This white paper describes SABiosciences' Oligo GEArray DNA microarray system. Key points:
1) Oligo GEArrays profile gene expression of focused gene panels (113-440 genes) related to biological pathways or disease states, providing a more economical alternative to genome-wide microarrays.
2) The system uses flexible target labeling kits, membrane arrays, and a simple chemiluminescent detection method, allowing it to be easily adopted in any laboratory.
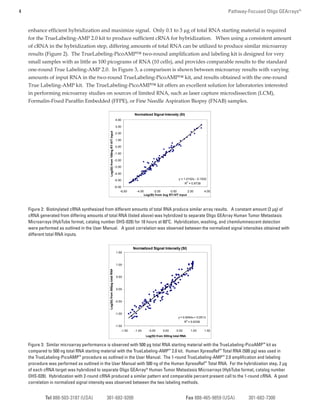
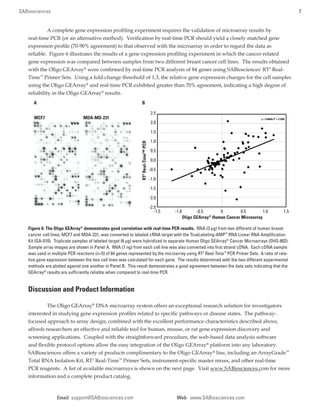
3) Data shows the Oligo GEArray has high sensitivity (can detect 10 fM targets), a wide dynamic range, and is highly reproducible, with technical replicates yielding CVs less than 10%.