Axiom® Genome-Wide CHB 1 & CHB 2 Array Plate Set
•
1 like•414 views
High coverage of common and rare variants to detect disease associations in Han Chinese populations
Report
Share
Report
Share
Download to read offline
Recommended
Next Generation Diagnostics: Potential Clinical Applications of Illumina’sTec...

Rob Kelley, MBA
Global Sales Manager
Translational Genomics
September 27, 2011
2009 09 08 Wiltshire Ipit Seminar Slides

Slideshow accompanying Associate Professor Tim Wiltshire's seminar on September 8, 2009.
Moderna at 35th Annual J.P. Morgan Healthcare Conference

Moderna at 35th Annual J.P. Morgan Healthcare Conference
Monday, January 9, 2017 at 2:30 p.m. PT
Recommended
Next Generation Diagnostics: Potential Clinical Applications of Illumina’sTec...

Rob Kelley, MBA
Global Sales Manager
Translational Genomics
September 27, 2011
2009 09 08 Wiltshire Ipit Seminar Slides

Slideshow accompanying Associate Professor Tim Wiltshire's seminar on September 8, 2009.
Moderna at 35th Annual J.P. Morgan Healthcare Conference

Moderna at 35th Annual J.P. Morgan Healthcare Conference
Monday, January 9, 2017 at 2:30 p.m. PT
Specific inhibition of CK2α from an anchor outside the active site

The development of selective inhibitors of protein kinases is challenging because of the significant conservation of the ATP binding site. Here, we describe a new mechanism by which the protein kinase CK2α can be selectively inhibited using features outside the ATP site. We have identified a new binding site for small molecules on CK2α adjacent to the ATP site and behind the αD loop, termed the αD pocket. An elaborated fragment anchored in this site has been linked with a low affinity fragment binding in the ATP site, creating a novel and selective inhibitor (CAM4066) that binds CK2α with a Kd of 320 nM and shows significantly improved selectivity compared to other CK2α inhibitors. CAM4066 shows target engagement in several cell lines and similar potency to clinical trial candidate CX4945. Our data demonstrate that targeting a poorly conserved, cryptic pocket allows inhibition of CK2α via a novel mechanism, enabling the development of a new generation of selective CK2α inhibitors.
Assessment of TP53 Mutation Status in Breast Tumor Tissue using the "Ion Ampl...

Assessment of TP53 Mutation Status in Breast Tumor Tissue using the "Ion Ampl...Thermo Fisher Scientific
TP53 mutations are found in 30% of breast tumors and are associated with poor prognosis in distinct subtypes of breast cancer. Direct sequencing is commonly used to obtain TP53 mutation status in tumor tissue, but has limitations in detection level and is time-consuming. Methods targeting hotspots is insufficient for TP53 analysis since the mutations are widely spread along the gene1. Here we describe the development of the Ion TorrentTM next-generation semiconductor sequencing and Ion AmpliSeqTM technology (Life TechnologiesTM).Dr. Yao-Wei Huang - Here we go again? Emergence of a novel swine enteric alph...

Here we go again? Emergence of a novel swine enteric alphacoronavirus (SeACov) in Southern China - Dr. Yao-Wei Huang, Zhejiang University, from the 2017 North American PRRS/National Swine Improvement Federation Joint Meeting, December 1‐3, 2017, Chicago, Illinois, USA.
More presentations at http://www.swinecast.com/2017-north-american-prrs-nsif-joint-meeting
Identify Compounds that Rescue Disease Relevant Mutant Membrane Proteins

Learn about diseases caused by protein misfolding and how you can screen for compounds, known as pharmacochaperones, that rescue misfolded proteins and could be used as therapeutics.
Effects of CTGF Overexpression on Fibrosis-Related Phenotypes

Wound healing consists of three phases: inflammation, proliferation, and remodeling. During proliferation and remodeling, new extracellular matrix (ECM) is laid down by cells called fibroblasts. ECM consists of proteins that provide structural support in tissues. ECM production and fibroblast migration and proliferation are stimulated by connective tissue growth factor (CTGF). During wound healing, CTGF is induced by transforming growth factor beta (TGF-beta) to stimulate this response. When TGF-beta and CTGF are overexpressed, excessive amounts of ECM proteins are laid down, forming scar tissue, also known as fibrosis. Fibrotic tissue in organs can impair healthy function, leading to organ failure. To better understand how CTGF functions in fibrotic tissue, cells with varying levels of CTGF expression will be examined in a cell model of wounding. We will measure gene expression and observe cells for changes in fibrosis-related phenotypes. We hypothesize that increased CTGF expression will lead to enhanced cell growth, motility, and ECM production, as compared to controls. Results will provide a foundation for further research into the effects of CTGF variations on fibrosis-related phenotypes.
Nous reptes de la Medicina de Precisió

Conferència a càrrec d'Enriqueta Felip. Oncòloga de l'Hospital Vall d'Hebron de Barcelona. Líder en la investigació de càncer de pulmó al VHIO. Membre de la Junta Directiva de la Societat Espanyola d'Oncologia Mèdica i de l'European Society for Medical Oncology. En el marc de la Jornada "Els nous reptes de la Medicina de Precisió" organitzada el 12 de novembre per la Societat Catalana de Gestió Sanitària
Current and Novel Immuno-Oncology Drug Evaluation Methods via Humanized Mouse...

Dr. Bin Xie discusses the current immuno-oncology drug development landscape, different humanized mouse models available for drug testing, and the investigation of potential mechanisms via imaging mass cytometry.
Since the first immune checkpoint blocker ipilimumab was approved by the US FDA in 2011, more drug companies have sought to develop their own immune therapy drugs. Humanized peripheral blood mononuclear cell (PBMC) reconstitution in immune deficient mice is becoming a valuable model for evaluating therapeutic antibodies, especially bispecific antibodies (BsAbs), which can mediate immune cells as well as target a tumor antigen.
However, this model has several drawbacks, including a limited dosing window due to graft-versus-host-disease and insufficient natural immune cell infiltration. This has hindered wide application of the model in the development of multiple immune checkpoint inhibitors or immune agonists.
To overcome these issues, LIDE has developed a unique human PBMC/cancer cell co-transfer model which can generate three-dimensional huPBMC-infiltrated tumor tissue for immunotherapy. This model has successfully been used to evaluate the biological function of several signaling proteins and biomarkers in multiple cancers, such as melanoma, breast cancer, and lung cancer.
In this webinar, Dr. Bin Xie discusses the current immuno-oncology drug development landscape, different humanized models available for drug testing, evaluates real-world case studies, and describes the investigation of potential mechanisms by imaging mass cytometry.
Key Topics Include:
- Introduction to immuno-oncology drug development and the importance of using humanized mouse models to address scientific questions
- Evaluation of current IO platforms and new methods from LIDE, including analysis of several case studies
- Understanding the spatiotemporal interaction between tissue-infiltrating immune cells and cancer cells via imaging mass cytometry
Identification and characterization of micro rn as expressed in hiv

My masters project presented at the University of Cape Town, a study to my knowledge not conducted before.
Introducing Genomic Testing Cooperative (GTC)

A quick introduction of Genomic Testing Cooperative (GTC).
We strive to offer advanced genomic testing to communities everywhere at an affordable price.
We utilize a cooperative business model that enables academic and commercial laboratories to help physicians treat patients locally with the most advanced precision medicine.
We embrace disruptive deep learning and advanced technology to create scalable efficiencies, incomparable precision, and a more personalized approach to patient care.
https://www.genomictestingcooperative.com
Axiom® Biobank Genotyping Arrays

A choice of population-optimized GWAS imputation grids, combined with exome, loss-of-function, ADME, and eQTL markers in one high-coverage, high-value, customizable design.
Axiom™ Genome-Wide ASI 1 Array Plate

The first array that maximizes coverage of common and rare variants in East Asian populations
Axiom™ Genome-Wide CEU 1 Array Plate

The first array to maximize power for European populations with high whole-genome coverage of common and rare variants
Axiom® Genome-Wide LAT 1 Array World Array 4

Coverage-Optimized World Arrays: next-generation designs combining GWAS, replication, and fine-mapping into one array. Dense marker coverage of disease-associated SNPs, genes, and regions, plus whole-genome coverage of common and rare variants. Optimized for diverse ancestries including European, African and Native American.
A novel method for building custom ampli seq panels using optimized pcr primers 

A novel method for building custom ampli seq panels using optimized pcr primers Thermo Fisher Scientific
AmpliSeq™ is a next generation sequencing library preparation method for targeted re-sequencing that utilizes highly multiplexed PCR to amplify regions of interest. A key to successful AmpliSeq libraries is the primer panel used for target amplification. Until now primers have been available as pre-assembled ready-to-use panels, or as custom made-to-order panels. We describe a new process for creating customized panels consisting of optimized and verified PCR primers. The primer sets are available as whole genes (i.e., all of the primers needed to create libraries that cover the entire coding regions of genes) and are selectable on the ampliseq.com website by either uploading gene lists or choosing genes from disease research areas.
We show NGS sequencing data from 10 disease research-oriented panels, including newborn screening research and inherited cancer research, assembled from individual pre-verified gene sets. Panel performance data include coverage uniformity, reproducibility, and sensitivity and positive predictive value of variant calling. To demonstrate flexibility of panel content and performance, the coverage uniformity of the 59 genes recommended by the American College of Medical Genetics and Genomics for reporting of incidental findings (ACMG59) was evaluated by themselves and with up to 135 additional genes and shown to be ≥ 97% in all contexts. We also demonstrate the robustness of this method using a variety of sample types (fresh, frozen, and dried blood, cheek swabs) with both manual and fully automated library preparation methods. For Research use only. Not for use in diagnostic procedures.
VarSeq 2.6.0: Advancing Pharmacogenomics and Genomic Analysis

In the rapidly evolving field of genomic analysis, staying current with the latest research, data sources, and test advancements is crucial. In this webinar, we review how VarSeq addresses the needs to stay on top of the latest with the release of VarSeq 2.6.0.
This release features an exome-optimized workflow for LOH and CNV calling as well as the introduction of VSPGx to produce pharmacogenomic reports for gene panels as well as exomes and genomes. With the recent release of gnomAD v4, we have had many requests for the integration of this large update to the most population frequency source. With VarSeq 2.6.0, the latest version of gnomAD has been integrated into VSClinical and the updated tracks spans beyond variants to cover CNVs and gene scores to update all your workflows to the latest data.
In this webcast, we will cover.
Improved VS-CNV performance and updated exome analysis workflows.
Pharmacogenomics in action: Utilizing VSPGx for exome and genome assessments.
gnomAD v4 in practice: Updated automated and manual variant interpretation workflows.
Join us for an insightful session on the latest VarSeq 2.6.0 features, bringing you the most up-to-date data and workflows for your genomic analysis.
GIAB for AMP GeT-RM Forum

GIAB for AMP GeT-RM Forum - How well can you detect difficult variants? Benchmarking with Genome in a Bottle
More Related Content
What's hot
Specific inhibition of CK2α from an anchor outside the active site

The development of selective inhibitors of protein kinases is challenging because of the significant conservation of the ATP binding site. Here, we describe a new mechanism by which the protein kinase CK2α can be selectively inhibited using features outside the ATP site. We have identified a new binding site for small molecules on CK2α adjacent to the ATP site and behind the αD loop, termed the αD pocket. An elaborated fragment anchored in this site has been linked with a low affinity fragment binding in the ATP site, creating a novel and selective inhibitor (CAM4066) that binds CK2α with a Kd of 320 nM and shows significantly improved selectivity compared to other CK2α inhibitors. CAM4066 shows target engagement in several cell lines and similar potency to clinical trial candidate CX4945. Our data demonstrate that targeting a poorly conserved, cryptic pocket allows inhibition of CK2α via a novel mechanism, enabling the development of a new generation of selective CK2α inhibitors.
Assessment of TP53 Mutation Status in Breast Tumor Tissue using the "Ion Ampl...

Assessment of TP53 Mutation Status in Breast Tumor Tissue using the "Ion Ampl...Thermo Fisher Scientific
TP53 mutations are found in 30% of breast tumors and are associated with poor prognosis in distinct subtypes of breast cancer. Direct sequencing is commonly used to obtain TP53 mutation status in tumor tissue, but has limitations in detection level and is time-consuming. Methods targeting hotspots is insufficient for TP53 analysis since the mutations are widely spread along the gene1. Here we describe the development of the Ion TorrentTM next-generation semiconductor sequencing and Ion AmpliSeqTM technology (Life TechnologiesTM).Dr. Yao-Wei Huang - Here we go again? Emergence of a novel swine enteric alph...

Here we go again? Emergence of a novel swine enteric alphacoronavirus (SeACov) in Southern China - Dr. Yao-Wei Huang, Zhejiang University, from the 2017 North American PRRS/National Swine Improvement Federation Joint Meeting, December 1‐3, 2017, Chicago, Illinois, USA.
More presentations at http://www.swinecast.com/2017-north-american-prrs-nsif-joint-meeting
Identify Compounds that Rescue Disease Relevant Mutant Membrane Proteins

Learn about diseases caused by protein misfolding and how you can screen for compounds, known as pharmacochaperones, that rescue misfolded proteins and could be used as therapeutics.
Effects of CTGF Overexpression on Fibrosis-Related Phenotypes

Wound healing consists of three phases: inflammation, proliferation, and remodeling. During proliferation and remodeling, new extracellular matrix (ECM) is laid down by cells called fibroblasts. ECM consists of proteins that provide structural support in tissues. ECM production and fibroblast migration and proliferation are stimulated by connective tissue growth factor (CTGF). During wound healing, CTGF is induced by transforming growth factor beta (TGF-beta) to stimulate this response. When TGF-beta and CTGF are overexpressed, excessive amounts of ECM proteins are laid down, forming scar tissue, also known as fibrosis. Fibrotic tissue in organs can impair healthy function, leading to organ failure. To better understand how CTGF functions in fibrotic tissue, cells with varying levels of CTGF expression will be examined in a cell model of wounding. We will measure gene expression and observe cells for changes in fibrosis-related phenotypes. We hypothesize that increased CTGF expression will lead to enhanced cell growth, motility, and ECM production, as compared to controls. Results will provide a foundation for further research into the effects of CTGF variations on fibrosis-related phenotypes.
Nous reptes de la Medicina de Precisió

Conferència a càrrec d'Enriqueta Felip. Oncòloga de l'Hospital Vall d'Hebron de Barcelona. Líder en la investigació de càncer de pulmó al VHIO. Membre de la Junta Directiva de la Societat Espanyola d'Oncologia Mèdica i de l'European Society for Medical Oncology. En el marc de la Jornada "Els nous reptes de la Medicina de Precisió" organitzada el 12 de novembre per la Societat Catalana de Gestió Sanitària
Current and Novel Immuno-Oncology Drug Evaluation Methods via Humanized Mouse...

Dr. Bin Xie discusses the current immuno-oncology drug development landscape, different humanized mouse models available for drug testing, and the investigation of potential mechanisms via imaging mass cytometry.
Since the first immune checkpoint blocker ipilimumab was approved by the US FDA in 2011, more drug companies have sought to develop their own immune therapy drugs. Humanized peripheral blood mononuclear cell (PBMC) reconstitution in immune deficient mice is becoming a valuable model for evaluating therapeutic antibodies, especially bispecific antibodies (BsAbs), which can mediate immune cells as well as target a tumor antigen.
However, this model has several drawbacks, including a limited dosing window due to graft-versus-host-disease and insufficient natural immune cell infiltration. This has hindered wide application of the model in the development of multiple immune checkpoint inhibitors or immune agonists.
To overcome these issues, LIDE has developed a unique human PBMC/cancer cell co-transfer model which can generate three-dimensional huPBMC-infiltrated tumor tissue for immunotherapy. This model has successfully been used to evaluate the biological function of several signaling proteins and biomarkers in multiple cancers, such as melanoma, breast cancer, and lung cancer.
In this webinar, Dr. Bin Xie discusses the current immuno-oncology drug development landscape, different humanized models available for drug testing, evaluates real-world case studies, and describes the investigation of potential mechanisms by imaging mass cytometry.
Key Topics Include:
- Introduction to immuno-oncology drug development and the importance of using humanized mouse models to address scientific questions
- Evaluation of current IO platforms and new methods from LIDE, including analysis of several case studies
- Understanding the spatiotemporal interaction between tissue-infiltrating immune cells and cancer cells via imaging mass cytometry
Identification and characterization of micro rn as expressed in hiv

My masters project presented at the University of Cape Town, a study to my knowledge not conducted before.
Introducing Genomic Testing Cooperative (GTC)

A quick introduction of Genomic Testing Cooperative (GTC).
We strive to offer advanced genomic testing to communities everywhere at an affordable price.
We utilize a cooperative business model that enables academic and commercial laboratories to help physicians treat patients locally with the most advanced precision medicine.
We embrace disruptive deep learning and advanced technology to create scalable efficiencies, incomparable precision, and a more personalized approach to patient care.
https://www.genomictestingcooperative.com
What's hot (11)
Medinfo2013 - An RDF/OWL Knowledge Base for Query Answering and Decision Supp...

Medinfo2013 - An RDF/OWL Knowledge Base for Query Answering and Decision Supp...
Specific inhibition of CK2α from an anchor outside the active site

Specific inhibition of CK2α from an anchor outside the active site
Assessment of TP53 Mutation Status in Breast Tumor Tissue using the "Ion Ampl...

Assessment of TP53 Mutation Status in Breast Tumor Tissue using the "Ion Ampl...
Dr. Yao-Wei Huang - Here we go again? Emergence of a novel swine enteric alph...

Dr. Yao-Wei Huang - Here we go again? Emergence of a novel swine enteric alph...
Identify Compounds that Rescue Disease Relevant Mutant Membrane Proteins

Identify Compounds that Rescue Disease Relevant Mutant Membrane Proteins
Effects of CTGF Overexpression on Fibrosis-Related Phenotypes

Effects of CTGF Overexpression on Fibrosis-Related Phenotypes
Current and Novel Immuno-Oncology Drug Evaluation Methods via Humanized Mouse...

Current and Novel Immuno-Oncology Drug Evaluation Methods via Humanized Mouse...
Identification and characterization of micro rn as expressed in hiv

Identification and characterization of micro rn as expressed in hiv
Similar to Axiom® Genome-Wide CHB 1 & CHB 2 Array Plate Set
Axiom® Biobank Genotyping Arrays

A choice of population-optimized GWAS imputation grids, combined with exome, loss-of-function, ADME, and eQTL markers in one high-coverage, high-value, customizable design.
Axiom™ Genome-Wide ASI 1 Array Plate

The first array that maximizes coverage of common and rare variants in East Asian populations
Axiom™ Genome-Wide CEU 1 Array Plate

The first array to maximize power for European populations with high whole-genome coverage of common and rare variants
Axiom® Genome-Wide LAT 1 Array World Array 4

Coverage-Optimized World Arrays: next-generation designs combining GWAS, replication, and fine-mapping into one array. Dense marker coverage of disease-associated SNPs, genes, and regions, plus whole-genome coverage of common and rare variants. Optimized for diverse ancestries including European, African and Native American.
A novel method for building custom ampli seq panels using optimized pcr primers 

A novel method for building custom ampli seq panels using optimized pcr primers Thermo Fisher Scientific
AmpliSeq™ is a next generation sequencing library preparation method for targeted re-sequencing that utilizes highly multiplexed PCR to amplify regions of interest. A key to successful AmpliSeq libraries is the primer panel used for target amplification. Until now primers have been available as pre-assembled ready-to-use panels, or as custom made-to-order panels. We describe a new process for creating customized panels consisting of optimized and verified PCR primers. The primer sets are available as whole genes (i.e., all of the primers needed to create libraries that cover the entire coding regions of genes) and are selectable on the ampliseq.com website by either uploading gene lists or choosing genes from disease research areas.
We show NGS sequencing data from 10 disease research-oriented panels, including newborn screening research and inherited cancer research, assembled from individual pre-verified gene sets. Panel performance data include coverage uniformity, reproducibility, and sensitivity and positive predictive value of variant calling. To demonstrate flexibility of panel content and performance, the coverage uniformity of the 59 genes recommended by the American College of Medical Genetics and Genomics for reporting of incidental findings (ACMG59) was evaluated by themselves and with up to 135 additional genes and shown to be ≥ 97% in all contexts. We also demonstrate the robustness of this method using a variety of sample types (fresh, frozen, and dried blood, cheek swabs) with both manual and fully automated library preparation methods. For Research use only. Not for use in diagnostic procedures.
VarSeq 2.6.0: Advancing Pharmacogenomics and Genomic Analysis

In the rapidly evolving field of genomic analysis, staying current with the latest research, data sources, and test advancements is crucial. In this webinar, we review how VarSeq addresses the needs to stay on top of the latest with the release of VarSeq 2.6.0.
This release features an exome-optimized workflow for LOH and CNV calling as well as the introduction of VSPGx to produce pharmacogenomic reports for gene panels as well as exomes and genomes. With the recent release of gnomAD v4, we have had many requests for the integration of this large update to the most population frequency source. With VarSeq 2.6.0, the latest version of gnomAD has been integrated into VSClinical and the updated tracks spans beyond variants to cover CNVs and gene scores to update all your workflows to the latest data.
In this webcast, we will cover.
Improved VS-CNV performance and updated exome analysis workflows.
Pharmacogenomics in action: Utilizing VSPGx for exome and genome assessments.
gnomAD v4 in practice: Updated automated and manual variant interpretation workflows.
Join us for an insightful session on the latest VarSeq 2.6.0 features, bringing you the most up-to-date data and workflows for your genomic analysis.
GIAB for AMP GeT-RM Forum

GIAB for AMP GeT-RM Forum - How well can you detect difficult variants? Benchmarking with Genome in a Bottle
Genome in a Bottle- reference materials to benchmark challenging variants and...

Genome in a Bottle- reference materials to benchmark challenging variants and regions of the human genome
Development of a high-throughput high-density SNP genotyping array for bovine

Ali Pirani, Bioinformatics, Affymetrix.Our bioinformatics team first developed a workflow for validating de novo SNPs in animals. They have since applied this successfully in multiple plant species.
A next generation sequencing based sample-to-result pharmacogenomics research...

A next generation sequencing based sample-to-result pharmacogenomics research...Thermo Fisher Scientific
Pharmacogenomics (PGx) is the study of genetic variations in terms of their response to drugs. Variations in gene sequence or copy numbers may result in complete loss of function, partial decrease or increase in enzyme activity, or an altered affinity for substrates, which may in turn significantly impact drug efficacy. PGx studies are becoming increasingly important for precision medicine. We have developed a next generation sequencing (NGS) PGx research solution with increased flexibility on the assay targets and combined detection of SNP/INDEL genotyping and CNV using Ion AmpliSeq™ technology for low to medium throughput laboratories. With this highly multiplexed PGx research panel we can profile a set of 136 genetic markers in 40 known PGx related genes (Table 1) and determine CYP2D6 copy number variation (CNV, Figure 1) in a single reaction using Ion Torrent™ semiconductor sequencing.Characterizing Alzheimer’s Disease candidate genes and transcripts with targe...

Characterizing Alzheimer’s Disease candidate genes and transcripts with targe...Integrated DNA Technologies
Alzheimer’s disease (AD) is a devastating neurodegenerative disease that is genetically complex. Although great progress has been made in identifying fully penetrant mutations in genes that cause early-onset AD, these still represent a very small percentage of AD cases. Large-scale, genome-wide association studies (GWAS) have identified at least 20 additional genetic risk loci for the more common form: late-onset AD. However, the identified SNPs are typically not the actual risk variants, but are in linkage disequilibrium with the presumed causative variants [1].
To help identify causative genetic variants, we have combined highly accurate, long-read sequencing with hybrid-capture technology. In this collaborative webinar*, we present this method and show how combining IDT xGen® Lockdown® Probes with PacBio SMRT® Sequencing allows targeting and sequencing of candidate genes from genomic DNA and corresponding transcripts from cDNA. Using a panel of target capture probes for 35 AD candidate genes, we demonstrate the power of this approach by looking at data for two individuals with AD. Some additional benefits of this method include the ability to leverage long reads, phase heterozygous variants, and link corresponding transcript isoforms to their respective alleles.
Reference: 1. Van Cauwenberghe C, Van Broeckhoven C, Sleegers K. (2016) The genetic landscape of Alzheimer disease: clinical implications and perspectives. Genet Med, 18(5):421–430.
* This presentation represents a collaboration between Pacific Biosciences and Integrated DNA Technologies. The individual opinions expressed may not reflect shared opinions of Pacific Biosciences and Integrated DNA Technologies.Automating the ACMG Guidelines with VSClinical

Clinical Genetic testing requires a complex analysis using the totality of our knowledge about the clinical relevance of a variant and a gene. This includes bioinformatic evidence as well as clinical evidence. The ACMG Guidelines provided a framework in which to score variants based on this evidence, and while some of those scoring criteria require close consultation of the clinical context for a given patient, much of it can be automated.
In this webcast, we review how VSClinical automates the ACMG scoring guidelines while integrating the collective lab expertise from previously classified variants and preferences about genes. We will cover:
Using the ACMG Auto Classifier as part the filtering strategy for gene panels and trio workflows
How gene preferences such as the default transcript, inheritance model, and disorder are updated and saved from VSClinical and used in all future analysis
How the per-variant recommendation engine builds on the auto-classification with descriptive reasons for answering each criterion yes or no
Using the auto-interpretation to present the evidence for all scored criteria in a human-readable paragraph
Working with VSClinical’s self-learning knowledgebase that incorporates previously classified variants and genes inform the interpretation of new variants!
Heptotox biomarker discovery Ontomine

Use functional group frequency rules and biological informatics to mine hepatotoxicity biomarkers
Global Gene Expression Profiles from Bladder Tumor FFPE Samples

Cancer is a disease characterized by uncontrolled cell growth and proliferation. Recent advances in molecular medicine and cancer biology have changed the way research clinicians evaluate and consider treatment. Selected tumor biomarkers have been utilized as targets for drug therapy leading to better more effective treatment. Gene expression profiling has been used for identifying new biomarkers for tumor classification and driving decision making for better patient outcome in different tumor types. DNA microarrays have become a key method to acquire a comparative snapshot of the gene expression profile from test samples in a high throughput manner. Quantitative PCR and newer sequencing techniques are popular research alternatives offering highly accurate gene expression measurements, but with limitations due to cost, complex instrumentation and analysis needs. RNA extracted from formalin fixed paraffin embedded tissue (FFPE) creates considerable additional challenges in acquiring accurate gene expression measurements due to the highly fragmented and compromised integrity of FFPE RNA due to the fixation process.
Apac distributor training series 3 swift product for cancer study

Apac distributor training series 3 swift product for cancer study
Genome in a Bottle - Towards new benchmarks for the “dark matter” of the huma...

Genome in a Bottle: Towards new benchmarks for the “dark matter” of the human genome
Similar to Axiom® Genome-Wide CHB 1 & CHB 2 Array Plate Set (20)
A novel method for building custom ampli seq panels using optimized pcr primers 

A novel method for building custom ampli seq panels using optimized pcr primers
VarSeq 2.6.0: Advancing Pharmacogenomics and Genomic Analysis

VarSeq 2.6.0: Advancing Pharmacogenomics and Genomic Analysis
Genome in a Bottle- reference materials to benchmark challenging variants and...

Genome in a Bottle- reference materials to benchmark challenging variants and...
Development of a high-throughput high-density SNP genotyping array for bovine

Development of a high-throughput high-density SNP genotyping array for bovine
A next generation sequencing based sample-to-result pharmacogenomics research...

A next generation sequencing based sample-to-result pharmacogenomics research...
Characterizing Alzheimer’s Disease candidate genes and transcripts with targe...

Characterizing Alzheimer’s Disease candidate genes and transcripts with targe...
Global Gene Expression Profiles from Bladder Tumor FFPE Samples

Global Gene Expression Profiles from Bladder Tumor FFPE Samples
Apac distributor training series 3 swift product for cancer study

Apac distributor training series 3 swift product for cancer study
Genome in a Bottle - Towards new benchmarks for the “dark matter” of the huma...

Genome in a Bottle - Towards new benchmarks for the “dark matter” of the huma...
More from Affymetrix
SNP genotyping using the Affymetrix® Axiom® Genome-Wide Pan-African (PanAFR) ...

Affymetrix Research and Development Scientists Many GWAS arrays are optimized for European populations but significantly compromise study power when used in non-European cohorts. Read about the design strategy of the Axiom Pan-African Array, our leading solution for optimized whole-genome studies of African ancestries.
A SNP array for human population genetics studies

Yontao Lu, Teri Genschoreck, Swapan Mallick, Amy Ollmann, Nick Patterson, Yiping Zhan, Teresa Webster, David Reich Overview of the Human Origins Array, the first array developed with leading geneticists to enable rigorous population genetics studies.
FFPE Applications Solutions brochure

A Platform of comprehensive genomics solutions for analyzing FFPE archival tissues from whole-genome to single molecules
Download our publication archive list

Includes over 250 transcriptomics and genomics papers published from 2009 to 2011 in plants, animals, and other non-human species; sorted by species.
Designing GWAS arrays for efficient imputation-based coverage

Yiping Zhan, Bioinformatics, Affymetrix (presented at ASHG 2012).
Our proprietary, imputation-based SNP selection method creates population-optimized GWAS array designs that use fewer markers to deliver better 1000 Genomes coverage than pairwise tagSNP arrays.
Use of Affymetrix Arrays (GeneChip® Human Transcriptome 2.0 Array and Cytosca...

Giovanni Martinelli, PhD
Associate Professor of Haematology, Institute of Haematology and Medical Oncology, University of Bologna, Italy
From trials evaluating drugs to trials evaluating treatment algorithms – Focu...

Christophe Le Tourneau, MD, PhD
Medical Oncologist, Head of the Phase I Program, Institut Curie, France
Affymetrix OncoScan®* data analysis with Nexus Copy Number™

Zhiwei Che, PhD
Director of Application Science, BioDiscovery Inc, USA
Phenotypic identification of subclones in multiple myeloma with different gen...

Bruno Paiva, PhD
Scientific Co-Director of CIMA LAB Diagnostics and Director of the Unit of Cytometry, University of Navarra, Spain
Mitigating genotyping application note

Mitigating sequencing errors, monomorphs, and poor performing markers during de novo SNP selection for genotyping applications
Statistical methods for off-target variant genotyping on Affymetrix' Axiom Ar...

Teresa Webster, Bioinformatics, Affymetrix.Automated clustering and calling of ""Off-Target Variants,"" atypical cluster patterns caused by secondary variants in the probe hybridization region.
Allopolyploid Genotyping Algorithm on Affymetrix' Axiom Arrays

Hong Gao, Bioinformatics, Affymetrix.An automated algorithm that reduces genotype analysis to ~1 hour is described and applied to hexaploid wheat data.
SNP genotyping of markers with nearby secondary polymorphisms using Affymetri...

Laurent Bellon, Product Development, Affymetrix.Axiom® Genotyping Solution is uniquely suited to genotyping of highly polymorphic genomes as it is permissive to secondary variants as close as 10 bp from the target variant.
Best practices for genotyping analysis of plant and animal genomes with Affym...

Ali Pirani, Bioinformatics, Affymetrix.Overview of the best workflow to get from data to results in the shortest time.
SNP genotyping using Affymetrix' Axiom Genotyping Solution

Michael Shapero, Product Development, Affymetrix.Outline of the development and performance of the new Axiom® 384 high-throughput format for low-cost genotyping of up to 50,000 SNPs.
More from Affymetrix (17)
SNP genotyping using the Affymetrix® Axiom® Genome-Wide Pan-African (PanAFR) ...

SNP genotyping using the Affymetrix® Axiom® Genome-Wide Pan-African (PanAFR) ...
Designing GWAS arrays for efficient imputation-based coverage

Designing GWAS arrays for efficient imputation-based coverage
Use of Affymetrix Arrays (GeneChip® Human Transcriptome 2.0 Array and Cytosca...

Use of Affymetrix Arrays (GeneChip® Human Transcriptome 2.0 Array and Cytosca...
From trials evaluating drugs to trials evaluating treatment algorithms – Focu...

From trials evaluating drugs to trials evaluating treatment algorithms – Focu...
Affymetrix OncoScan®* data analysis with Nexus Copy Number™

Affymetrix OncoScan®* data analysis with Nexus Copy Number™
Phenotypic identification of subclones in multiple myeloma with different gen...

Phenotypic identification of subclones in multiple myeloma with different gen...
Statistical methods for off-target variant genotyping on Affymetrix' Axiom Ar...

Statistical methods for off-target variant genotyping on Affymetrix' Axiom Ar...
Allopolyploid Genotyping Algorithm on Affymetrix' Axiom Arrays

Allopolyploid Genotyping Algorithm on Affymetrix' Axiom Arrays
SNP genotyping of markers with nearby secondary polymorphisms using Affymetri...

SNP genotyping of markers with nearby secondary polymorphisms using Affymetri...
Best practices for genotyping analysis of plant and animal genomes with Affym...

Best practices for genotyping analysis of plant and animal genomes with Affym...
SNP genotyping using Affymetrix' Axiom Genotyping Solution

SNP genotyping using Affymetrix' Axiom Genotyping Solution
Recently uploaded
Astronomy Update- Curiosity’s exploration of Mars _ Local Briefs _ leadertele...

Article written for leader telegram
SCHIZOPHRENIA Disorder/ Brain Disorder.pdf

This pdf is about the Schizophrenia.
For more details visit on YouTube; @SELF-EXPLANATORY;
https://www.youtube.com/channel/UCAiarMZDNhe1A3Rnpr_WkzA/videos
Thanks...!
Structures and textures of metamorphic rocks

It is useful for the Under Graduating students for easy understanding and it's useful for the exam preparations.
The ASGCT Annual Meeting was packed with exciting progress in the field advan...

The ASGCT Annual Meeting was packed with exciting progress in the field advancing efforts to deliver highly promising therapies to more patients.
Nutraceutical market, scope and growth: Herbal drug technology

As consumer awareness of health and wellness rises, the nutraceutical market—which includes goods like functional meals, drinks, and dietary supplements that provide health advantages beyond basic nutrition—is growing significantly. As healthcare expenses rise, the population ages, and people want natural and preventative health solutions more and more, this industry is increasing quickly. Further driving market expansion are product formulation innovations and the use of cutting-edge technology for customized nutrition. With its worldwide reach, the nutraceutical industry is expected to keep growing and provide significant chances for research and investment in a number of categories, including vitamins, minerals, probiotics, and herbal supplements.
4. An Overview of Sugarcane White Leaf Disease in Vietnam.pdf

An overview of Sugarcane White Leaf Disease in Vietnam
Observation of Io’s Resurfacing via Plume Deposition Using Ground-based Adapt...

Since volcanic activity was first discovered on Io from Voyager images in 1979, changes
on Io’s surface have been monitored from both spacecraft and ground-based telescopes.
Here, we present the highest spatial resolution images of Io ever obtained from a groundbased telescope. These images, acquired by the SHARK-VIS instrument on the Large
Binocular Telescope, show evidence of a major resurfacing event on Io’s trailing hemisphere. When compared to the most recent spacecraft images, the SHARK-VIS images
show that a plume deposit from a powerful eruption at Pillan Patera has covered part
of the long-lived Pele plume deposit. Although this type of resurfacing event may be common on Io, few have been detected due to the rarity of spacecraft visits and the previously low spatial resolution available from Earth-based telescopes. The SHARK-VIS instrument ushers in a new era of high resolution imaging of Io’s surface using adaptive
optics at visible wavelengths.
(May 29th, 2024) Advancements in Intravital Microscopy- Insights for Preclini...

(May 29th, 2024) Advancements in Intravital Microscopy- Insights for Preclini...Scintica Instrumentation
Intravital microscopy (IVM) is a powerful tool utilized to study cellular behavior over time and space in vivo. Much of our understanding of cell biology has been accomplished using various in vitro and ex vivo methods; however, these studies do not necessarily reflect the natural dynamics of biological processes. Unlike traditional cell culture or fixed tissue imaging, IVM allows for the ultra-fast high-resolution imaging of cellular processes over time and space and were studied in its natural environment. Real-time visualization of biological processes in the context of an intact organism helps maintain physiological relevance and provide insights into the progression of disease, response to treatments or developmental processes.
In this webinar we give an overview of advanced applications of the IVM system in preclinical research. IVIM technology is a provider of all-in-one intravital microscopy systems and solutions optimized for in vivo imaging of live animal models at sub-micron resolution. The system’s unique features and user-friendly software enables researchers to probe fast dynamic biological processes such as immune cell tracking, cell-cell interaction as well as vascularization and tumor metastasis with exceptional detail. This webinar will also give an overview of IVM being utilized in drug development, offering a view into the intricate interaction between drugs/nanoparticles and tissues in vivo and allows for the evaluation of therapeutic intervention in a variety of tissues and organs. This interdisciplinary collaboration continues to drive the advancements of novel therapeutic strategies.
Richard's entangled aventures in wonderland

Since the loophole-free Bell experiments of 2020 and the Nobel prizes in physics of 2022, critics of Bell's work have retreated to the fortress of super-determinism. Now, super-determinism is a derogatory word - it just means "determinism". Palmer, Hance and Hossenfelder argue that quantum mechanics and determinism are not incompatible, using a sophisticated mathematical construction based on a subtle thinning of allowed states and measurements in quantum mechanics, such that what is left appears to make Bell's argument fail, without altering the empirical predictions of quantum mechanics. I think however that it is a smoke screen, and the slogan "lost in math" comes to my mind. I will discuss some other recent disproofs of Bell's theorem using the language of causality based on causal graphs. Causal thinking is also central to law and justice. I will mention surprising connections to my work on serial killer nurse cases, in particular the Dutch case of Lucia de Berk and the current UK case of Lucy Letby.
Multi-source connectivity as the driver of solar wind variability in the heli...

The ambient solar wind that flls the heliosphere originates from multiple
sources in the solar corona and is highly structured. It is often described
as high-speed, relatively homogeneous, plasma streams from coronal
holes and slow-speed, highly variable, streams whose source regions are
under debate. A key goal of ESA/NASA’s Solar Orbiter mission is to identify
solar wind sources and understand what drives the complexity seen in the
heliosphere. By combining magnetic feld modelling and spectroscopic
techniques with high-resolution observations and measurements, we show
that the solar wind variability detected in situ by Solar Orbiter in March
2022 is driven by spatio-temporal changes in the magnetic connectivity to
multiple sources in the solar atmosphere. The magnetic feld footpoints
connected to the spacecraft moved from the boundaries of a coronal hole
to one active region (12961) and then across to another region (12957). This
is refected in the in situ measurements, which show the transition from fast
to highly Alfvénic then to slow solar wind that is disrupted by the arrival of
a coronal mass ejection. Our results describe solar wind variability at 0.5 au
but are applicable to near-Earth observatories.
Nucleic Acid-its structural and functional complexity.

This presentation explores a brief idea about the structural and functional attributes of nucleotides, the structure and function of genetic materials along with the impact of UV rays and pH upon them.
extra-chromosomal-inheritance[1].pptx.pdfpdf![extra-chromosomal-inheritance[1].pptx.pdfpdf](data:image/gif;base64,R0lGODlhAQABAIAAAAAAAP///yH5BAEAAAAALAAAAAABAAEAAAIBRAA7)
![extra-chromosomal-inheritance[1].pptx.pdfpdf](data:image/gif;base64,R0lGODlhAQABAIAAAAAAAP///yH5BAEAAAAALAAAAAABAAEAAAIBRAA7)
Slide 1: Title Slide
Extrachromosomal Inheritance
Slide 2: Introduction to Extrachromosomal Inheritance
Definition: Extrachromosomal inheritance refers to the transmission of genetic material that is not found within the nucleus.
Key Components: Involves genes located in mitochondria, chloroplasts, and plasmids.
Slide 3: Mitochondrial Inheritance
Mitochondria: Organelles responsible for energy production.
Mitochondrial DNA (mtDNA): Circular DNA molecule found in mitochondria.
Inheritance Pattern: Maternally inherited, meaning it is passed from mothers to all their offspring.
Diseases: Examples include Leber’s hereditary optic neuropathy (LHON) and mitochondrial myopathy.
Slide 4: Chloroplast Inheritance
Chloroplasts: Organelles responsible for photosynthesis in plants.
Chloroplast DNA (cpDNA): Circular DNA molecule found in chloroplasts.
Inheritance Pattern: Often maternally inherited in most plants, but can vary in some species.
Examples: Variegation in plants, where leaf color patterns are determined by chloroplast DNA.
Slide 5: Plasmid Inheritance
Plasmids: Small, circular DNA molecules found in bacteria and some eukaryotes.
Features: Can carry antibiotic resistance genes and can be transferred between cells through processes like conjugation.
Significance: Important in biotechnology for gene cloning and genetic engineering.
Slide 6: Mechanisms of Extrachromosomal Inheritance
Non-Mendelian Patterns: Do not follow Mendel’s laws of inheritance.
Cytoplasmic Segregation: During cell division, organelles like mitochondria and chloroplasts are randomly distributed to daughter cells.
Heteroplasmy: Presence of more than one type of organellar genome within a cell, leading to variation in expression.
Slide 7: Examples of Extrachromosomal Inheritance
Four O’clock Plant (Mirabilis jalapa): Shows variegated leaves due to different cpDNA in leaf cells.
Petite Mutants in Yeast: Result from mutations in mitochondrial DNA affecting respiration.
Slide 8: Importance of Extrachromosomal Inheritance
Evolution: Provides insight into the evolution of eukaryotic cells.
Medicine: Understanding mitochondrial inheritance helps in diagnosing and treating mitochondrial diseases.
Agriculture: Chloroplast inheritance can be used in plant breeding and genetic modification.
Slide 9: Recent Research and Advances
Gene Editing: Techniques like CRISPR-Cas9 are being used to edit mitochondrial and chloroplast DNA.
Therapies: Development of mitochondrial replacement therapy (MRT) for preventing mitochondrial diseases.
Slide 10: Conclusion
Summary: Extrachromosomal inheritance involves the transmission of genetic material outside the nucleus and plays a crucial role in genetics, medicine, and biotechnology.
Future Directions: Continued research and technological advancements hold promise for new treatments and applications.
Slide 11: Questions and Discussion
Invite Audience: Open the floor for any questions or further discussion on the topic.
Recently uploaded (20)
Astronomy Update- Curiosity’s exploration of Mars _ Local Briefs _ leadertele...

Astronomy Update- Curiosity’s exploration of Mars _ Local Briefs _ leadertele...
PRESENTATION ABOUT PRINCIPLE OF COSMATIC EVALUATION

PRESENTATION ABOUT PRINCIPLE OF COSMATIC EVALUATION
The ASGCT Annual Meeting was packed with exciting progress in the field advan...

The ASGCT Annual Meeting was packed with exciting progress in the field advan...
Nutraceutical market, scope and growth: Herbal drug technology

Nutraceutical market, scope and growth: Herbal drug technology
4. An Overview of Sugarcane White Leaf Disease in Vietnam.pdf

4. An Overview of Sugarcane White Leaf Disease in Vietnam.pdf
Observation of Io’s Resurfacing via Plume Deposition Using Ground-based Adapt...

Observation of Io’s Resurfacing via Plume Deposition Using Ground-based Adapt...
(May 29th, 2024) Advancements in Intravital Microscopy- Insights for Preclini...

(May 29th, 2024) Advancements in Intravital Microscopy- Insights for Preclini...
Mammalian Pineal Body Structure and Also Functions

Mammalian Pineal Body Structure and Also Functions
Multi-source connectivity as the driver of solar wind variability in the heli...

Multi-source connectivity as the driver of solar wind variability in the heli...
Nucleic Acid-its structural and functional complexity.

Nucleic Acid-its structural and functional complexity.
Circulatory system_ Laplace law. Ohms law.reynaults law,baro-chemo-receptors-...

Circulatory system_ Laplace law. Ohms law.reynaults law,baro-chemo-receptors-...
In silico drugs analogue design: novobiocin analogues.pptx

In silico drugs analogue design: novobiocin analogues.pptx
Axiom® Genome-Wide CHB 1 & CHB 2 Array Plate Set
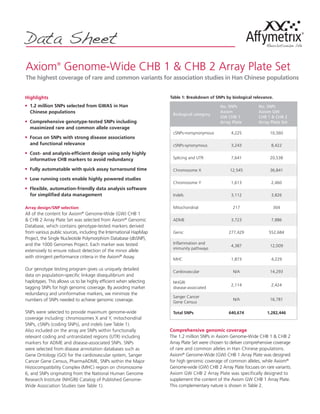
- 1. Axiom® Genome-Wide CHB 1 & CHB 2 Array Plate Set The highest coverage of rare and common variants for association studies in Han Chinese populations Highlights n 1.2 million SNPs selected from GWAS in Han Chinese populations n Comprehensive genotype-tested SNPs including maximized rare and common allele coverage n Focus on SNPs with strong disease associations and functional relevance n Cost- and analysis-efficient design using only highly informative CHB markers to avoid redundancy n Fully automatable with quick assay turnaround time n Low running costs enable highly powered studies n Flexible, automation-friendly data analysis software for simplified data management Array design/SNP selection All of the content for Axiom® Genome-Wide (GW) CHB 1 & CHB 2 Array Plate Set was selected from Axiom® Genomic Database, which contains genotype-tested markers derived from various public sources, including the International HapMap Project, the Single Nucleotide Polymorphism Database (dbSNP), and the 1000 Genomes Project. Each marker was tested extensively to ensure robust detection of the minor allele with stringent performance criteria in the Axiom® Assay. Our genotype testing program gives us uniquely detailed data on population-specific linkage disequilibrium and haplotypes. This allows us to be highly efficient when selecting tagging SNPs for high genomic coverage. By avoiding marker redundancy and uninformative markers, we minimize the numbers of SNPs needed to achieve genomic coverage. SNPs were selected to provide maximum genome-wide coverage including: chromosomes X and Y, mitochondrial SNPs, cSNPs (coding SNPs), and indels (see Table 1). Also included on the array are SNPs within functionally relevant coding and untranslated regions (UTR) including markers for ADME and disease-associated SNPs. SNPs were selected from disease annotation databases such as Gene Ontology (GO) for the cardiovascular system, Sanger Cancer Gene Census, PharmaADME, SNPs within the Major Histocompatibility Complex (MHC) region on chromosome 6, and SNPs originating from the National Human Genome Research Institute (NHGRI) Catalog of Published Genome- Wide Association Studies (see Table 1). Comprehensive genomic coverage The 1.2 million SNPs in Axiom Genome-Wide CHB 1 & CHB 2 Array Plate Set were chosen to deliver comprehensive coverage of rare and common alleles in Han Chinese populations. Axiom® Genome-Wide (GW) CHB 1 Array Plate was designed for high genomic coverage of common alleles, while Axiom® Genome-wide (GW) CHB 2 Array Plate focuses on rare variants. Axiom GW CHB 2 Array Plate was specifically designed to supplement the content of the Axiom GW CHB 1 Array Plate. This complementary nature is shown in Table 2. Data Sheet Table 1: Breakdown of SNPs by biological relevance. Biological category No. SNPs Axiom GW CHB 1 Array Plate No. SNPs Axiom GW CHB 1 & CHB 2 Array Plate Set cSNPs-nonsynonymous 4,225 10,560 cSNPs-synonymous 3,243 8,422 Splicing and UTR 7,641 20,538 Chromosome X 12,545 36,841 Chromosome Y 1,613 2,460 Indels 3,112 3,826 Mitochondrial 217 304 ADME 3,723 7,886 Genic 277,429 552,684 Inflammation and immunity pathways 4,387 12,009 MHC 1,873 4,229 Cardiovascular N/A 14,293 NHGRI disease-associated 2,114 2,424 Sanger Cancer Gene Census N/A 16,781 Total SNPs 640,674 1,282,446
- 2. Table 2: Genomic coverage of Axiom® GW CHB 1 Array Plate and Axiom® GW CHB 1 & CHB 2 Array Plate Set. Pairwise genomic coverage at r2 >0.8 for SNPs in the 1000 Genomes Project Phase 1 integrated release Dec 2011; Han Chinese in Beijing (CHB), Han Chinese South (CHS), and Japanese in Tokyo (JPT) alleles. Axiom GW CHB 1 Array Plate Axiom GW CHB 1 & CHB 2 Array Plate Set Other commercially available Chinese array Population MAF >2% MAF >5% MAF >2% MAF >5% MAF >2% MAF >5% CHB 66% 77% 80% 86% 72% 79% CHS 64% 75% 79% 86% 72% 79% JPT 64% 72% 78% 84% 72% 78% Superior assay performance As shown in Table 3, Axiom® Genome-Wide CHB 1 & CHB 2 Array Plate Set has superior technical performance offering high confidence for genome-wide association studies. Table 3: Performance achieved by Axiom GW CHB 1 & CHB 2 Array Plate Set. Calculated based on 190 Chinese and Japanese HapMap samples. Performance metric Specification Average SNP call rate >99% Average HapMap concordance >99.5% Average reproducibility >99.7% Ordering information Part number Description Details 901843 Axiom® Genome-Wide CHB 1 & CHB 2 Array Set Bundle Sufficient for processing 96 gDNA samples, includes: n One Axiom Genome-Wide CHB 1 96-Array Plate n One Axiom Genome-Wide CHB 2 96-Array Plate n Two Axiom® 2.0 Reagent Kits n Two Axiom® GeneTitan® Consumables Kits 901764 Axiom® Genome-Wide CHB 1 Array Plate Contains one Axiom Genome-Wide CHB 1 96-Array Plate 901842 Axiom® Genome-Wide CHB 2 Array Plate Contains one Axiom Genome-Wide CHB 2 96-Array Plate 901758 Axiom® 2.0 Reagent Kit Contains all reagents (except isopropanol) required to process 96 gDNA samples 901606 Axiom® GeneTitan® Consumables Kit Contains all GeneTitan® Instrument consumables required to process one Axiom® Array Plate 0000740 Axiom® Genotyping Services Sample processing and data generation services for gDNA samples (minimum 1,000 samples) using the Axiom Genotyping Solution www.affymetrix.com Please visit our website for international distributor contact information. “For Research Use Only. Not for use in diagnostic procedures.” P/N DNA01393 Rev. 1 ©Affymetrix, Inc. All rights reserved. Affymetrix® , Axiom® , Command Console® , CytoScan™ , DMET™ , GeneAtlas® , GeneChip® , GeneChip-compatible™ , GeneTitan® , Genotyping Console™ , myDesign™ , NetAffx® , OncoScan™ , Powered by Affymetrix™ , PrimeView™ , Procarta® , and QuantiGene® are trademarks or registered trademarks of Affymetrix, Inc. All other trademarks are the property of their respective owners. Products may be covered by one or more of the following patents: U.S. Patent Nos. 5,445,934; 5,744,305; 5,945,334; 6,140,044; 6,399,365; 6,420,169; 6,551,817; 6,733,977; 7,629,164; 7,790,389 and D430,024 and other U.S. or foreign patents. Products are manufactured and sold under license from OGT under 5,700,637 and 6,054,270. Affymetrix, Inc. Tel: +1-888-362-2447 Affymetrix UK Ltd. Tel: +44-(0)-1628-552550 Affymetrix Japan K.K. Tel: +81-(0)3-6430-4020 Panomics Products Tel: +1-877-PANOMICS www.panomics.com USB Products Tel: +1-800-321-9322 www.usb.affymetrix.com Quick assay turnaround time Axiom Genome-Wide CHB 1 & CHB 2 Array Plate Set is part of Axiom® Genotyping Solution, which uses Axiom® 2.0 Assay. The assay is fully automatable on Affymetrix GeneTitan® Multi- Channel Instrument for array processing. n Minimal operator intervention – load consumables, press start, then view analysis n High throughput – instrument operates overnight and can manage over 760 samples/week n Time efficient – less than 2.5 hours of hands-on time per 96 samples Flexible data analysis software Integrate Microsoft Windows GUI-based Genotyping Console™ Software (GTC) or automation-friendly command line-based Affymetrix Power Tools for completing the primary genotyping analysis. The flexible software workflow with simplified data management allows you to easily share your results and seamlessly integrate with third-party software packages. Sample types supported n gDNA from fresh blood n gDNA from saliva (collected using Oragene® DNA collection kits from DNA Genotek) n Whole-genome amplified (WGA) DNA (amplified from gDNA using QIAGEN Repli-g® kits)