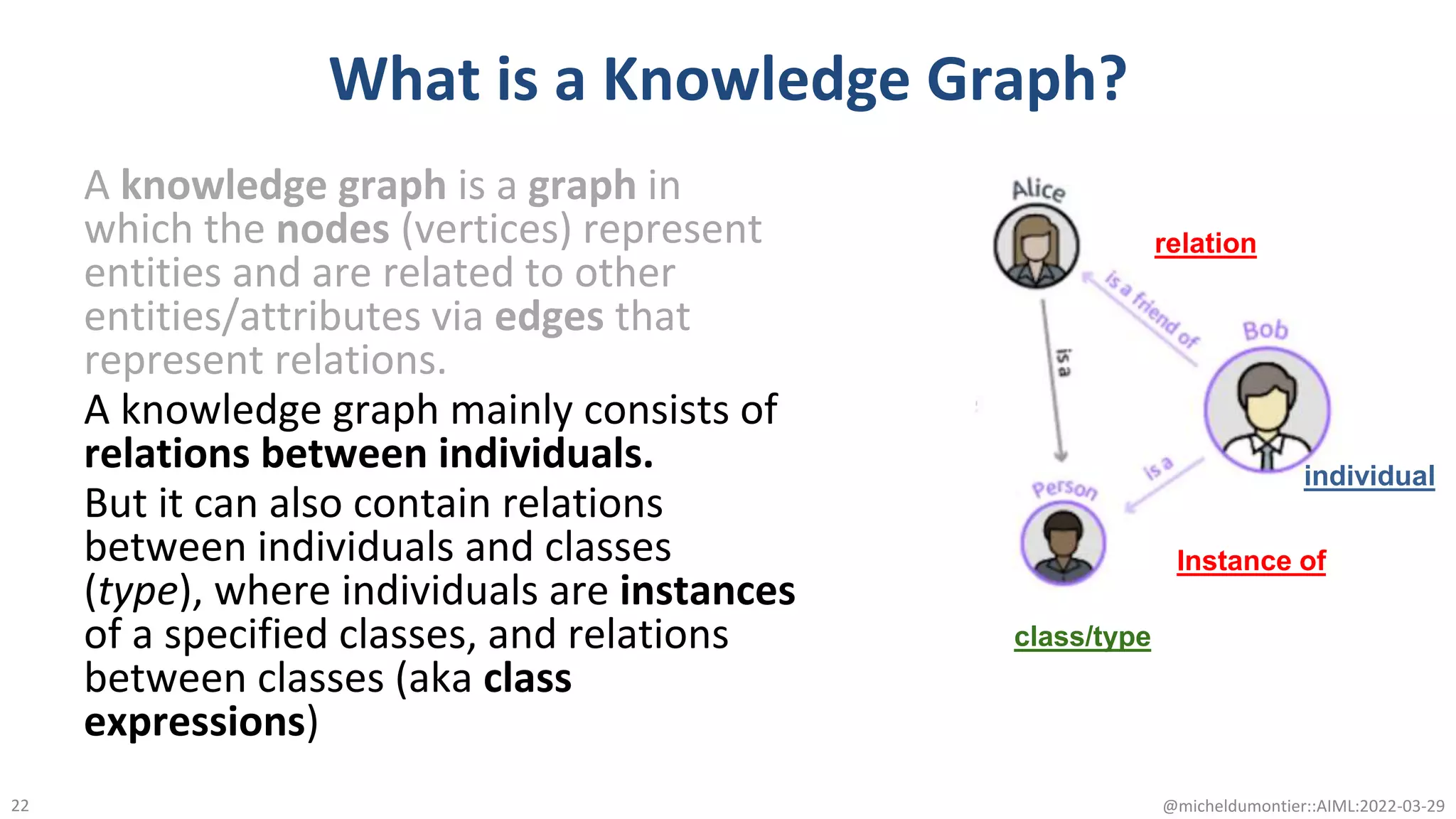
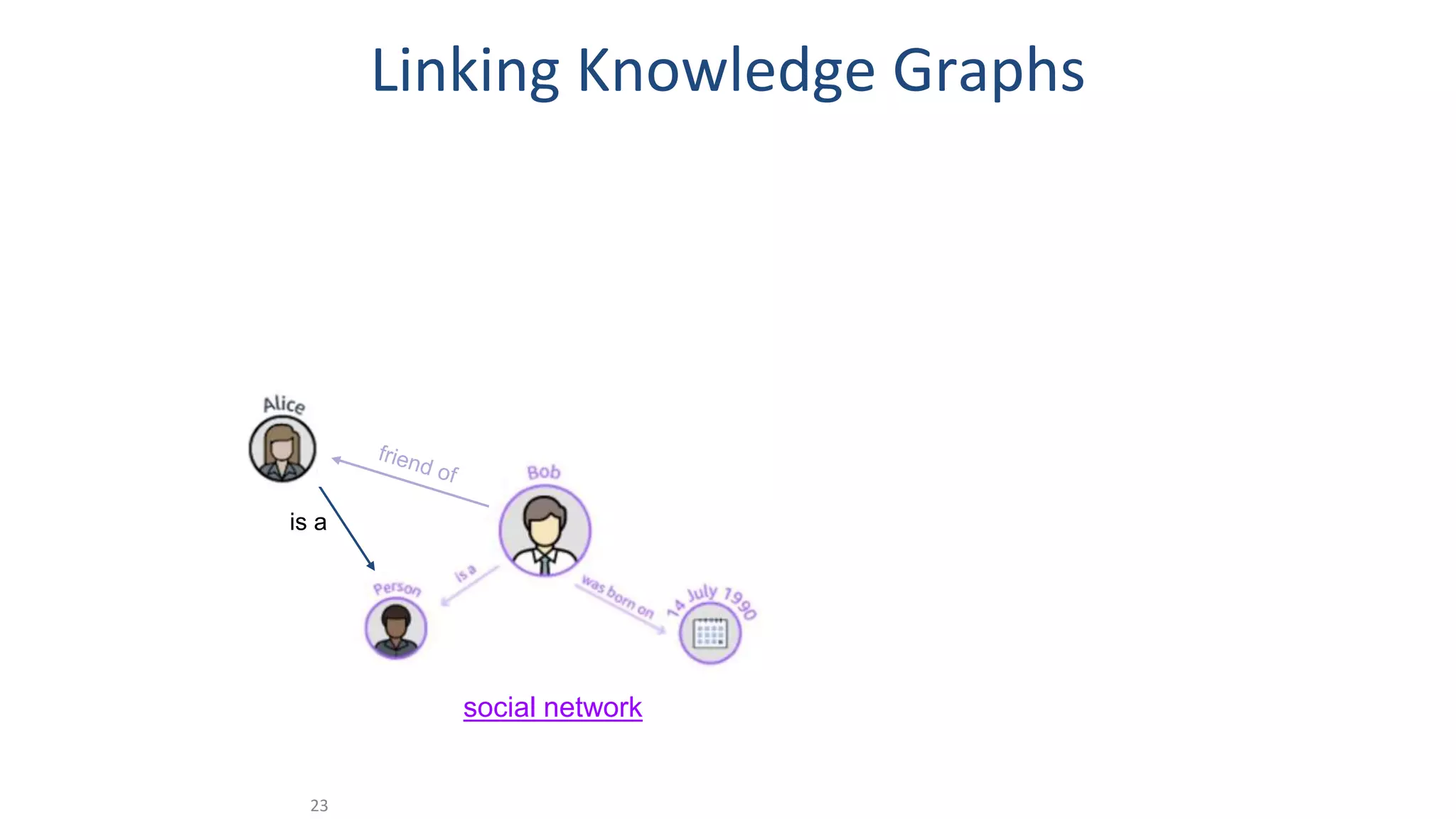
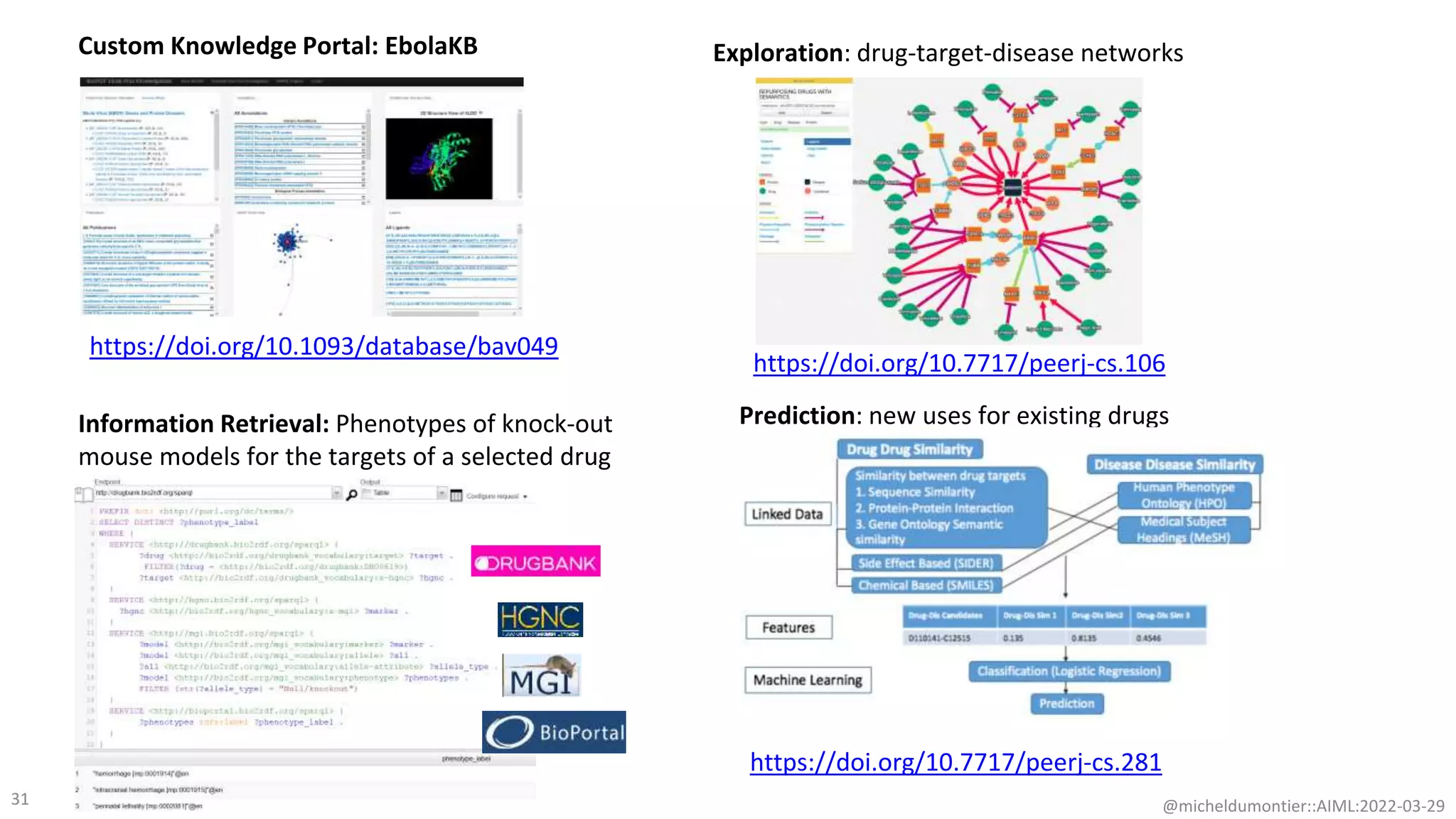
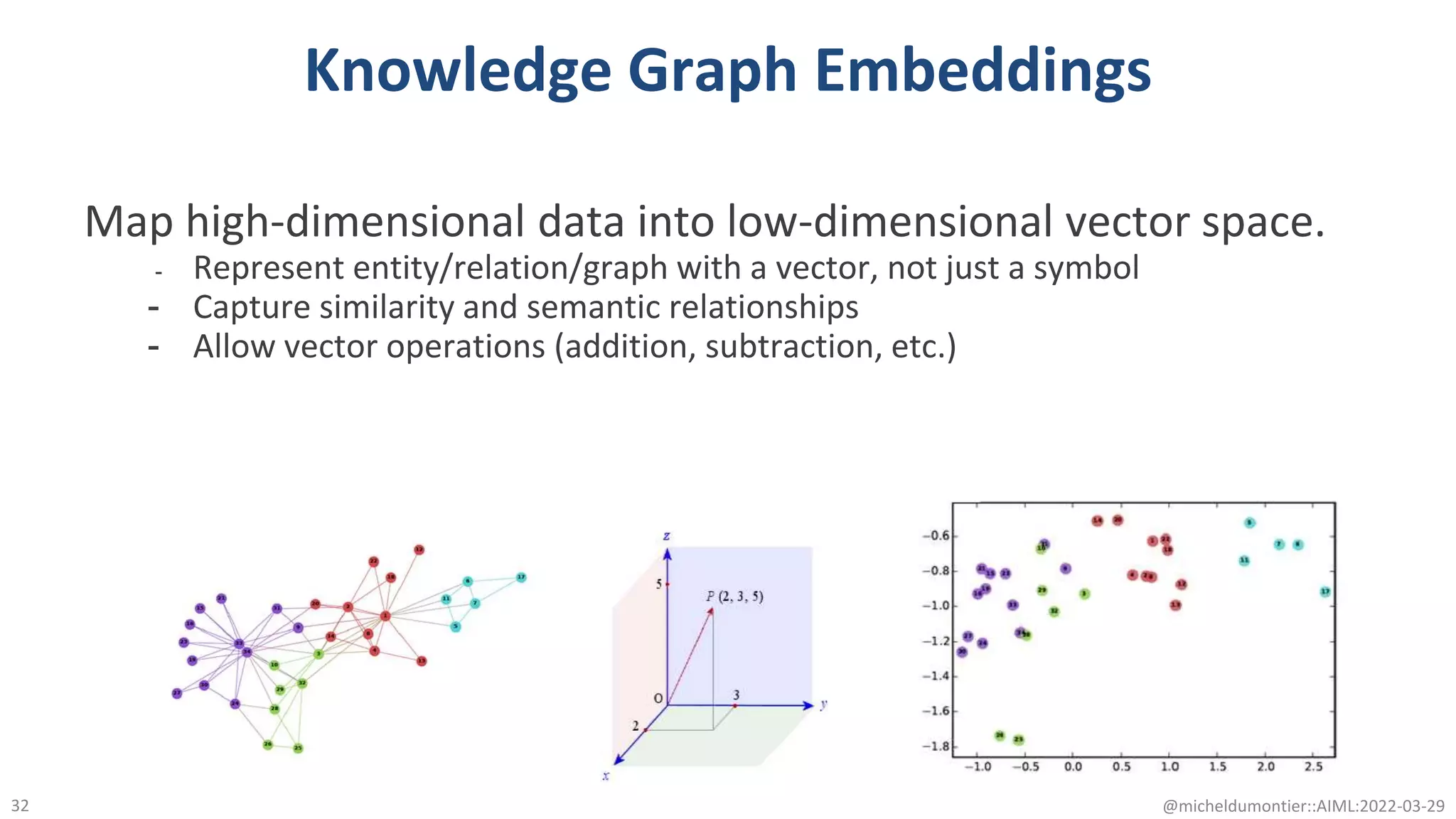
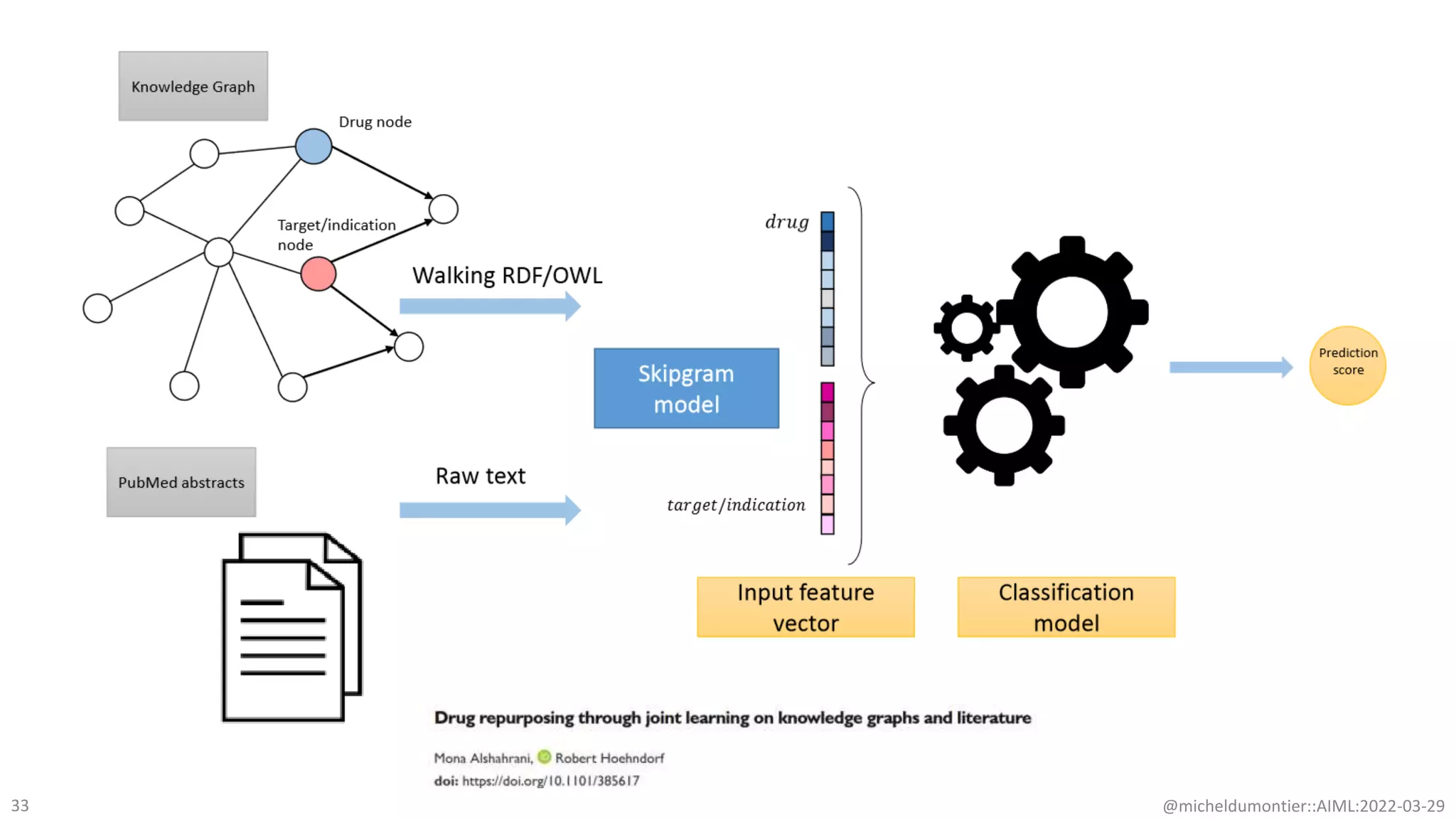
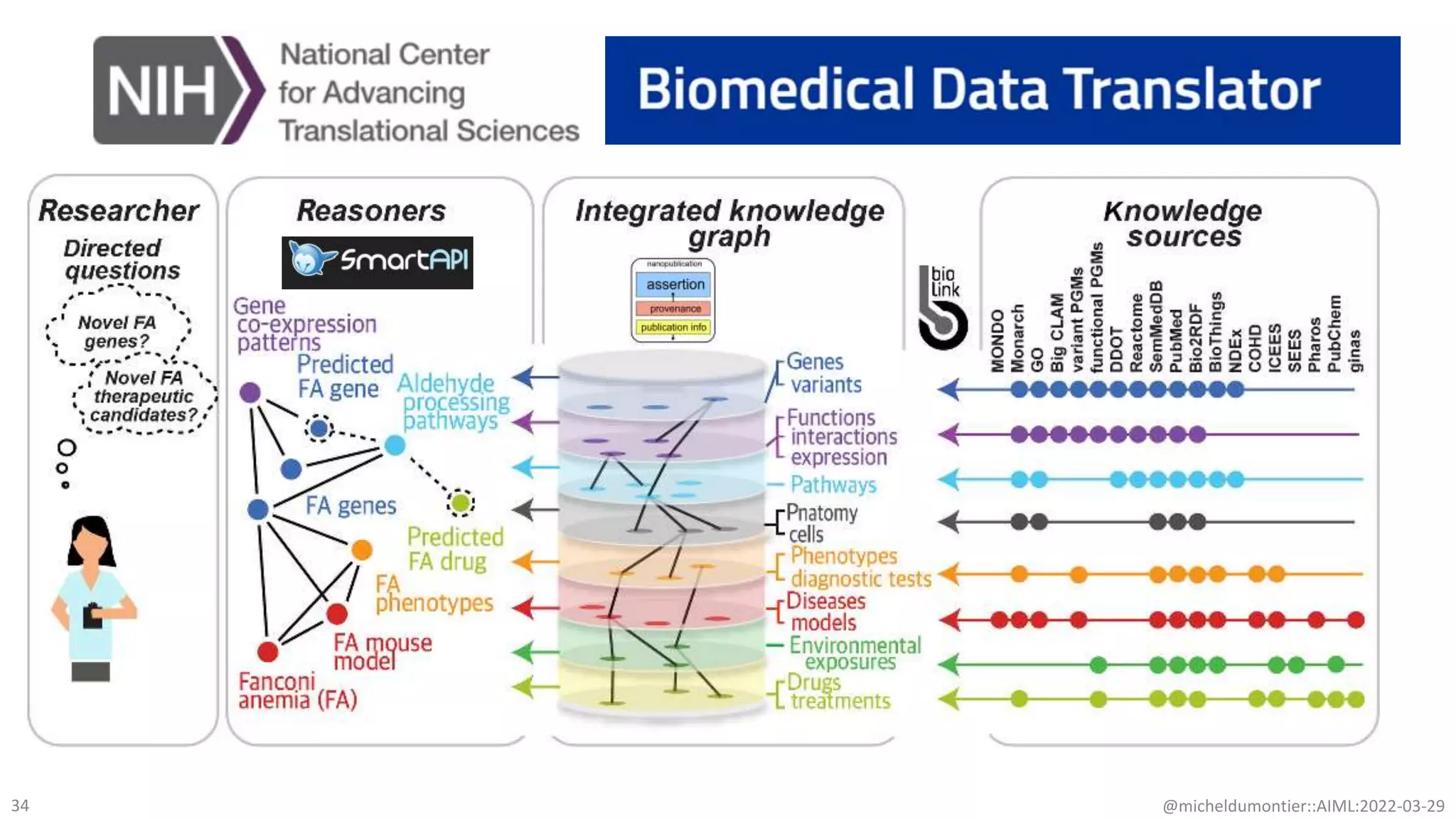
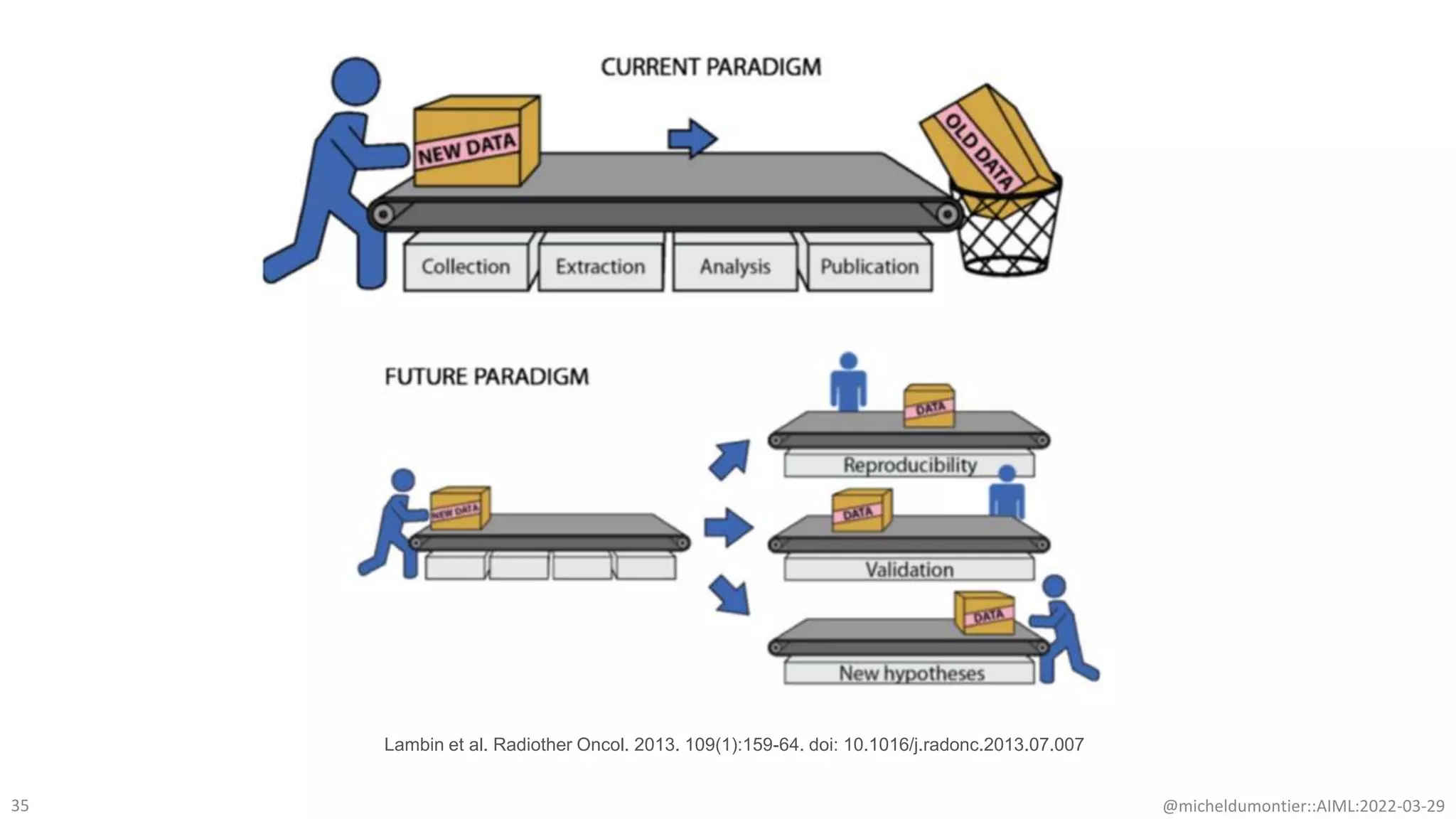
The document discusses the potential of data-driven discovery science facilitated by FAIR (Findable, Accessible, Interoperable, Reusable) knowledge graphs, emphasizing the importance of high-quality metadata and community standards for effective data reuse. It highlights challenges such as low reproducibility in research and the need for improved machine-human collaboration to enhance data discovery. The text advocates for a new paradigm in research that leverages existing data to accelerate discoveries in biomedicine and other fields.