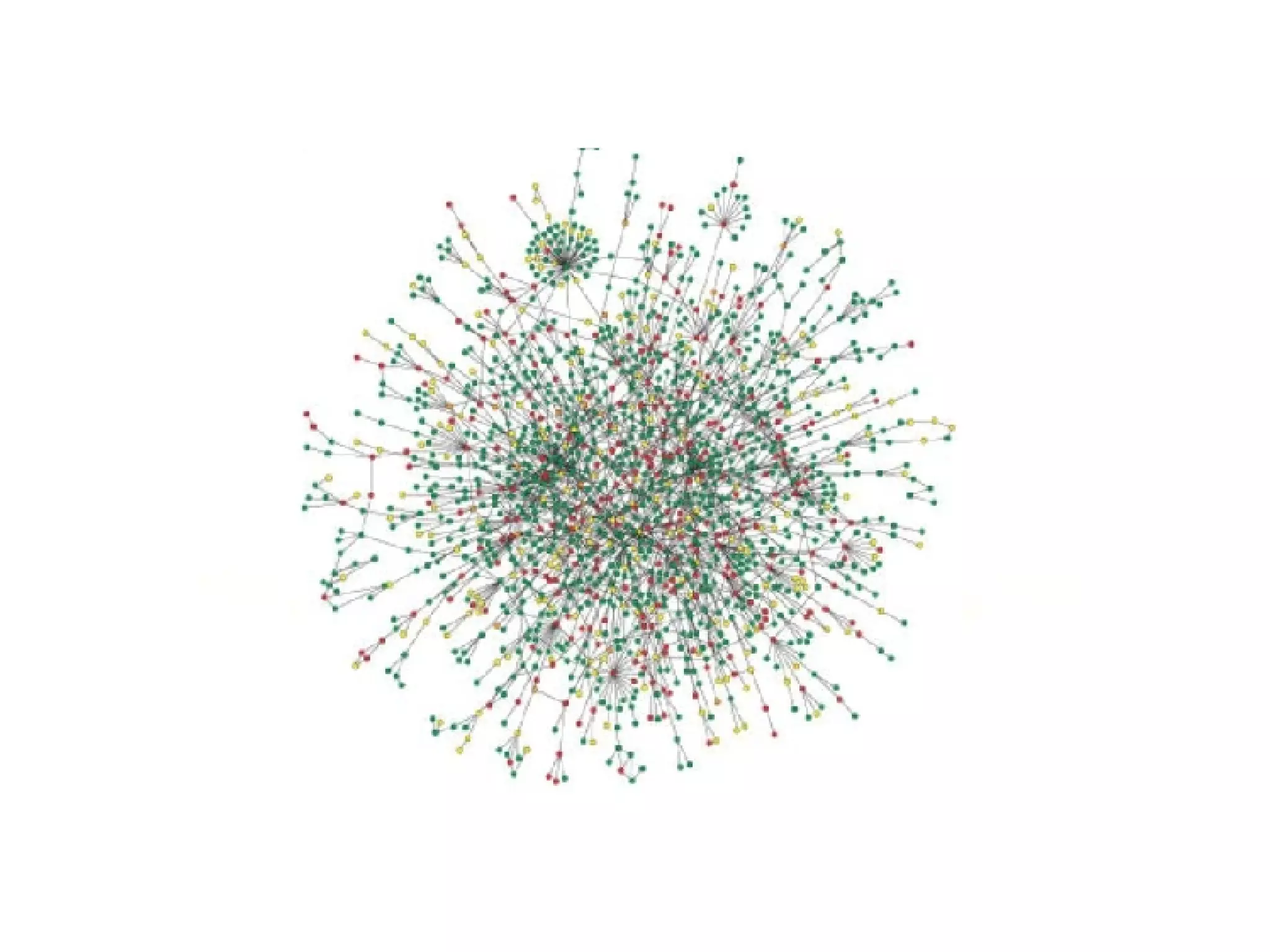
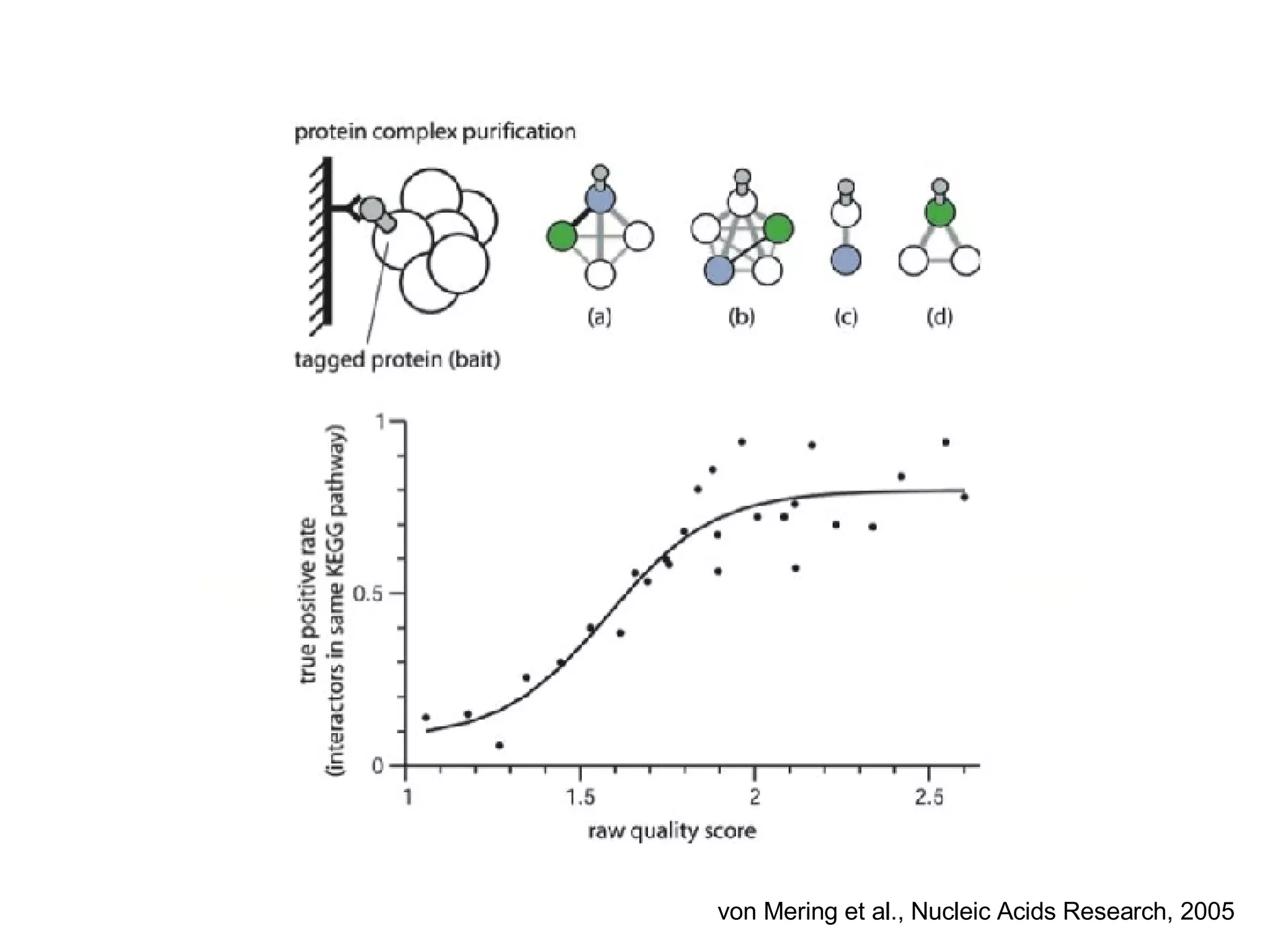
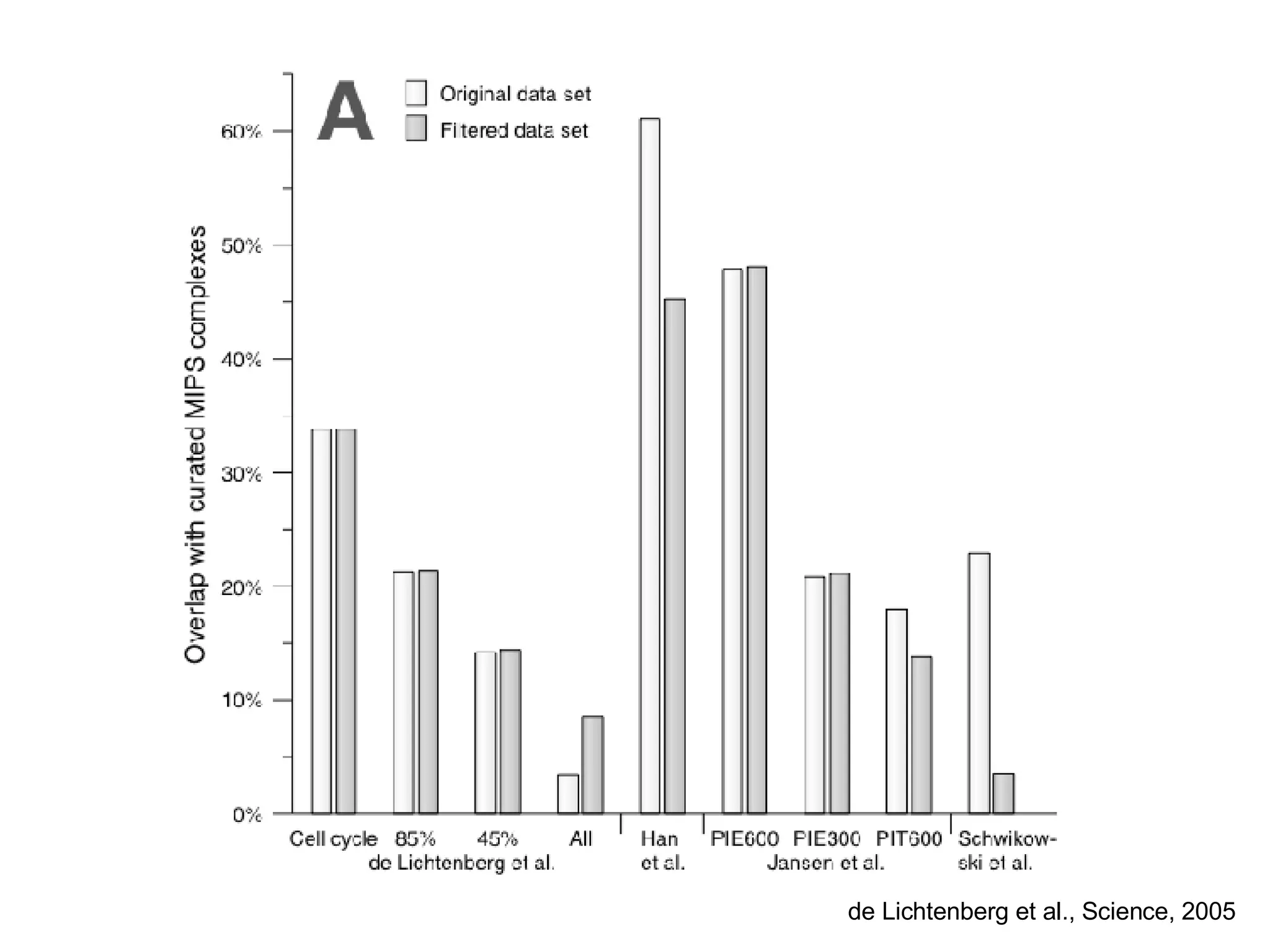
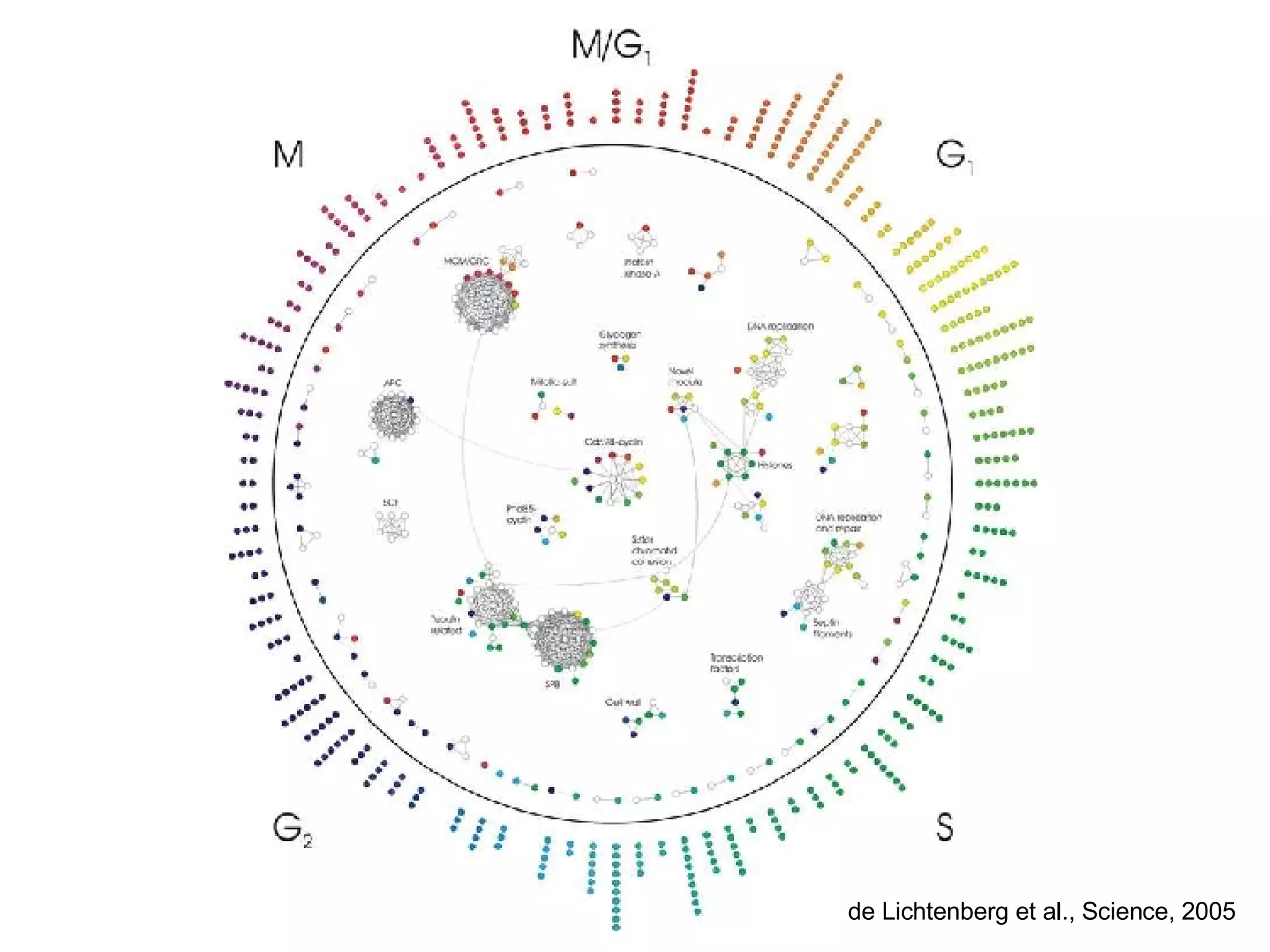
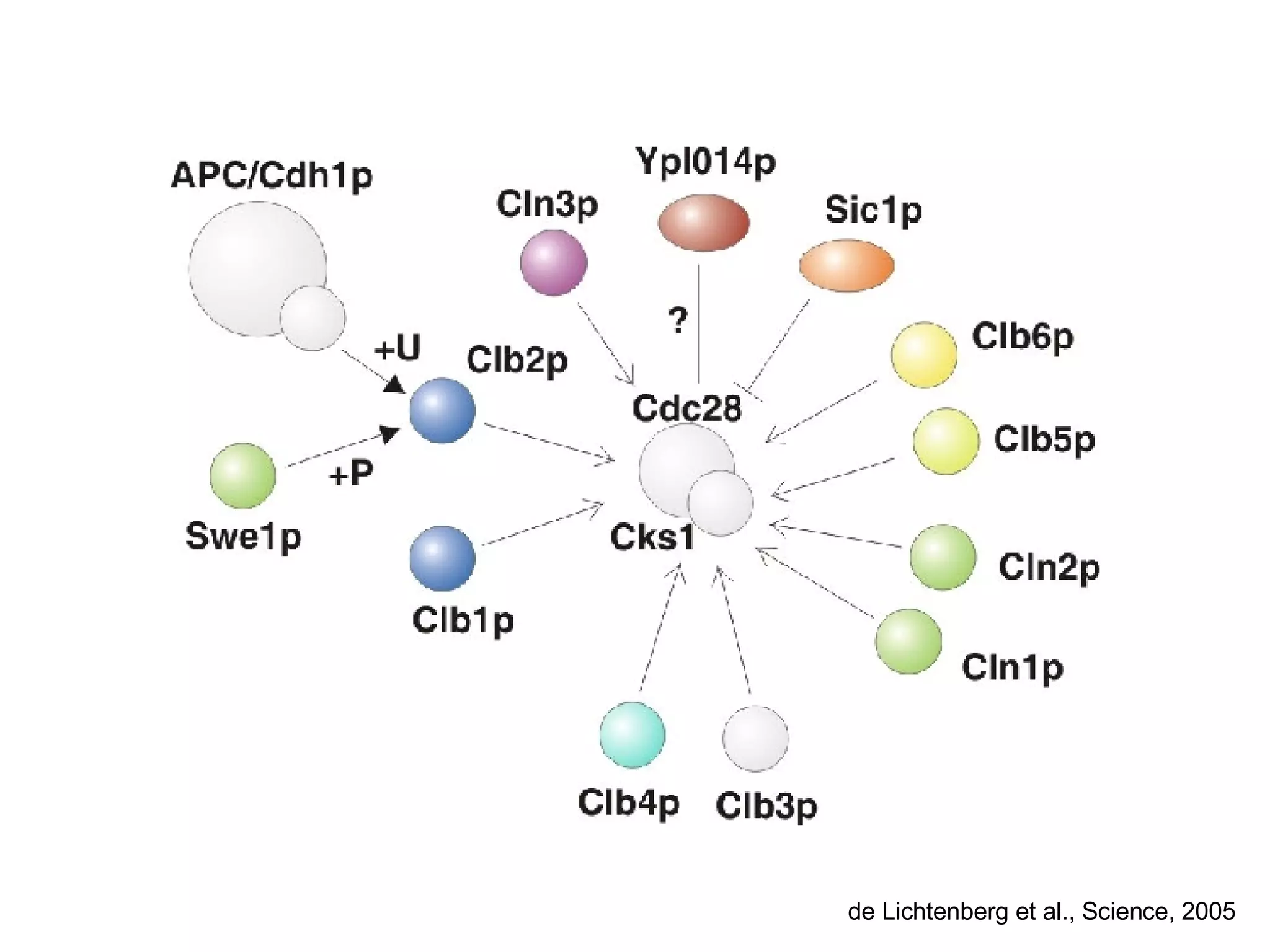
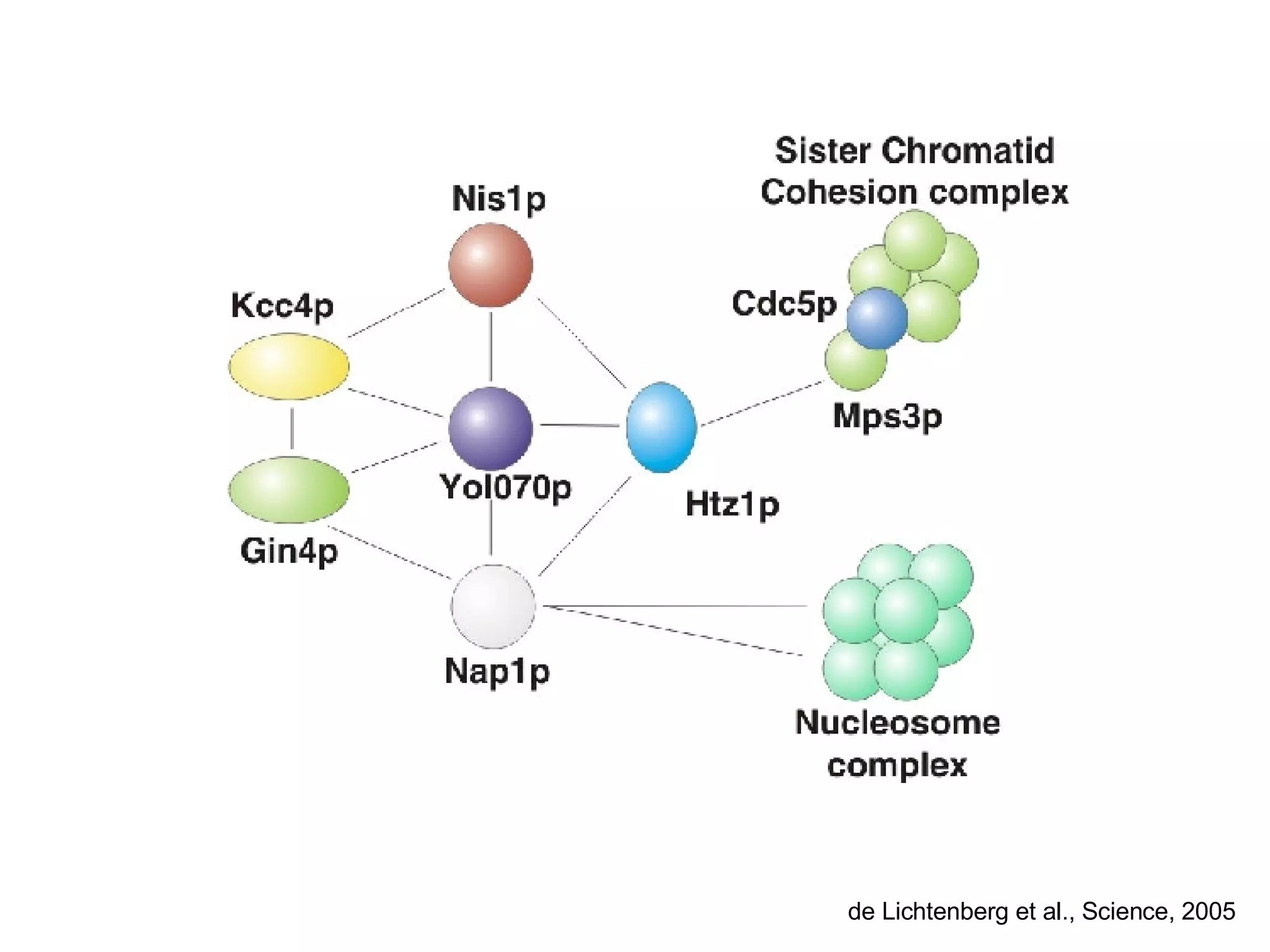
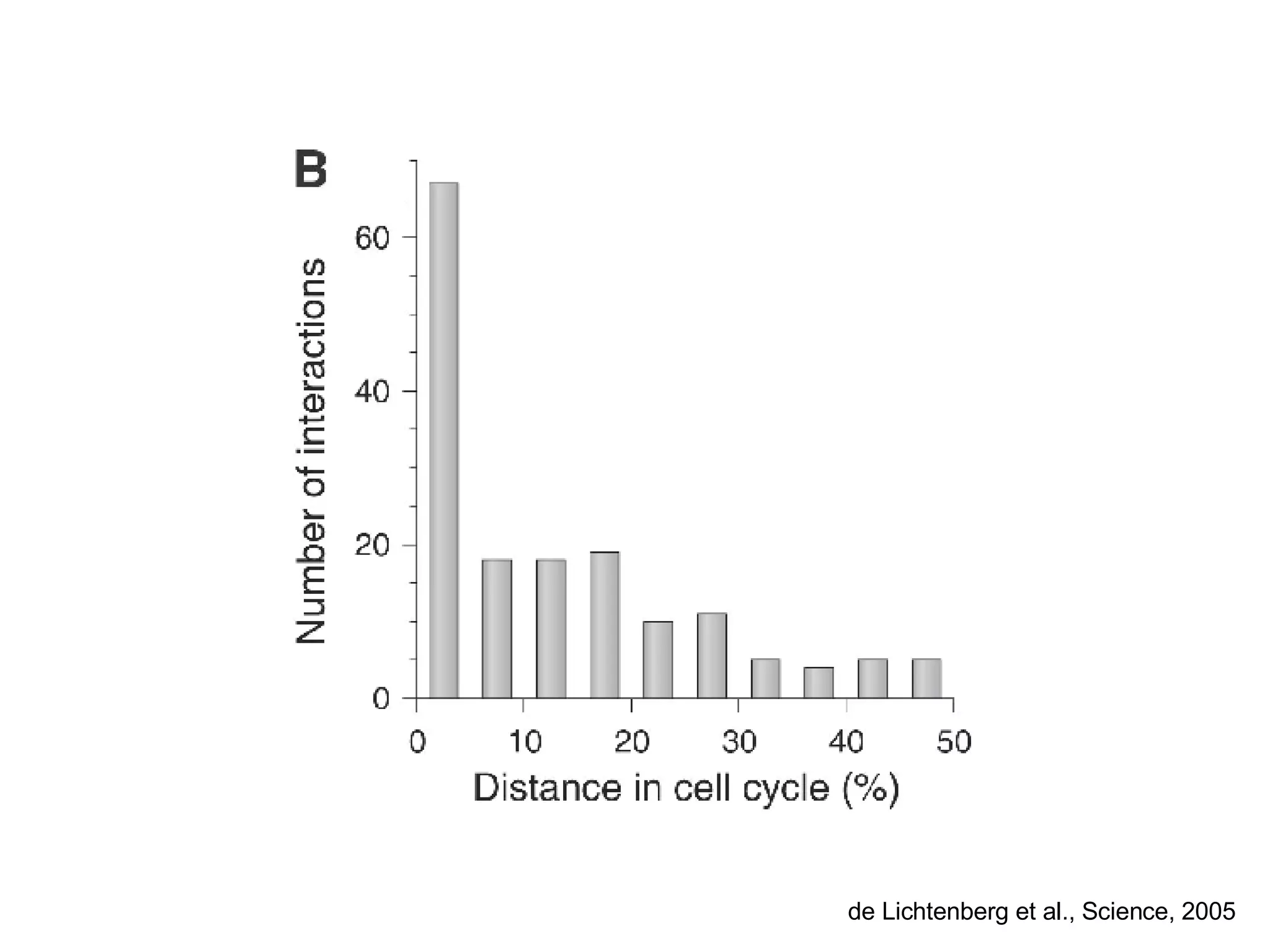
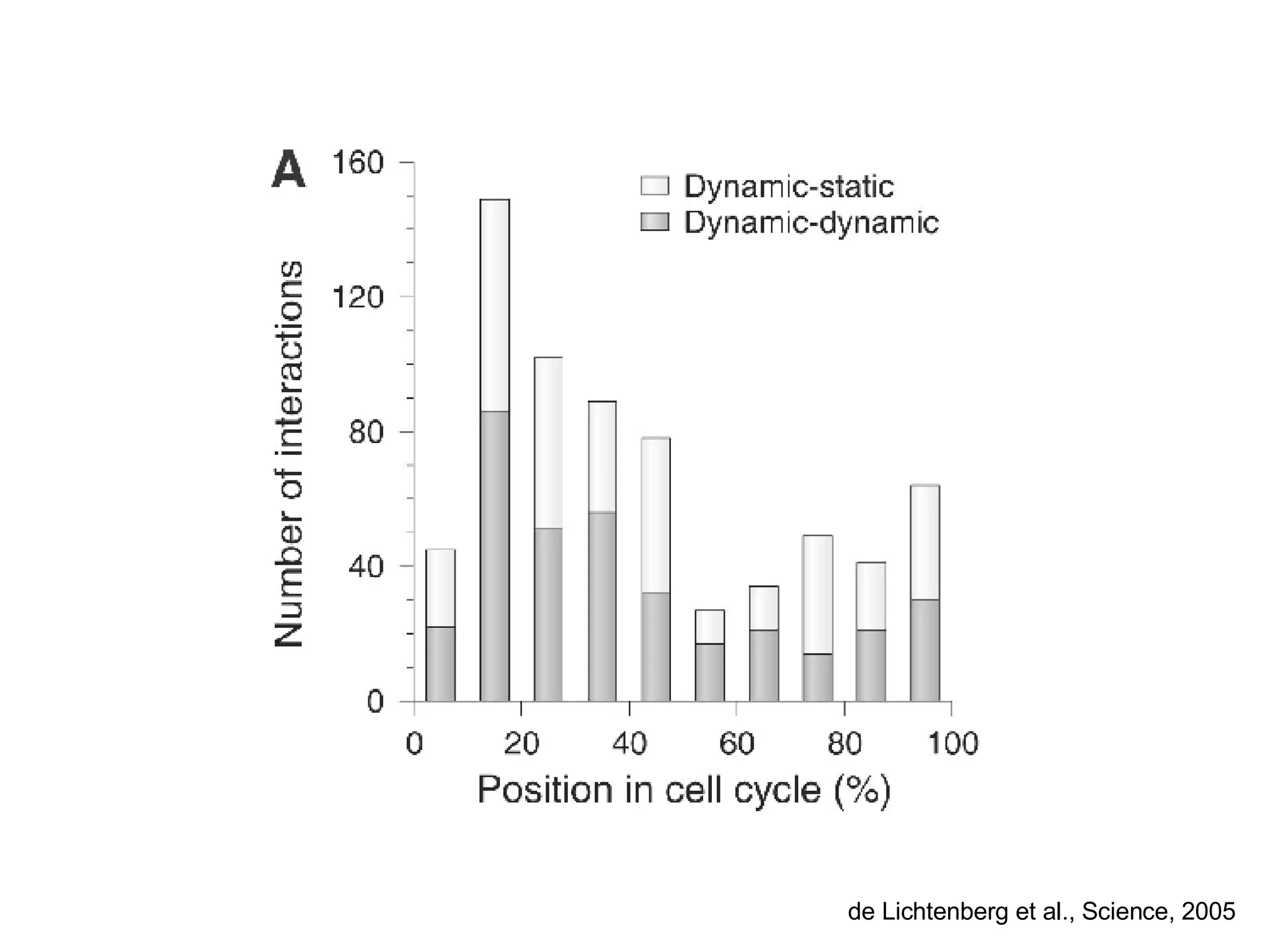
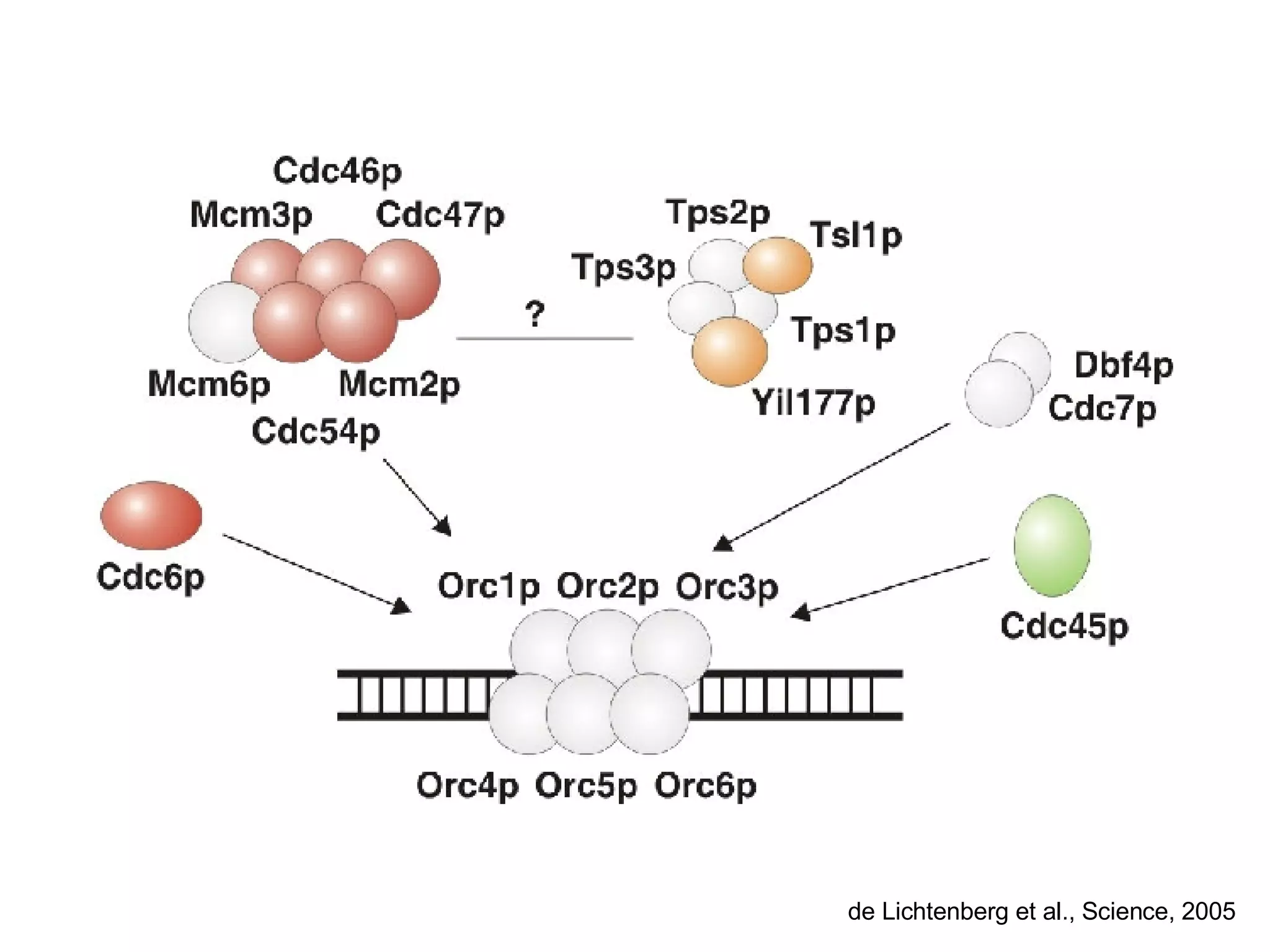
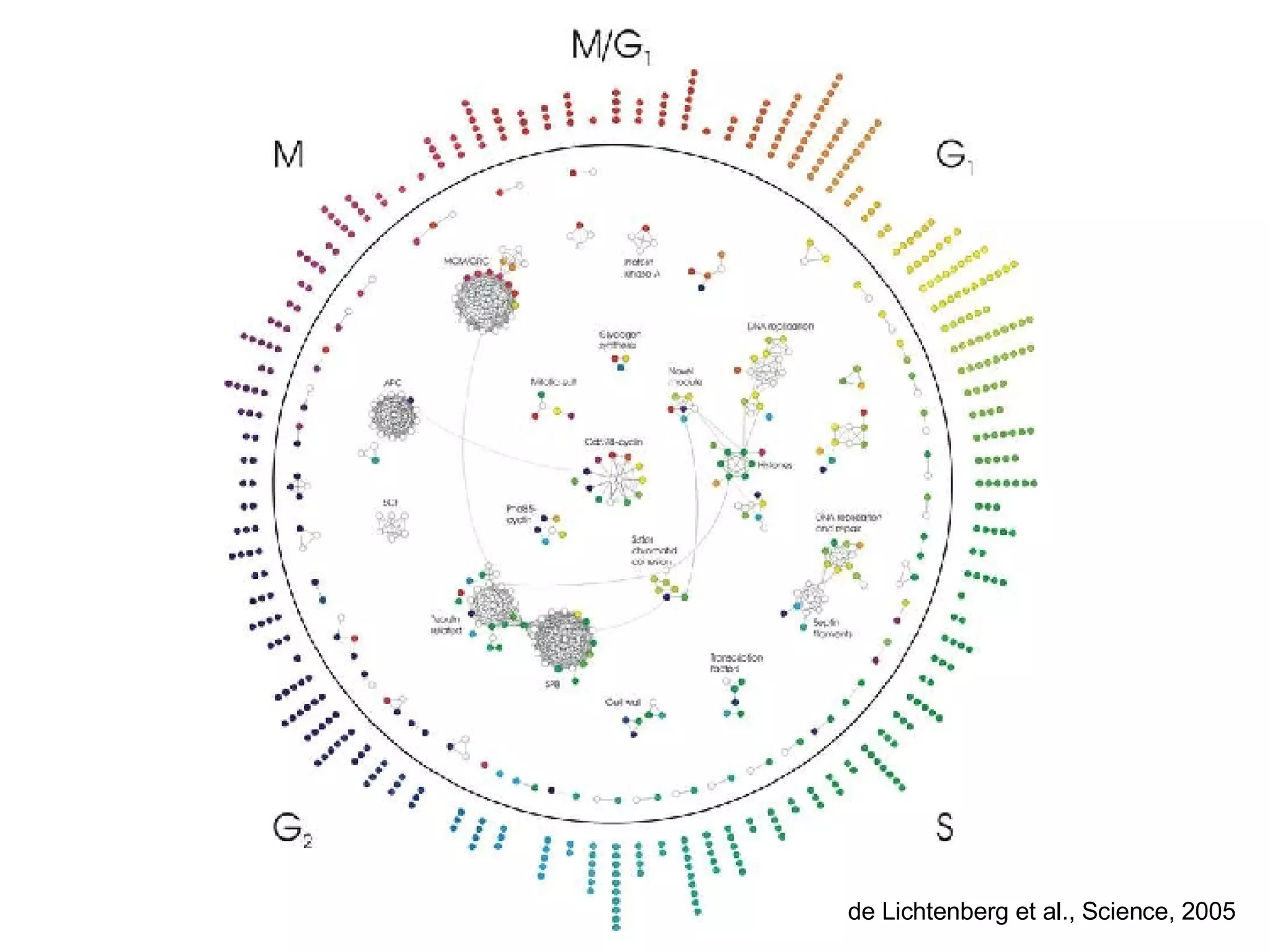
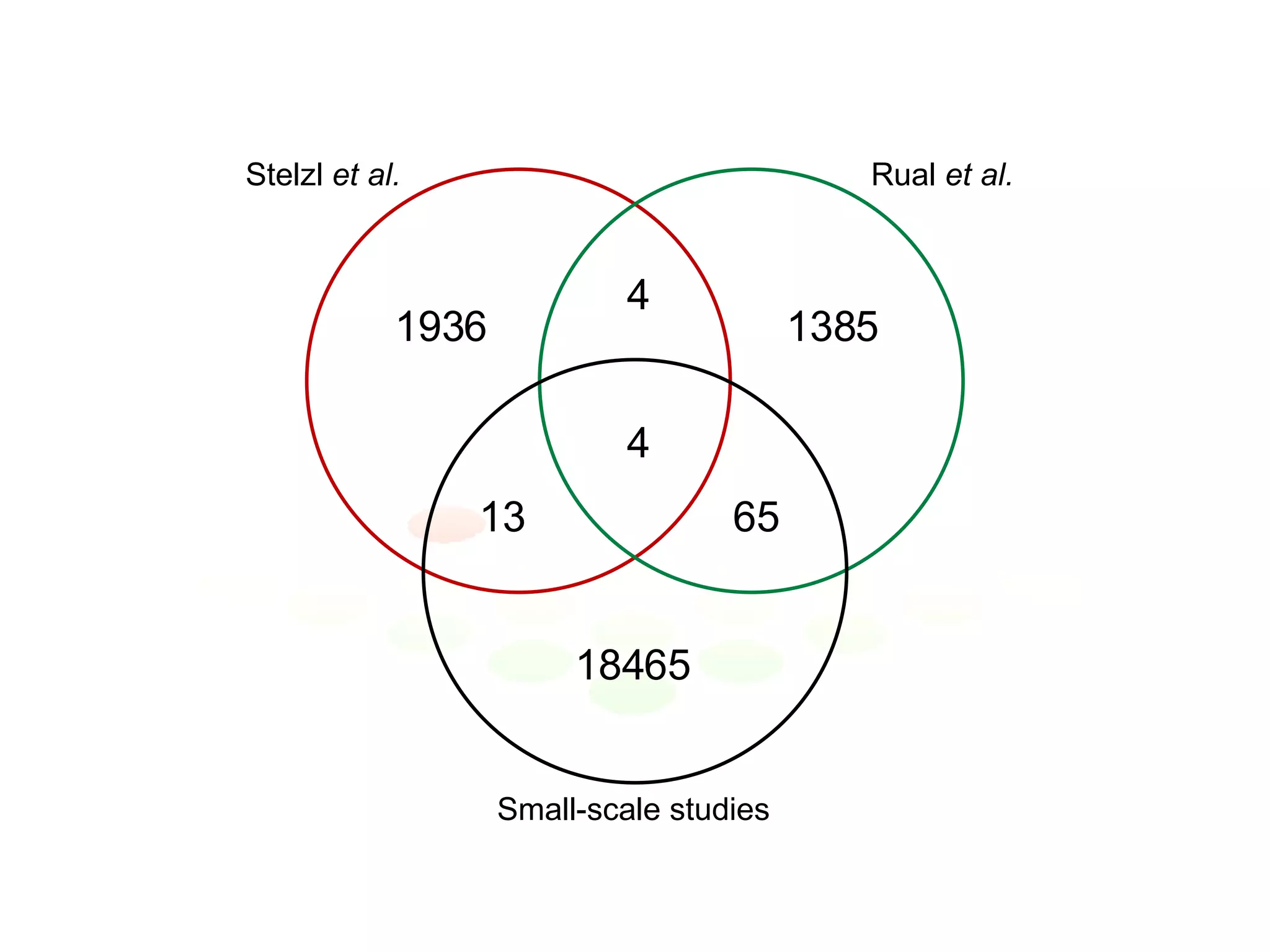
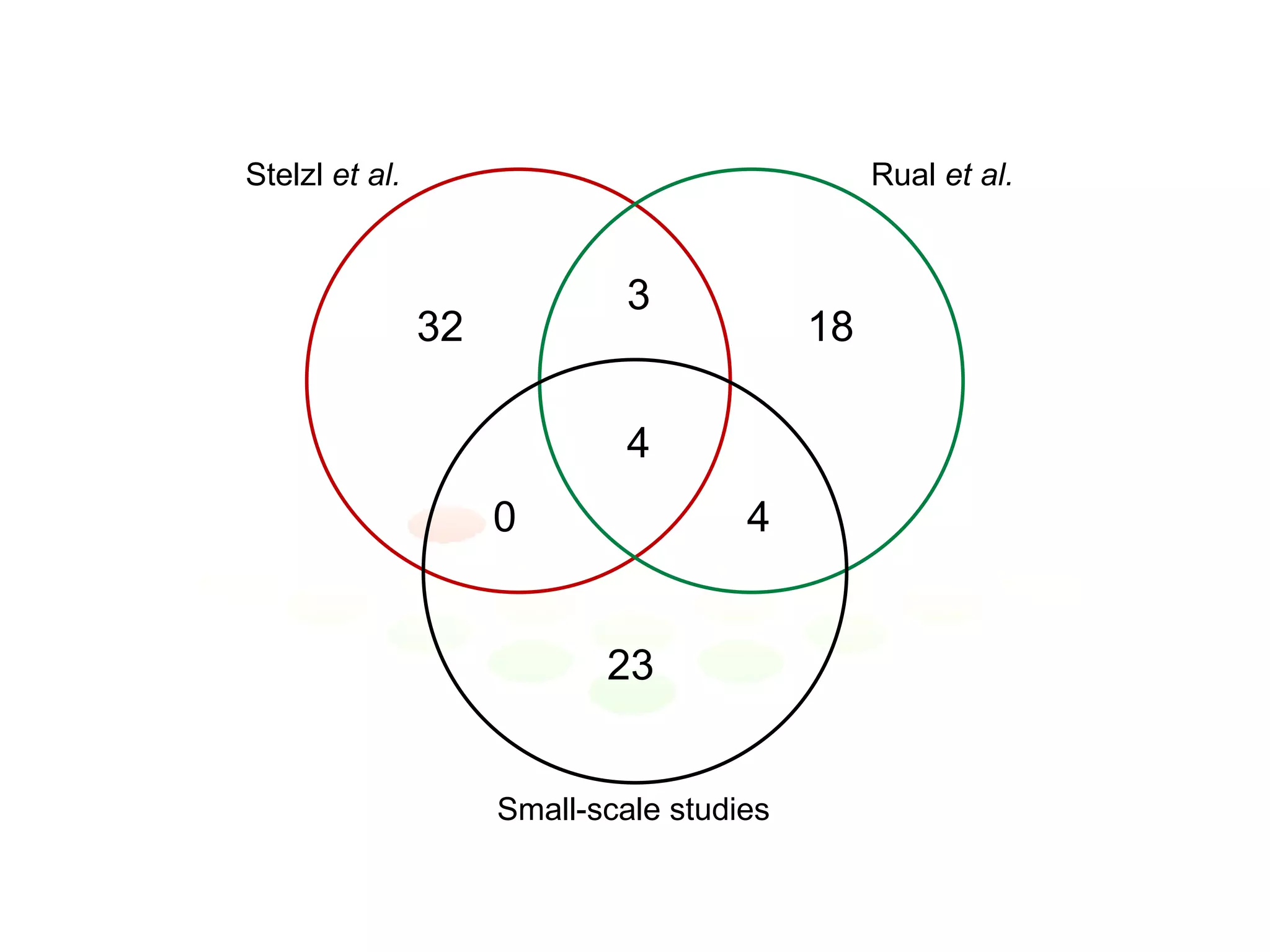
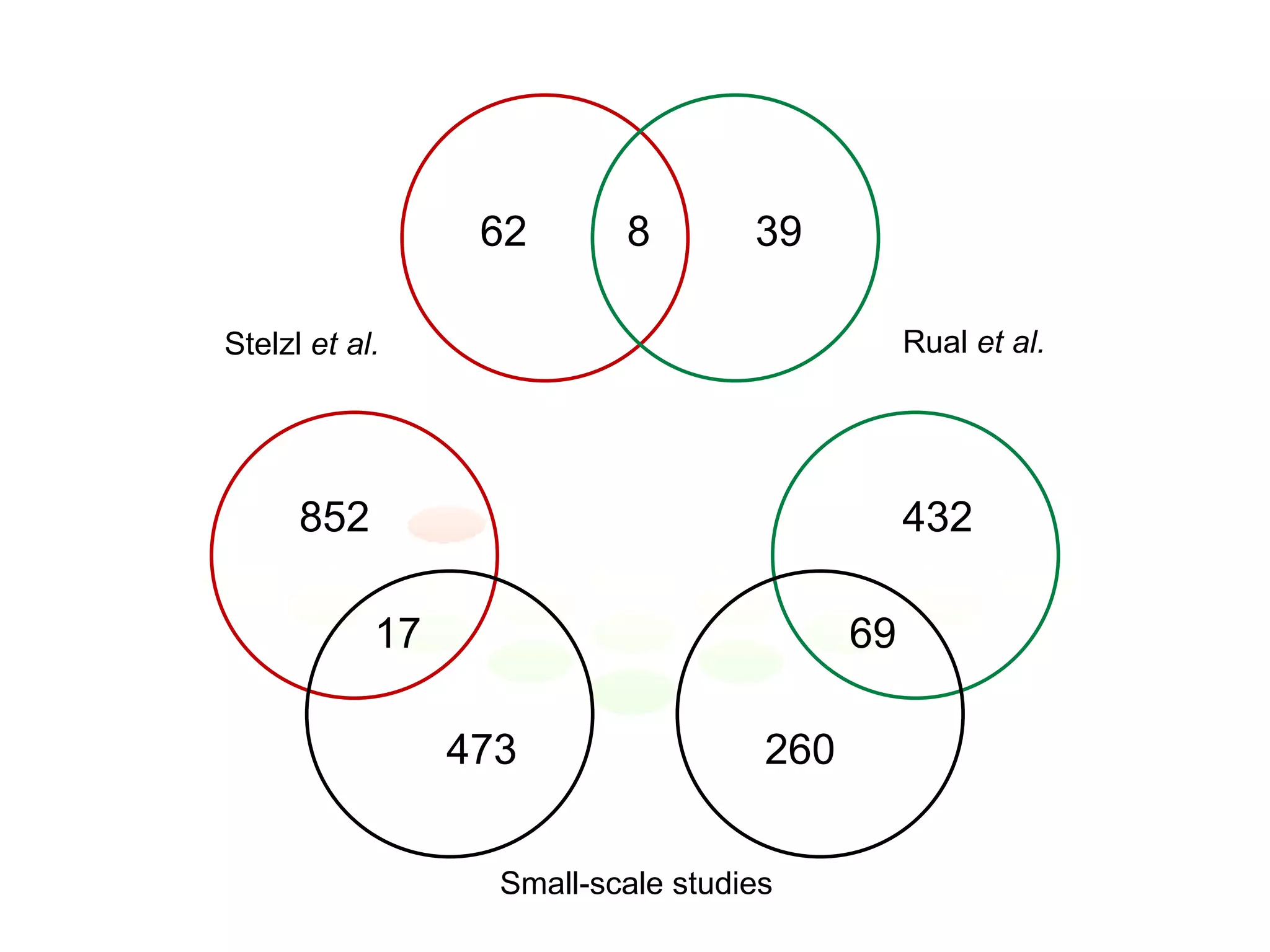
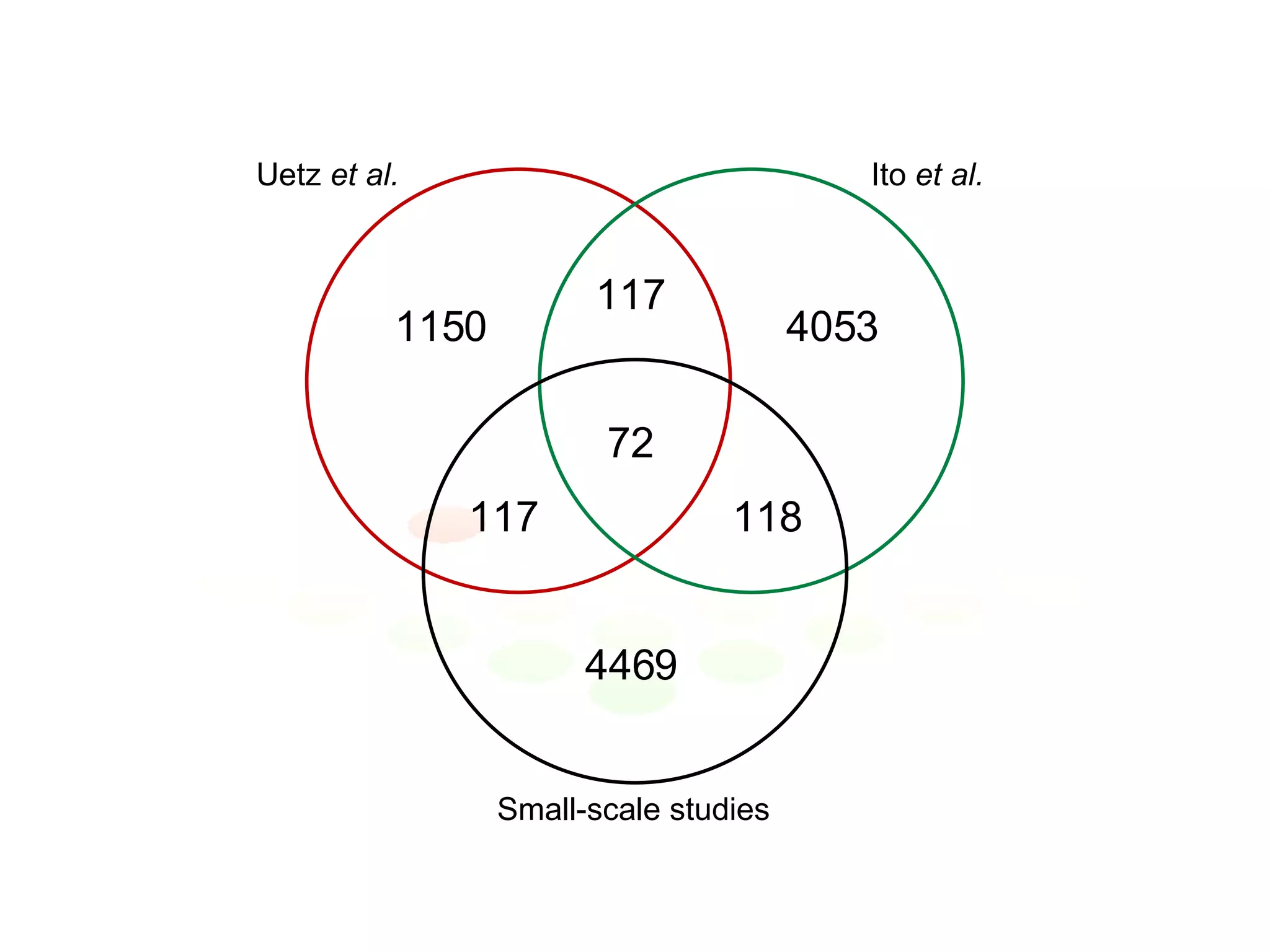
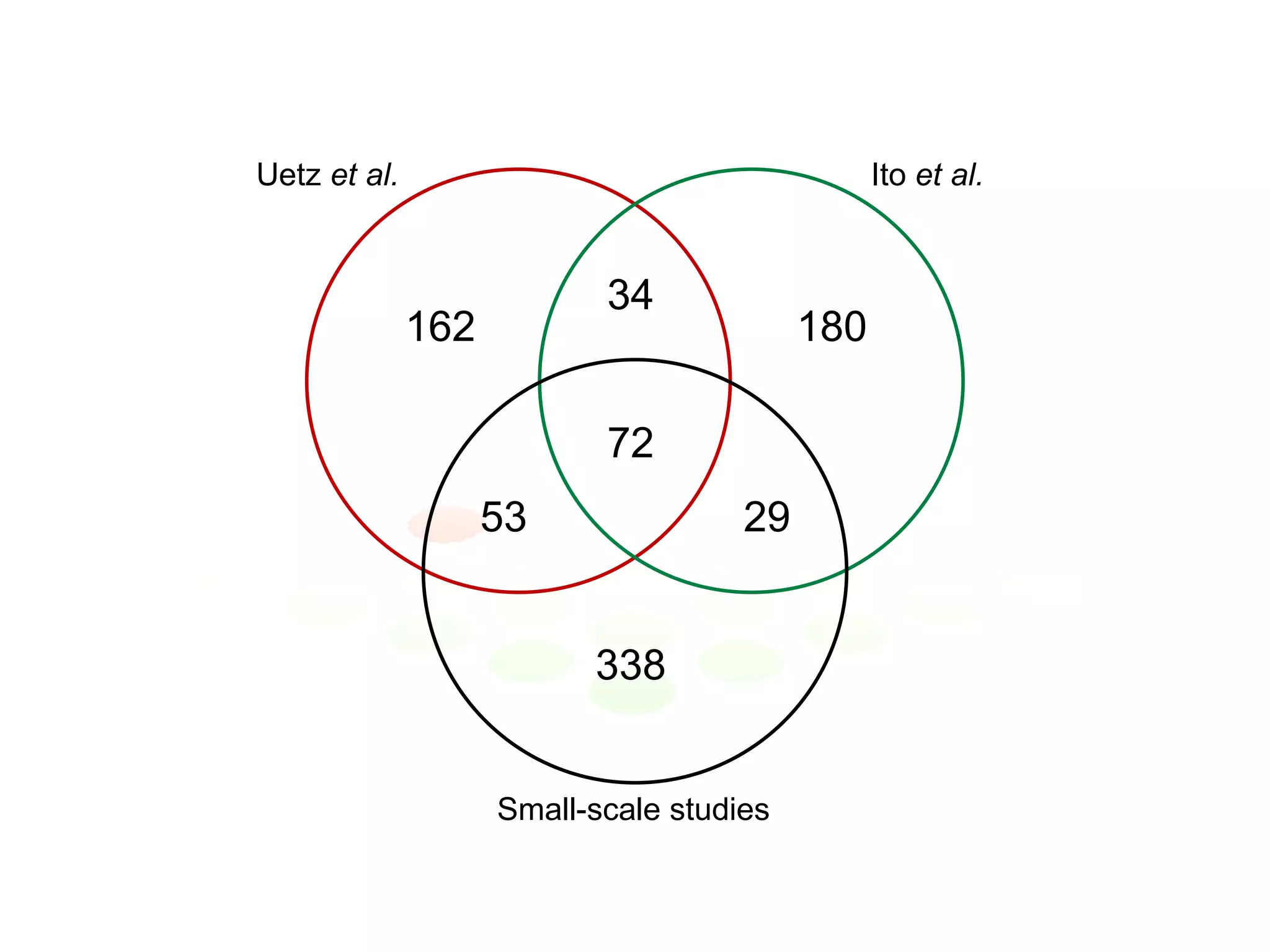
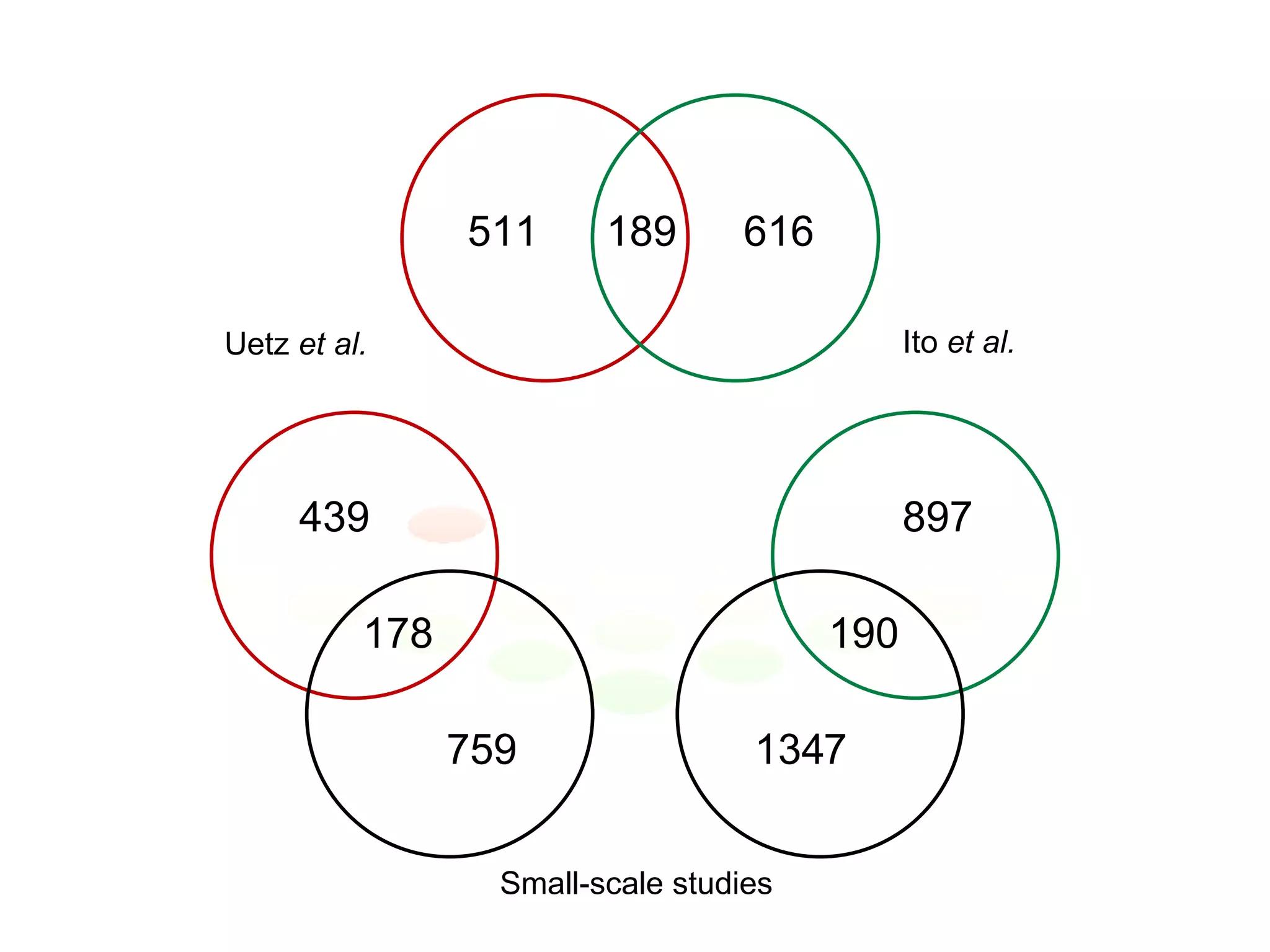
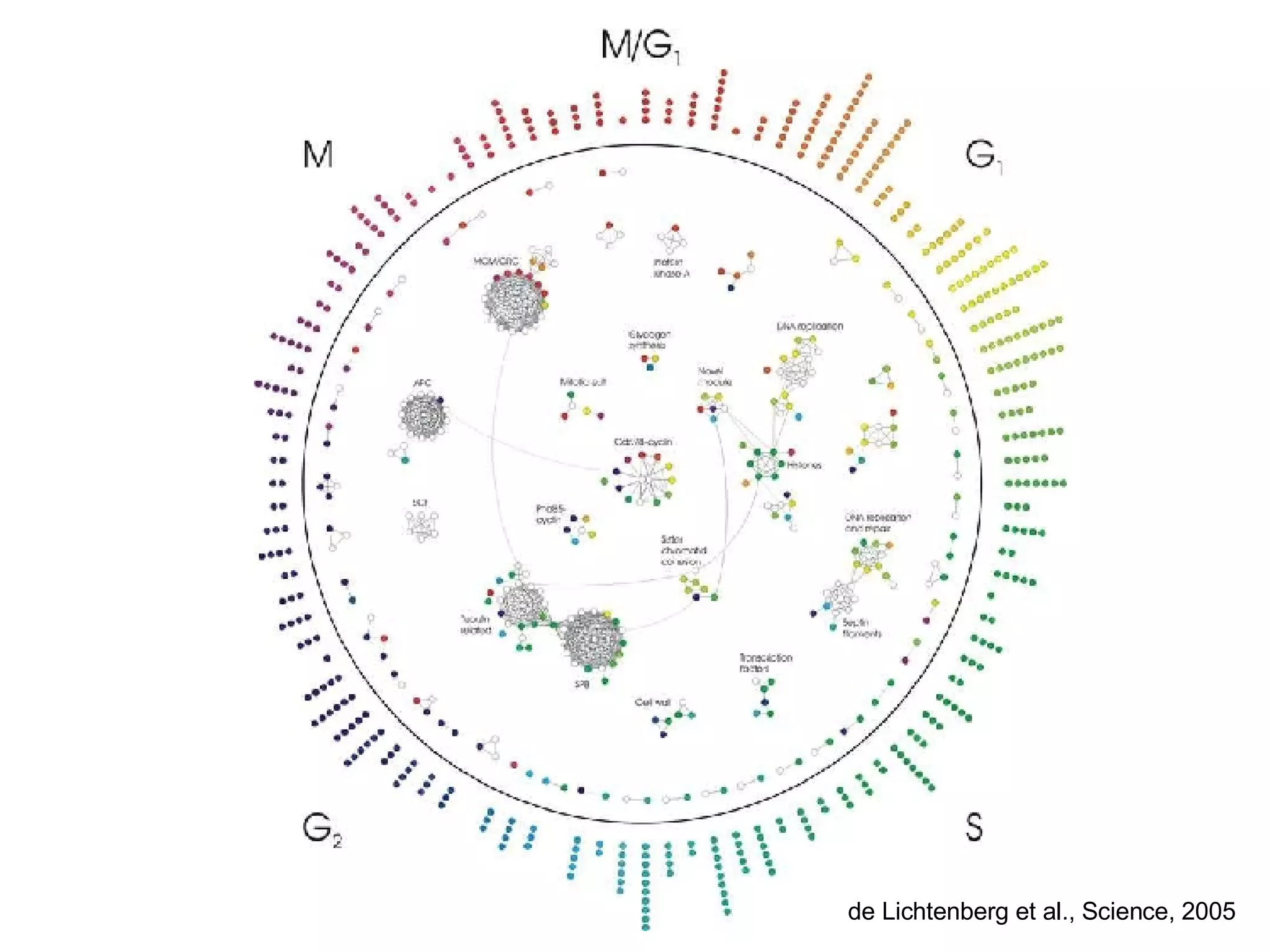
The document discusses the integration of diverse large-scale datasets to build comprehensive protein-protein interaction networks. It describes challenges with data from different sources having different identifiers, evidence types and quality. It also discusses methods used by STRING and other databases to combine data from curated databases, literature mining, primary datasets and transfer of interactions based on orthology. Examples are given of cell cycle studies in yeast that have analyzed periodically expressed genes and protein interactions.












































![Gene and protein names Cue words for entity recognition Verbs for relation extraction [ nxgene The GAL4 gene ] [ nxexpr T he expression of [ nxgene the cytochrome genes [ nxpg CYC1 and CYC7 ]]] is controlled by [ nxpg HAP1 ]](https://image.slidesharecdn.com/jen05talk10-1217969276804105-9/75/Integration-of-diverse-large-scale-datasets-56-2048.jpg)