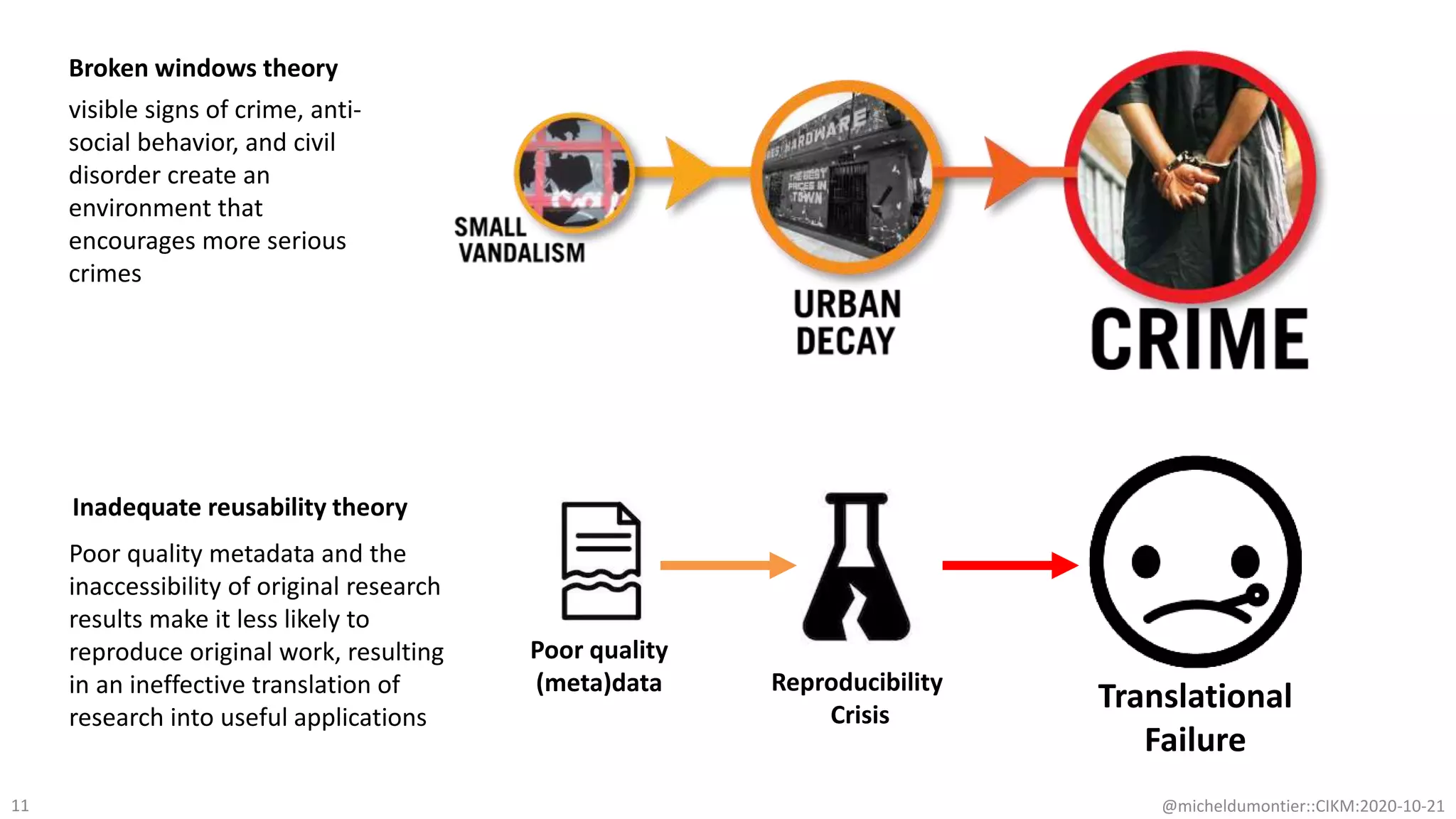
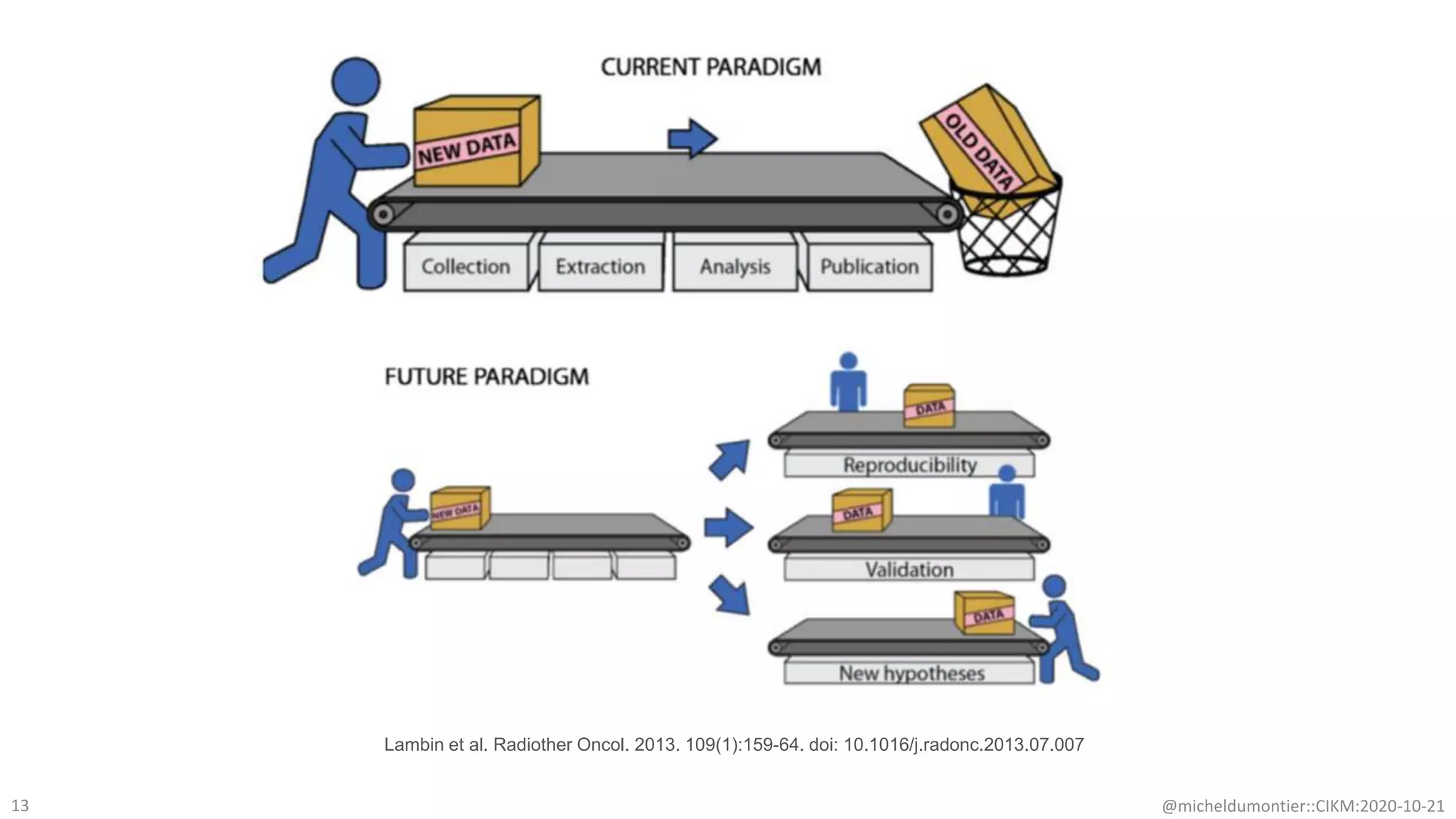
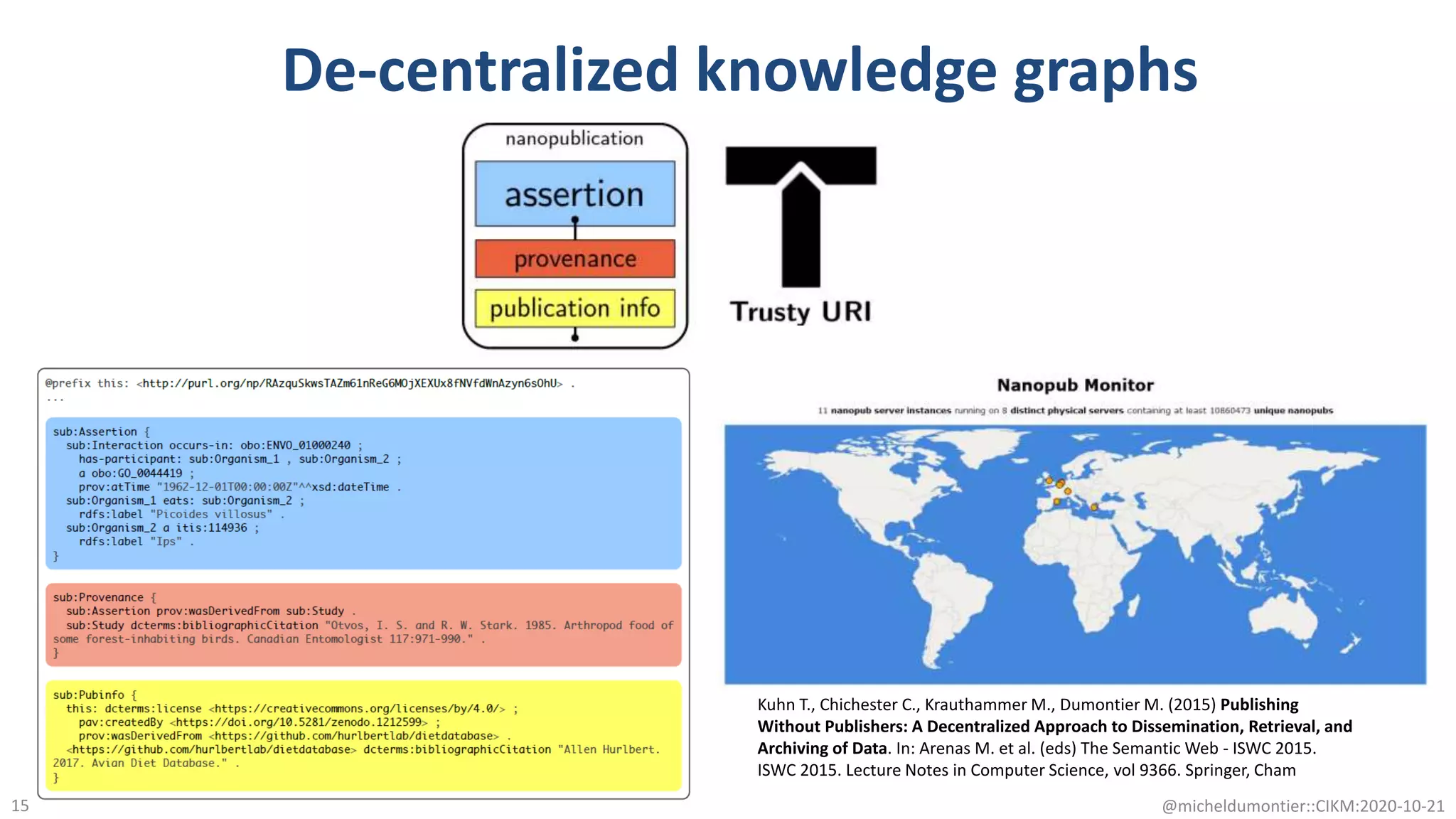
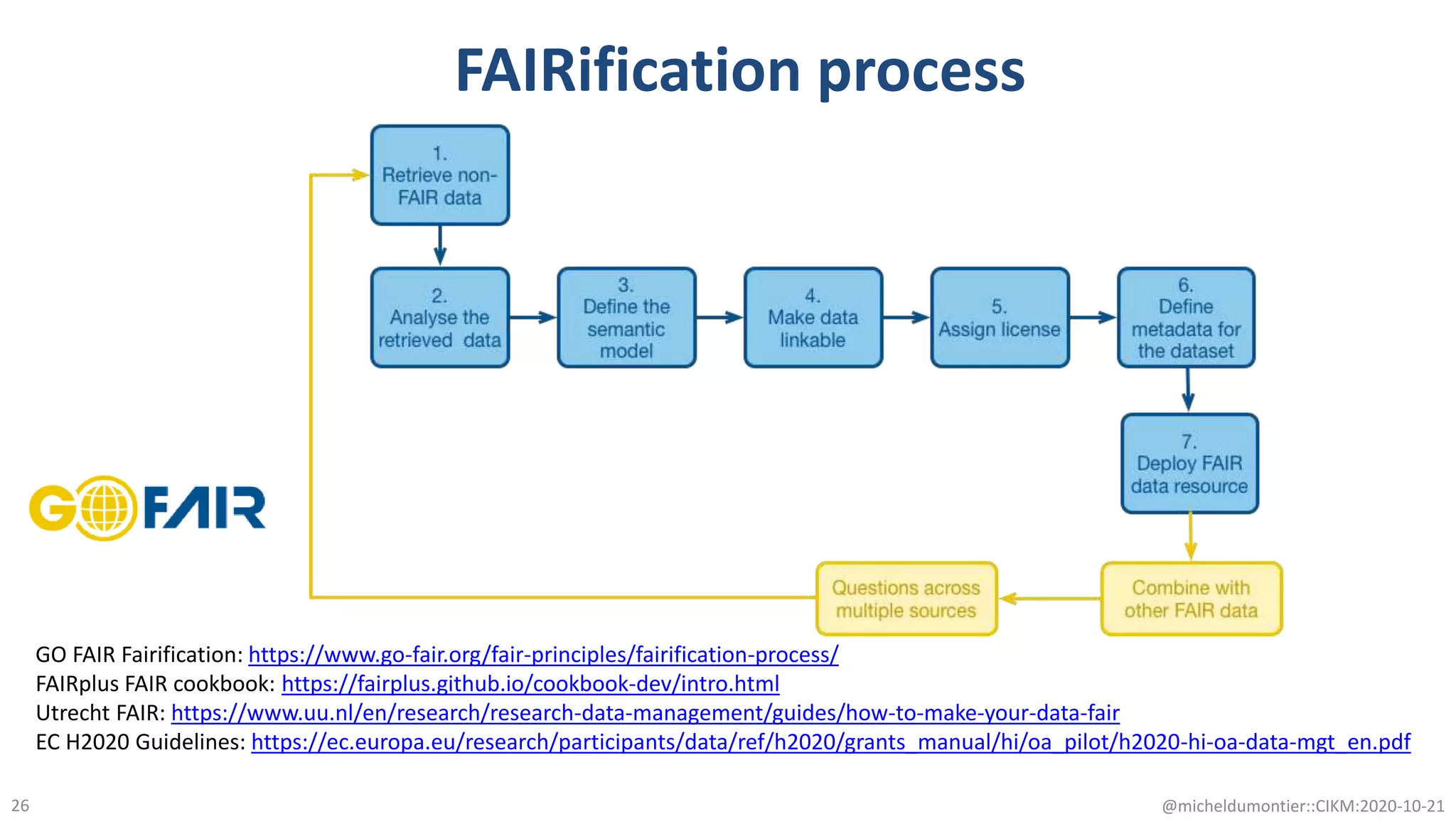
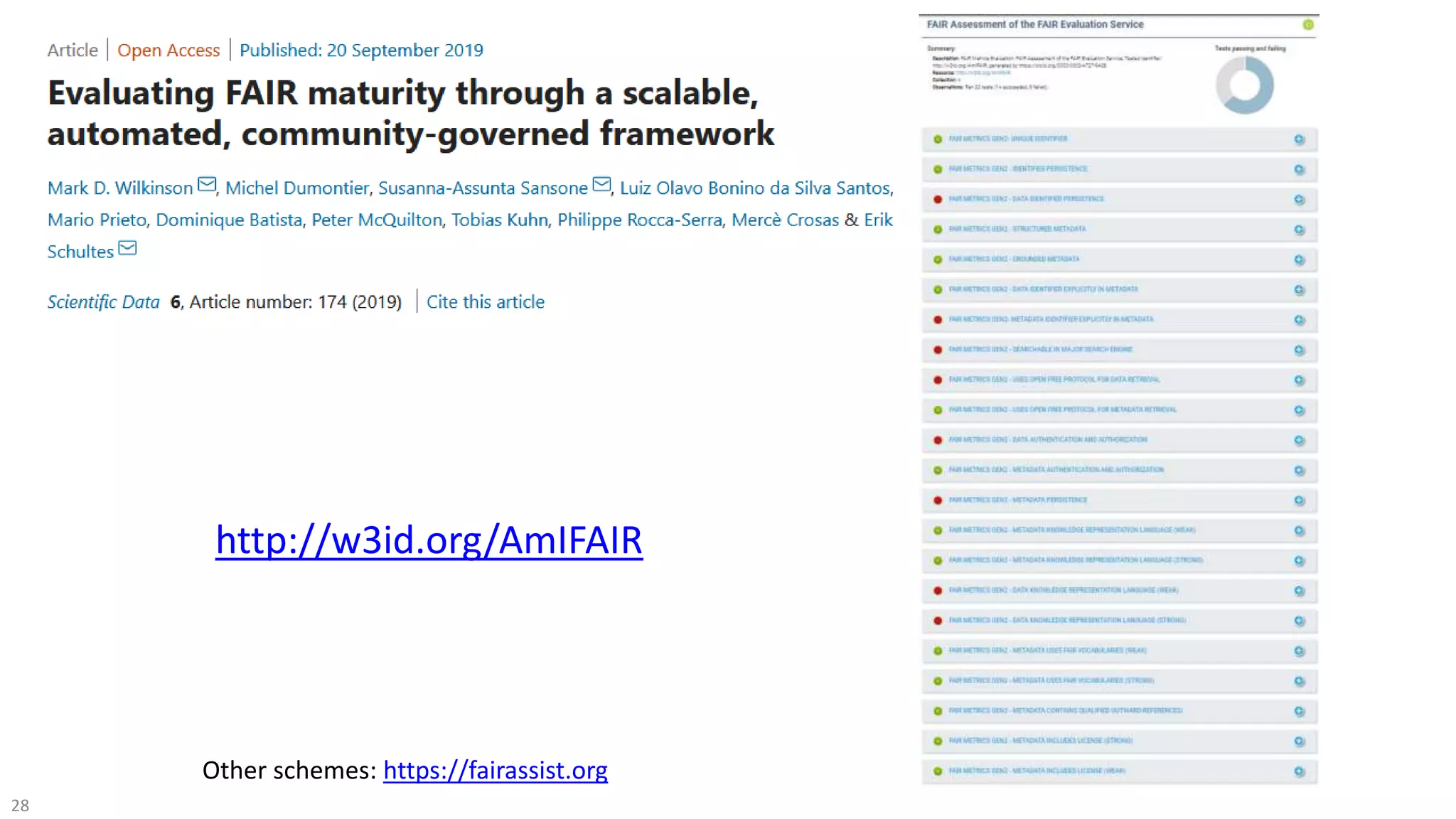
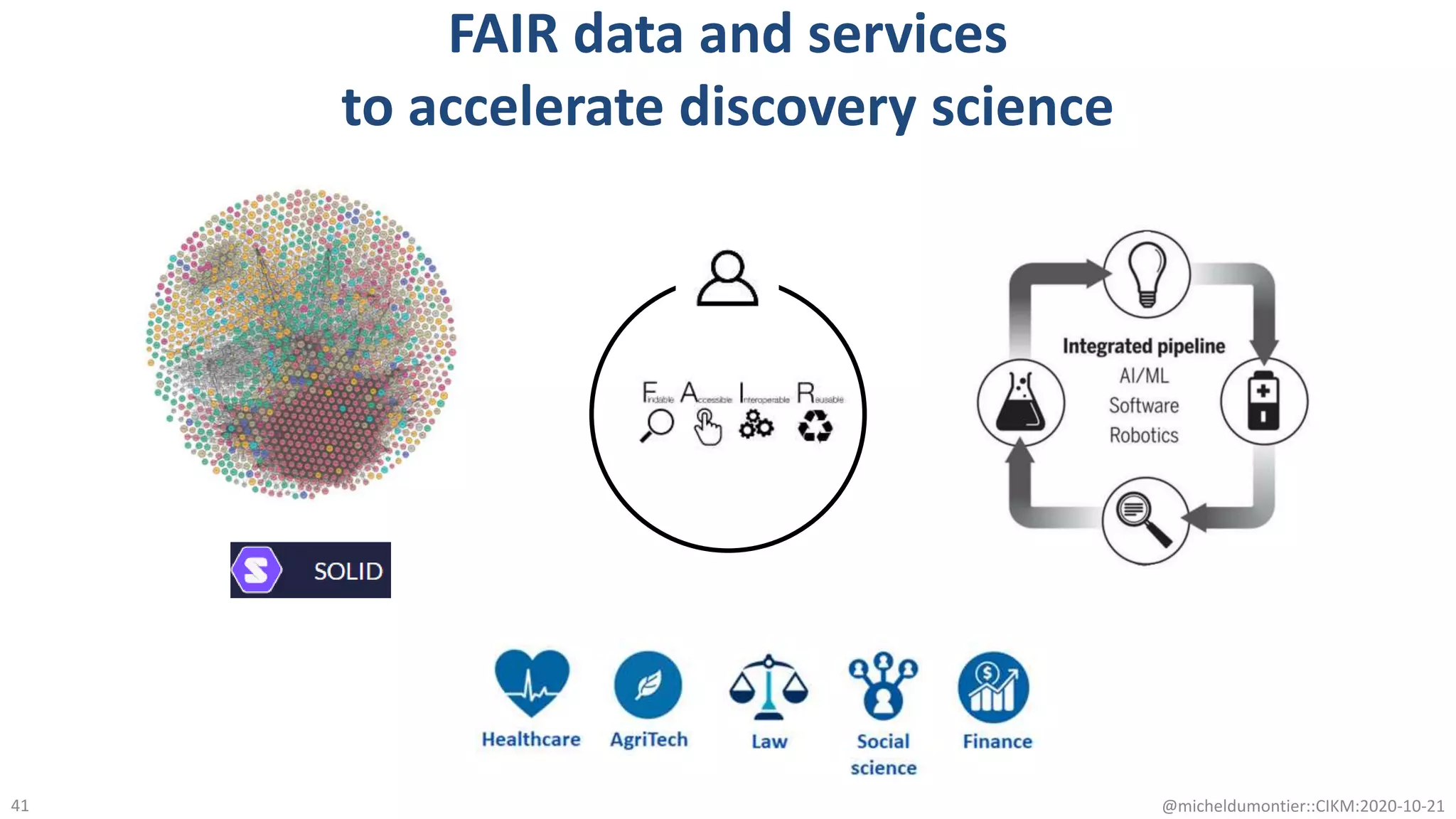
The document discusses the FAIR initiative aimed at enhancing the discovery and reuse of digital resources to accelerate scientific research, particularly in data science and biomedicine. It highlights the challenges of data reproducibility and the need for a robust infrastructure to support FAIR principles, including unique identifiers, quality metadata, and community standards. The document calls for new social, ethical, and technological frameworks to effectively utilize data for responsible research and innovation.