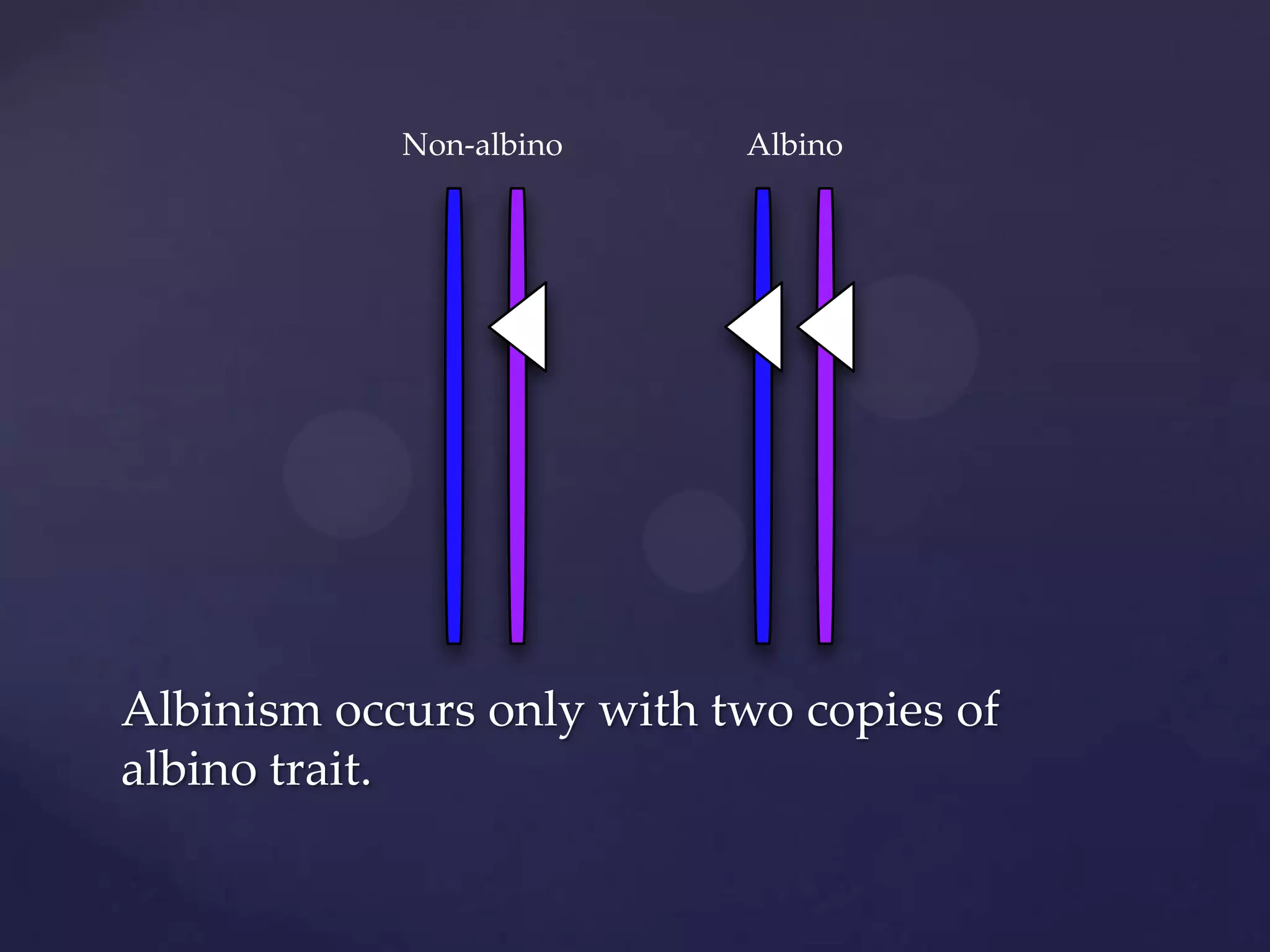
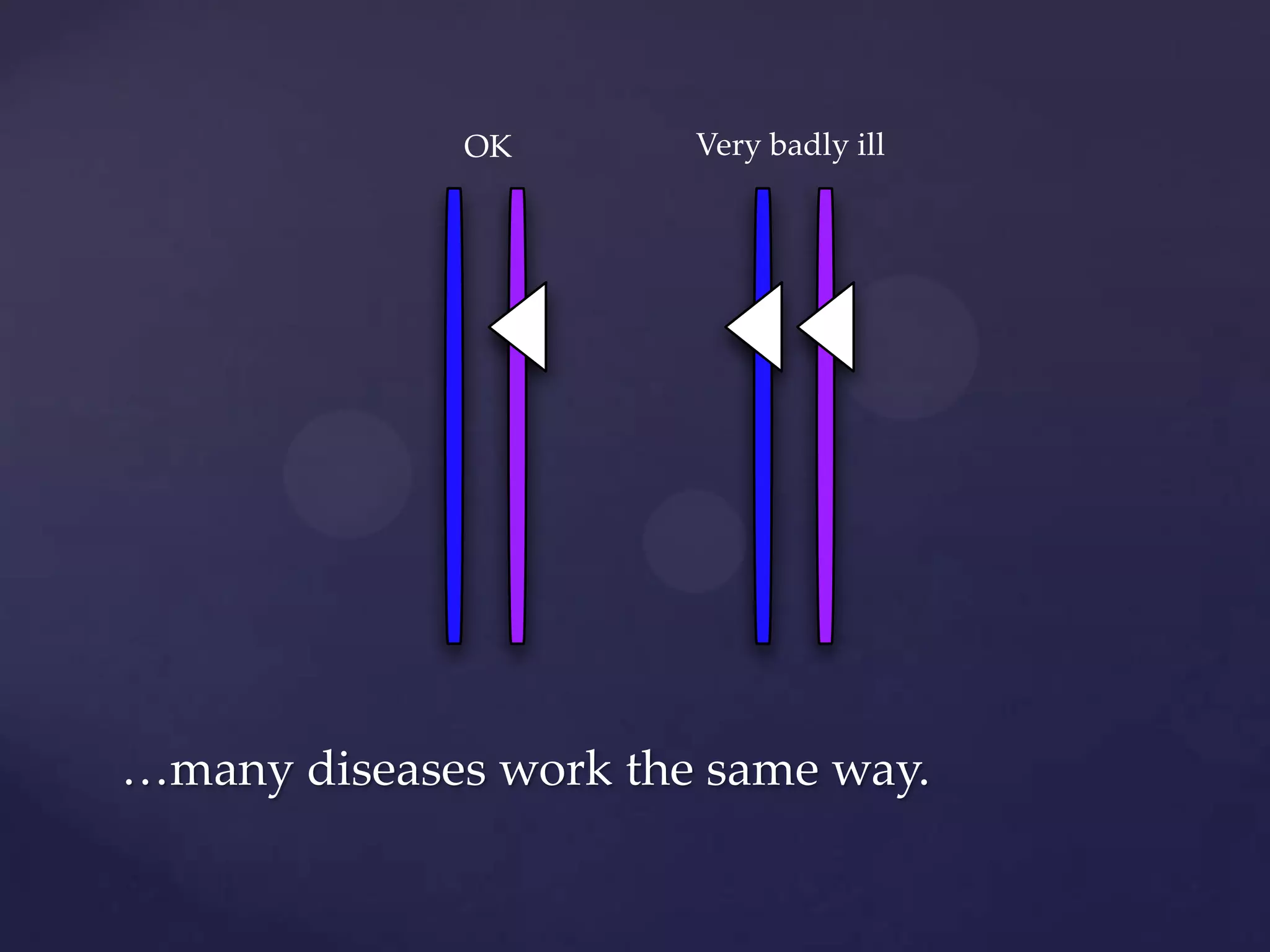
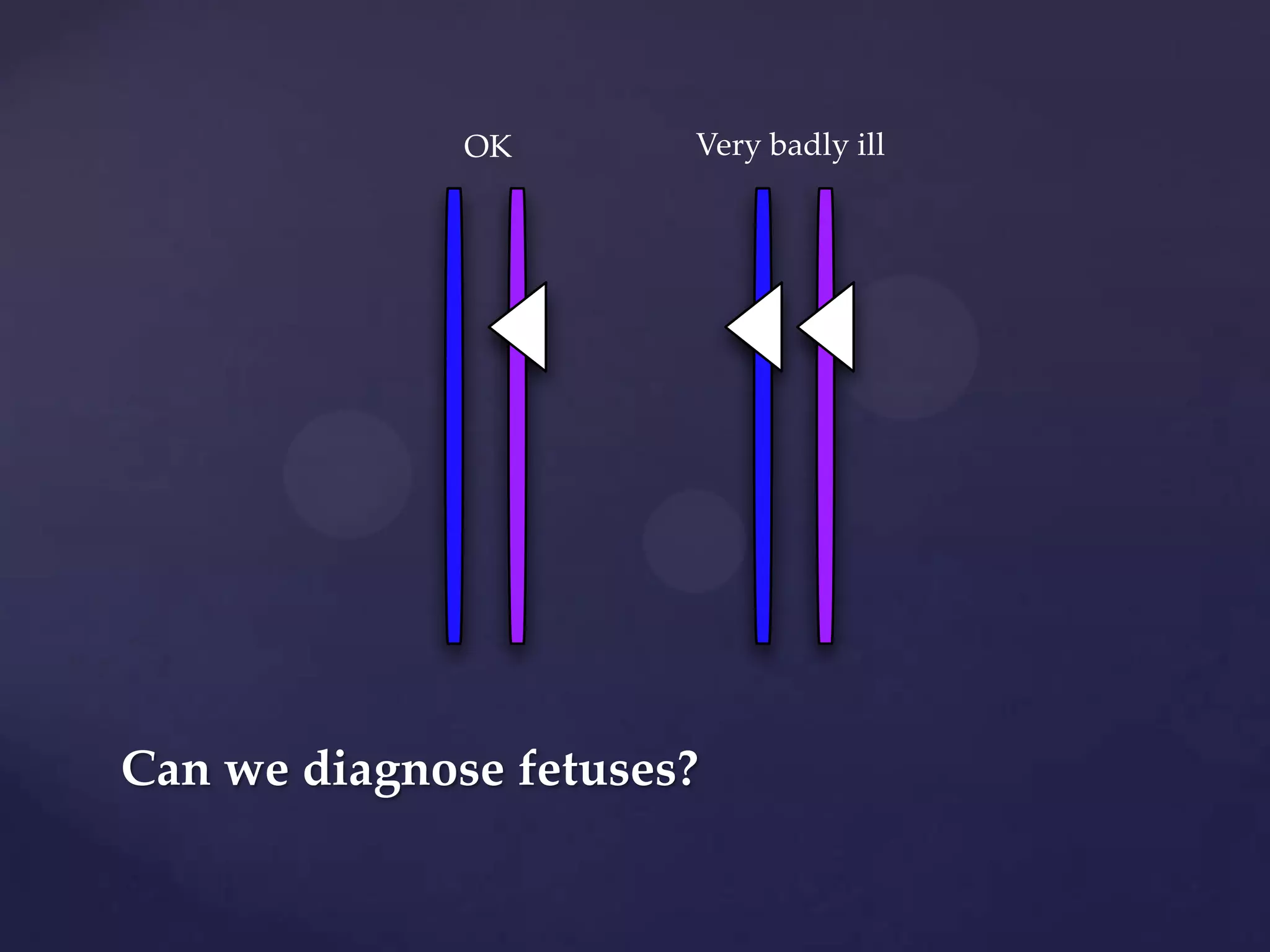
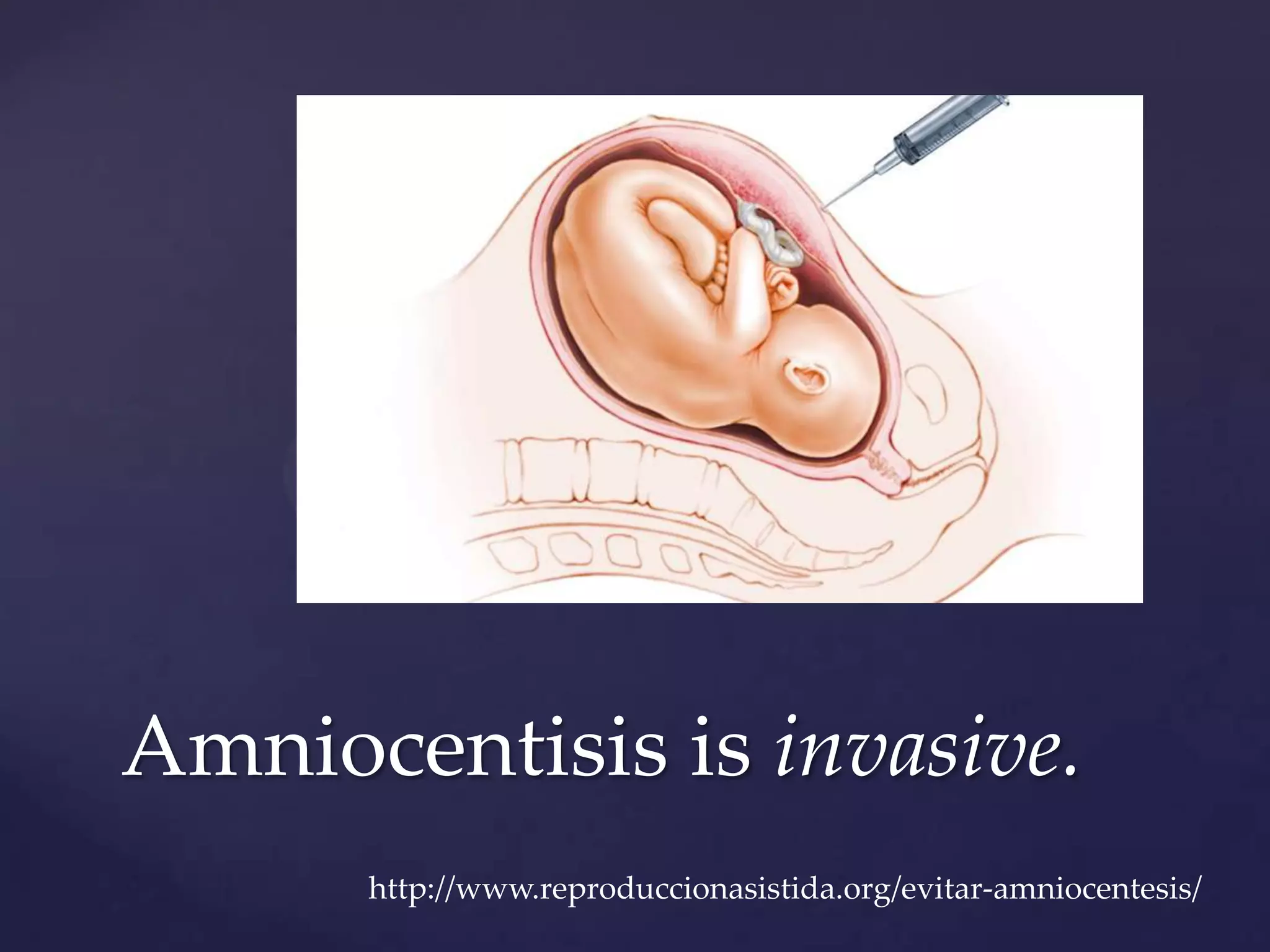
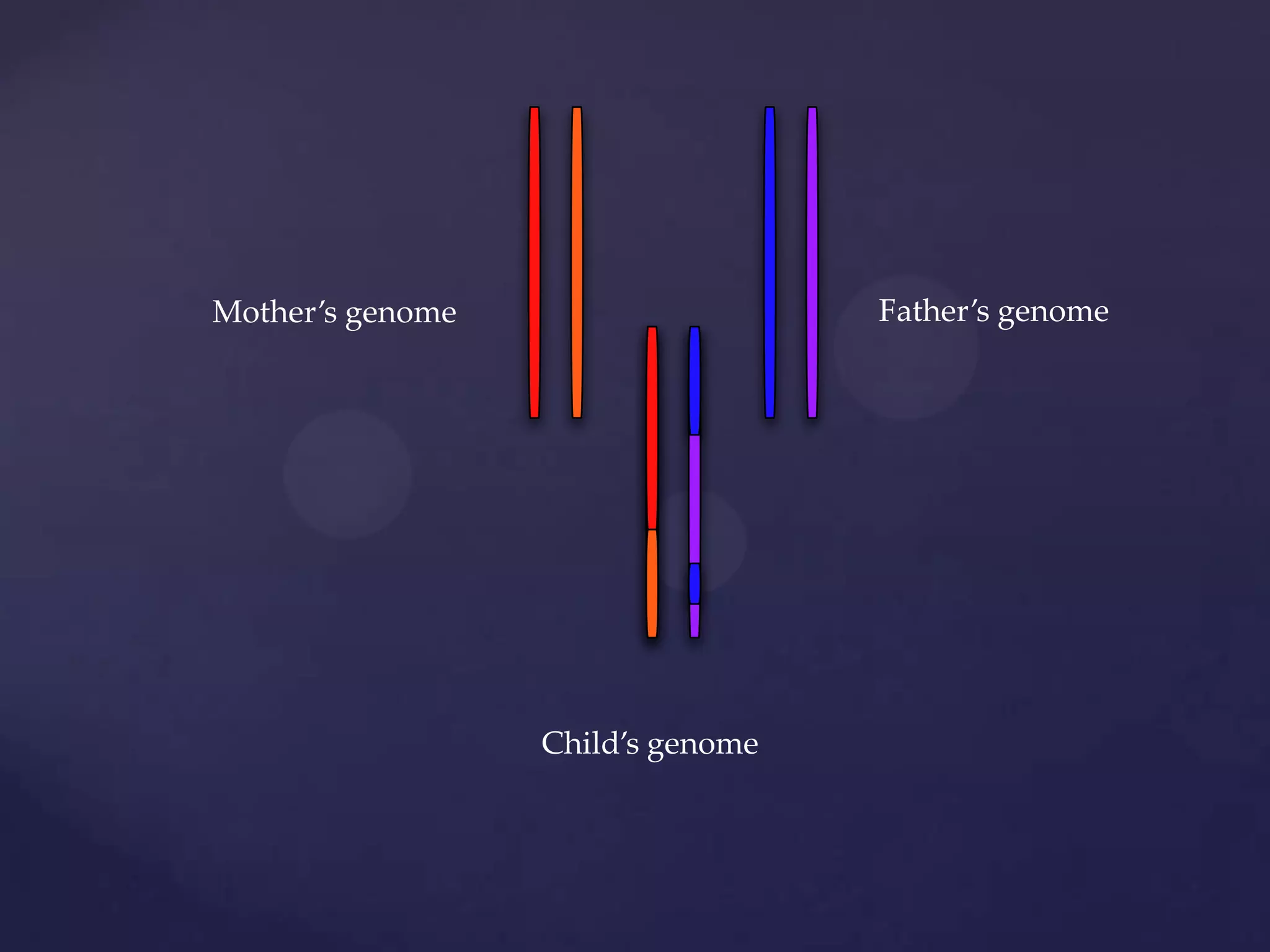
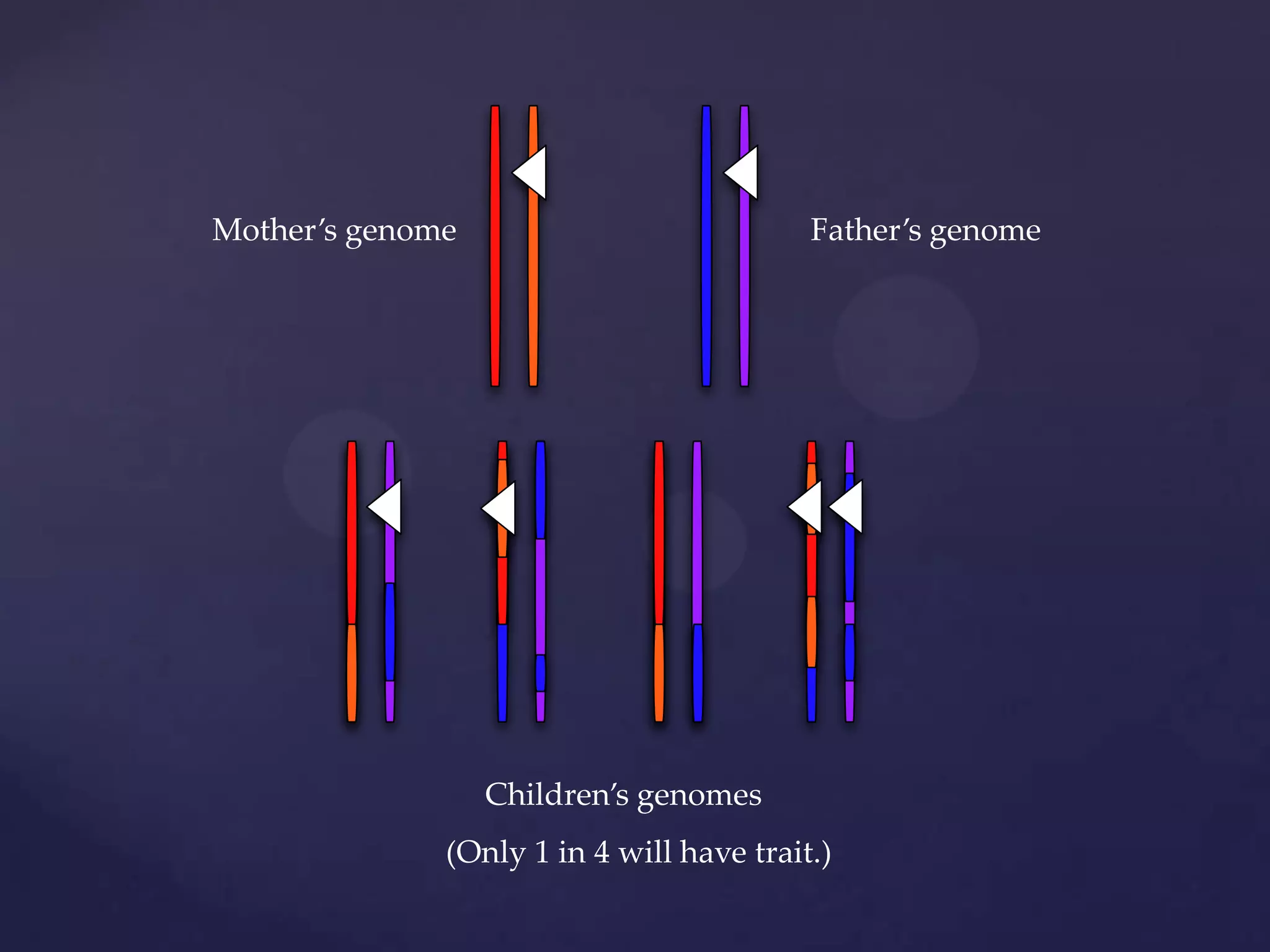
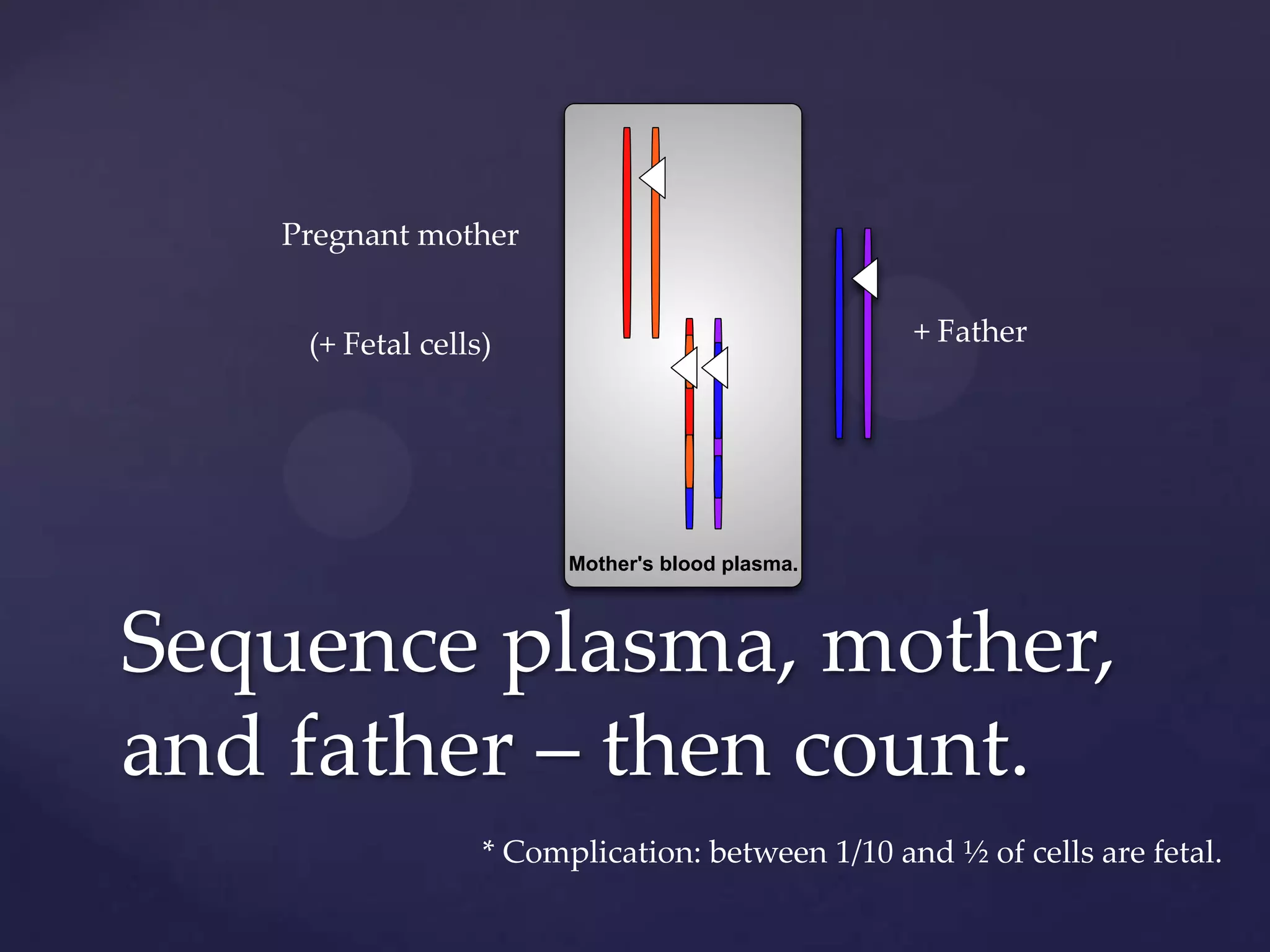
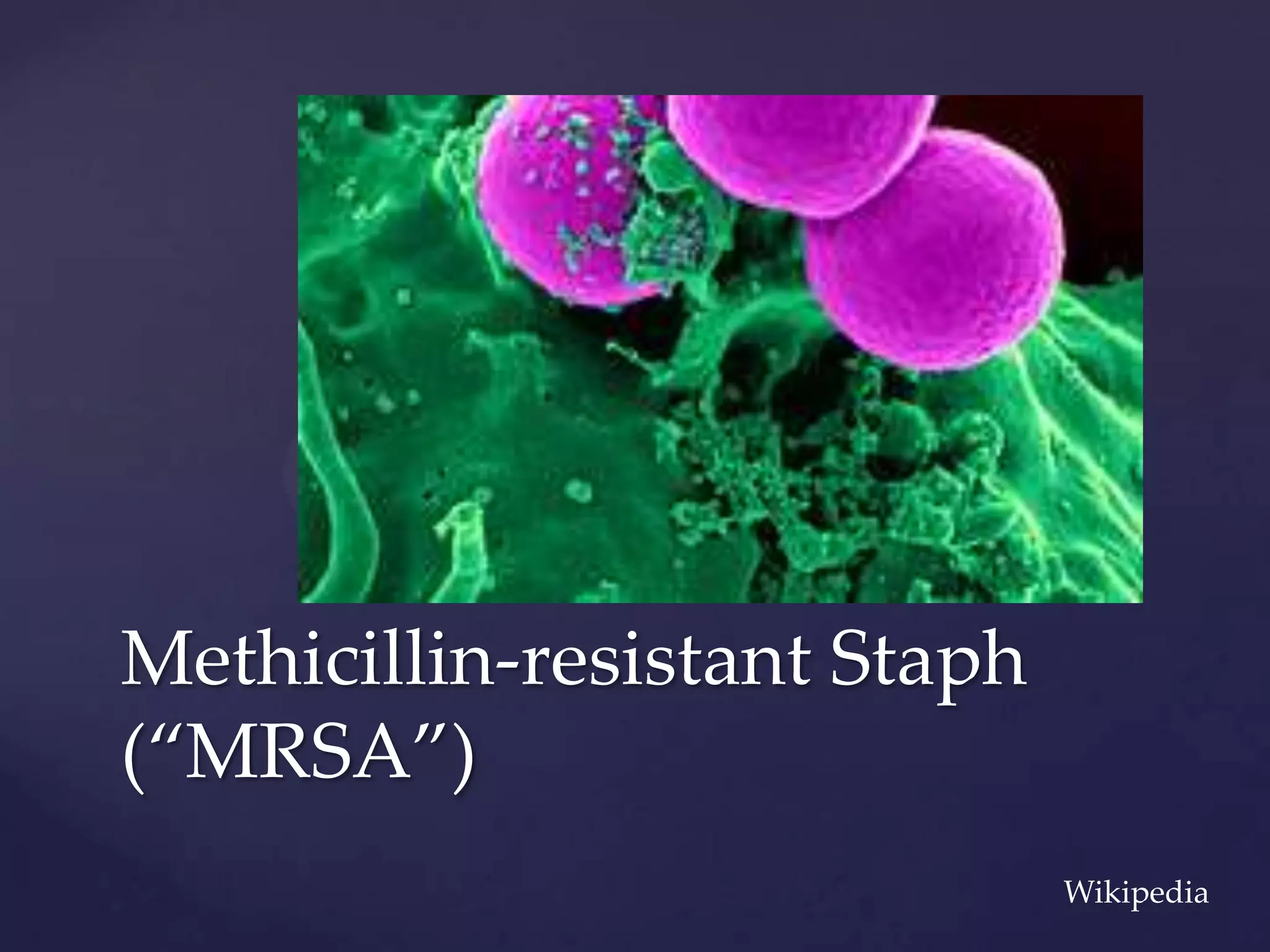
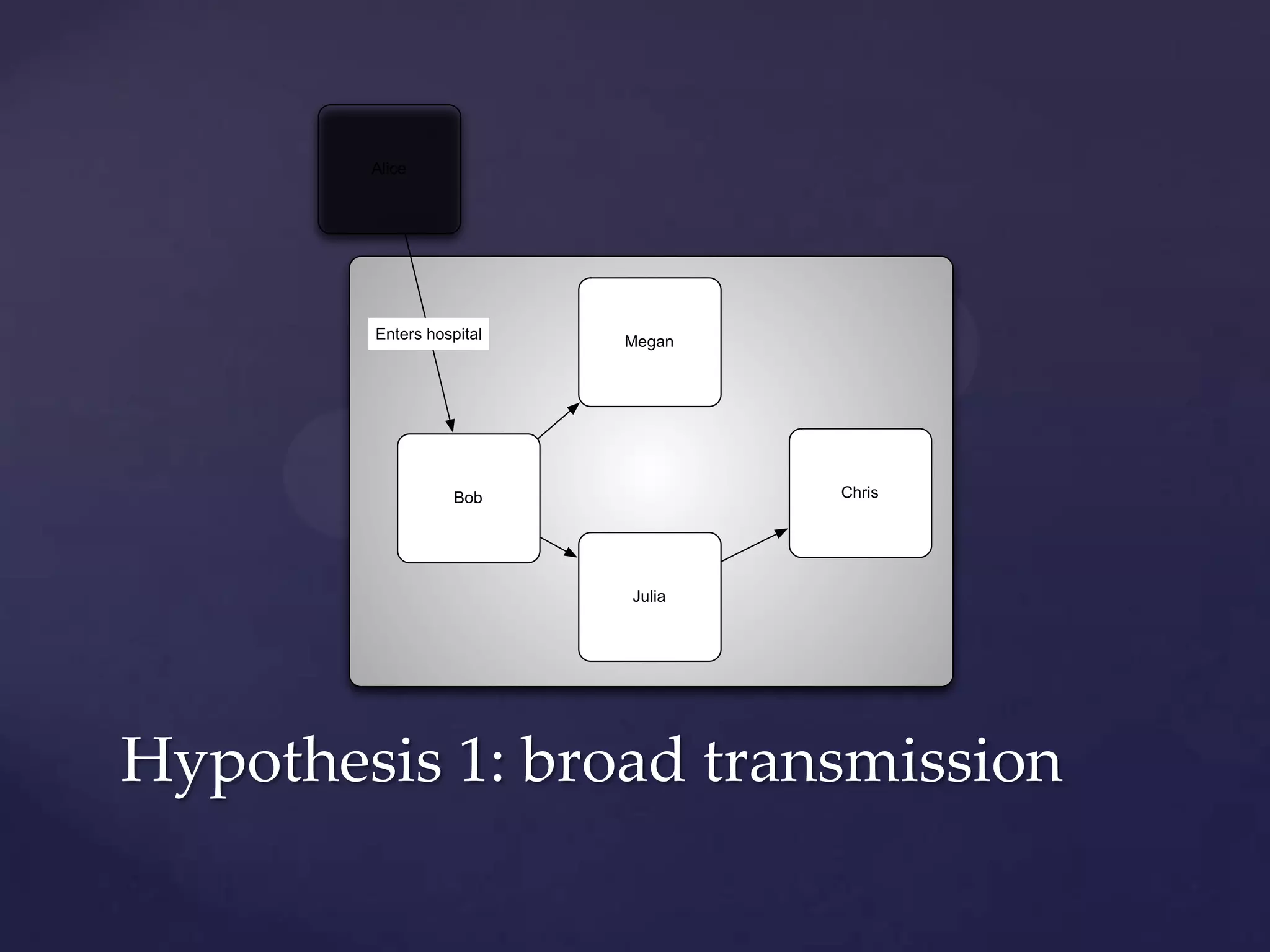
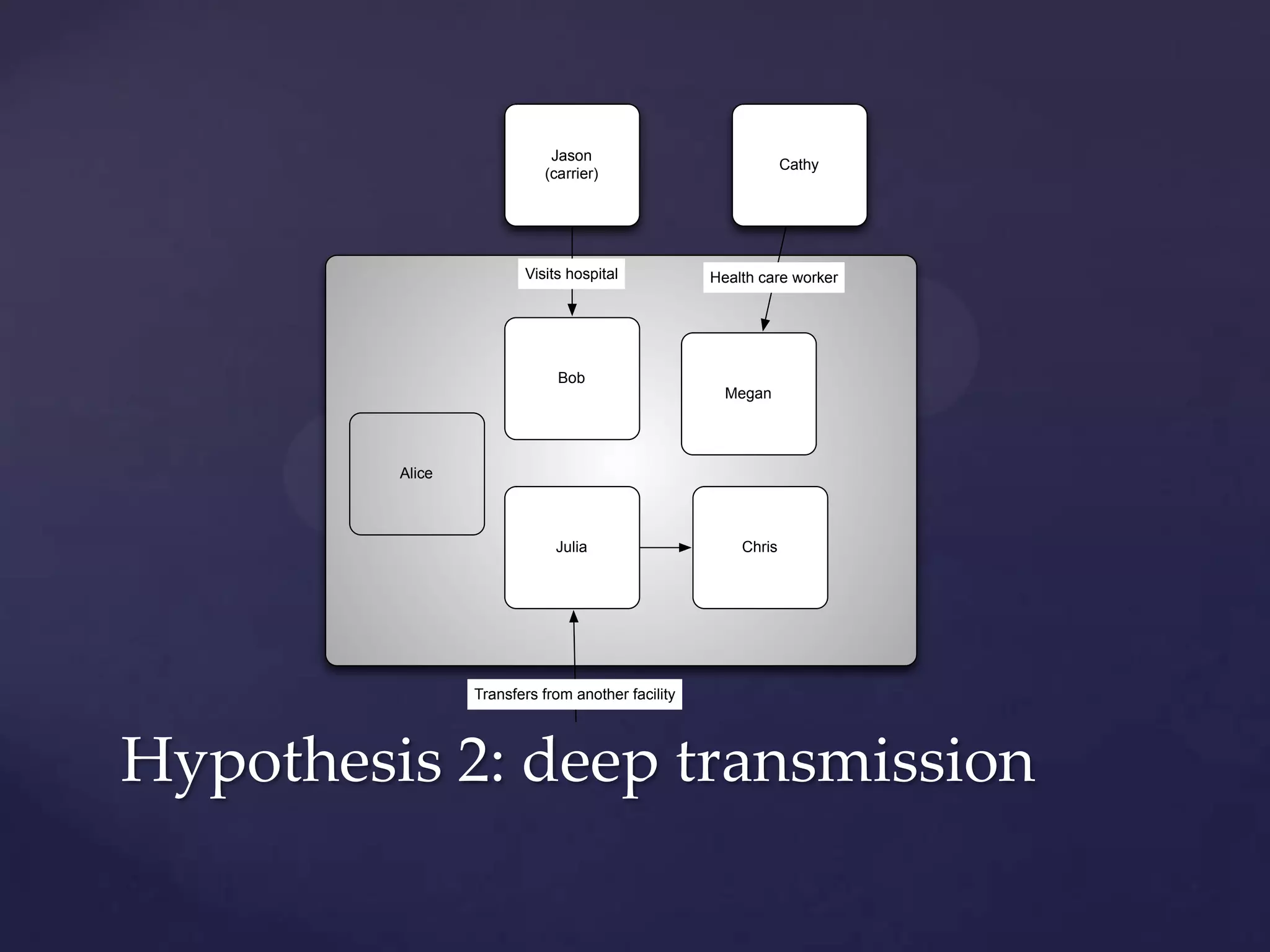
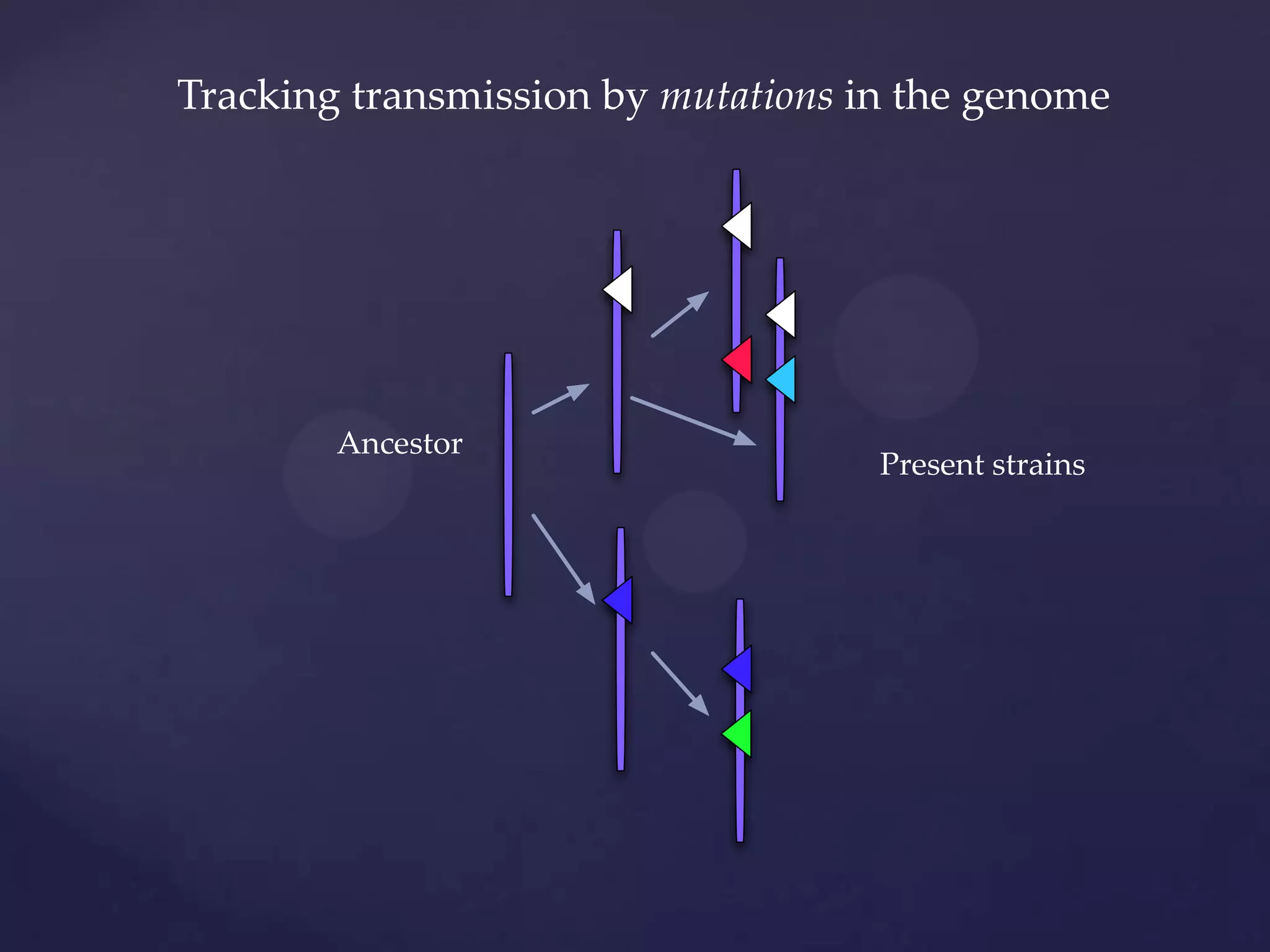
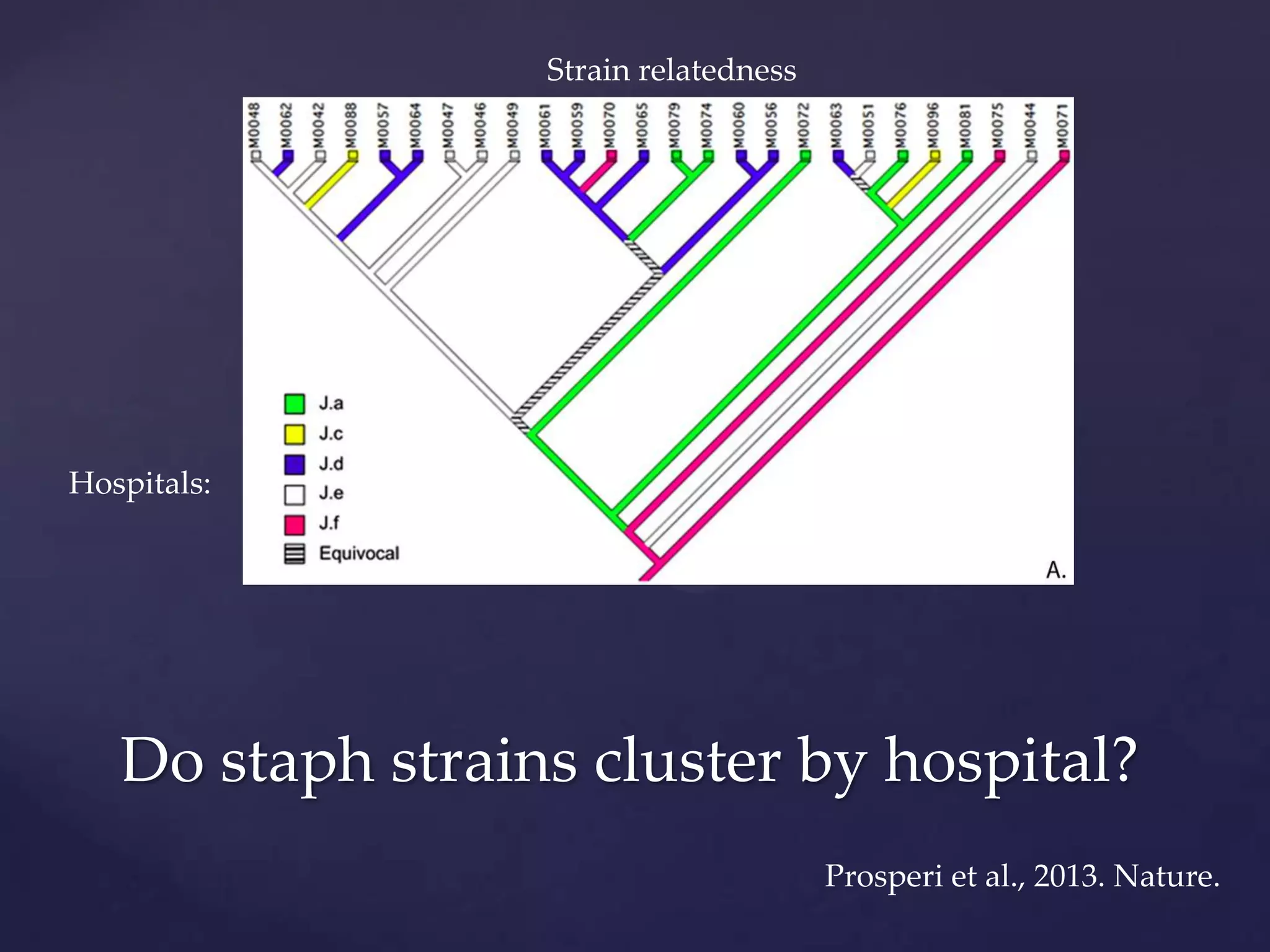
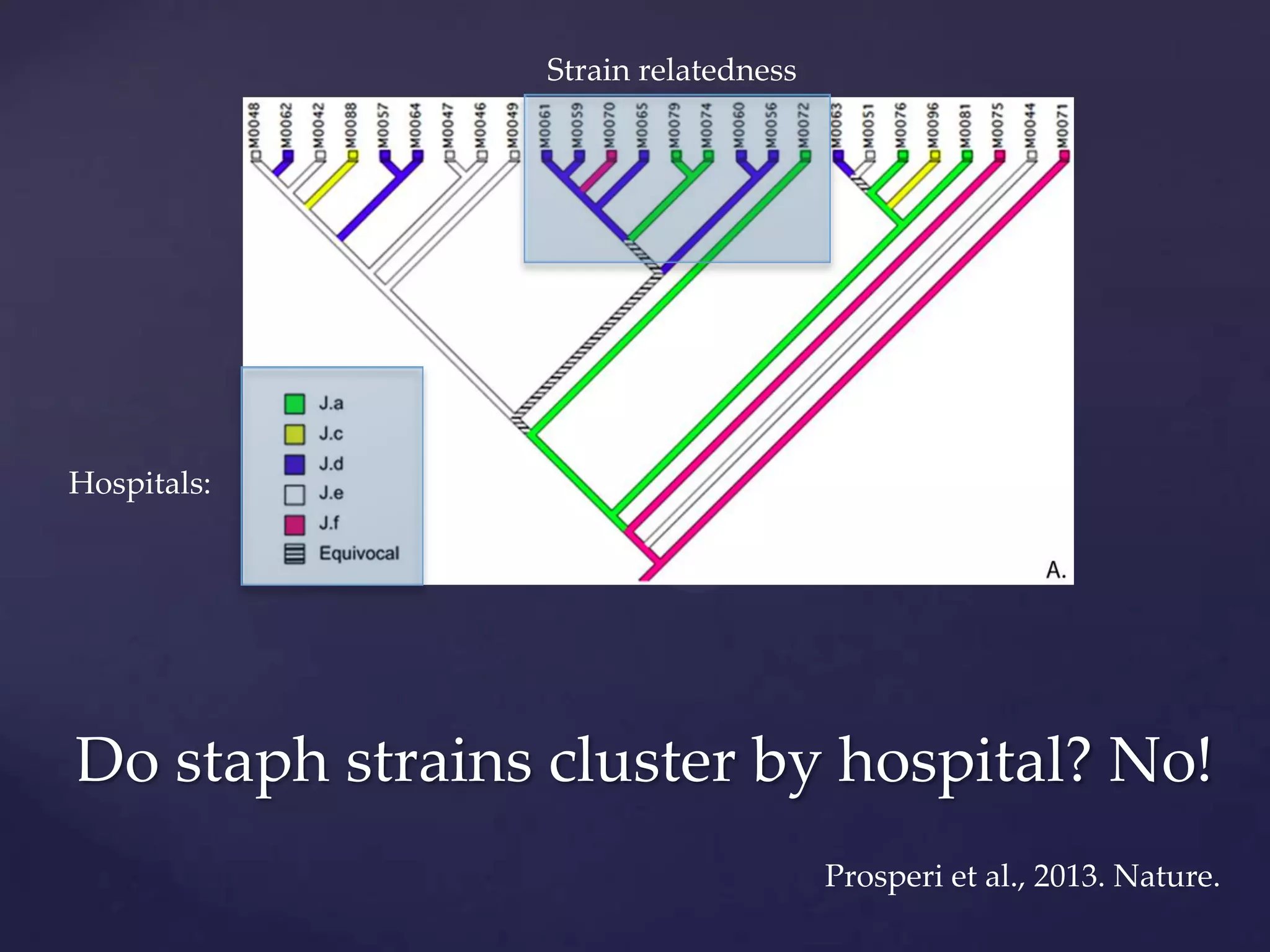
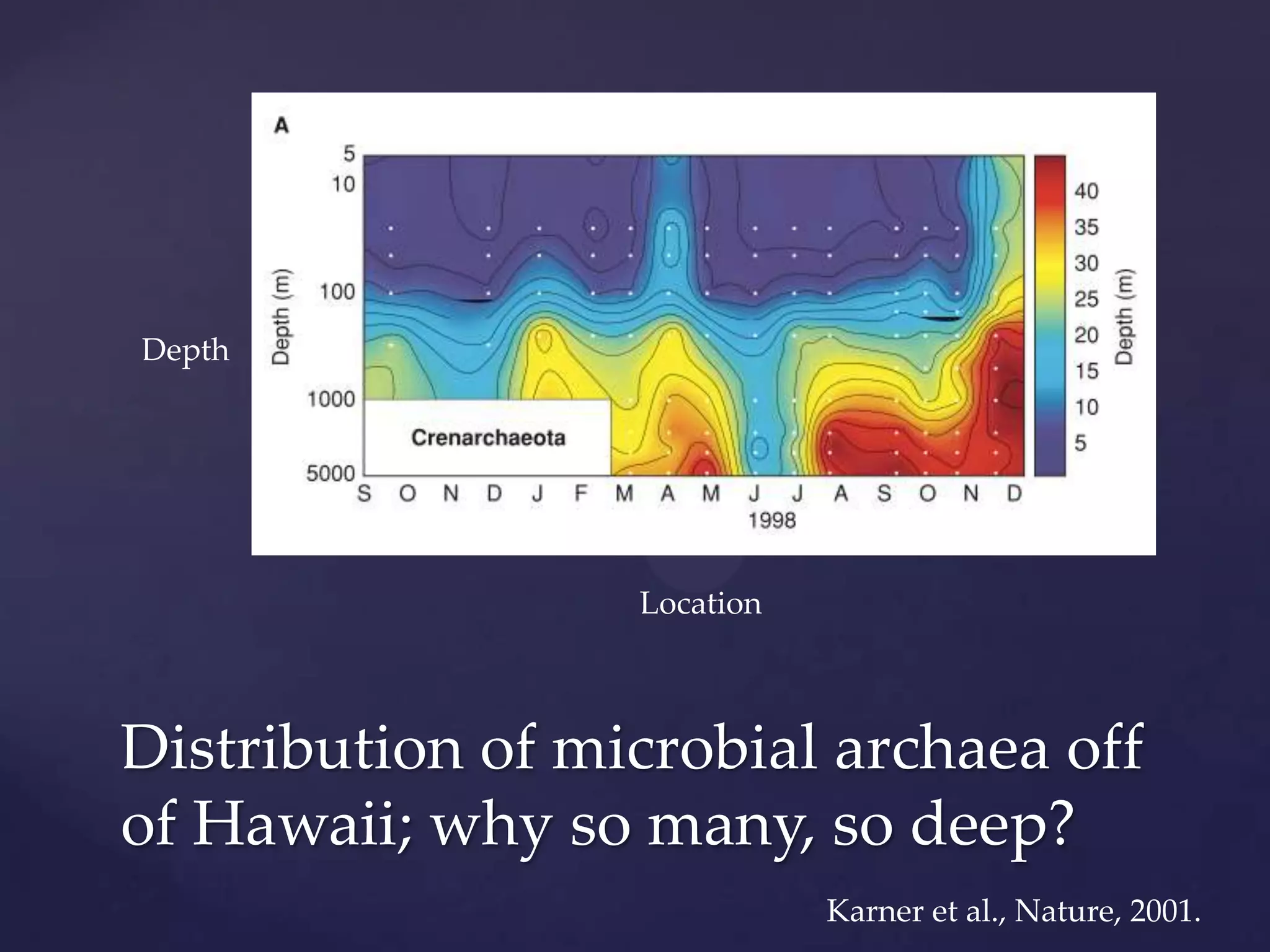
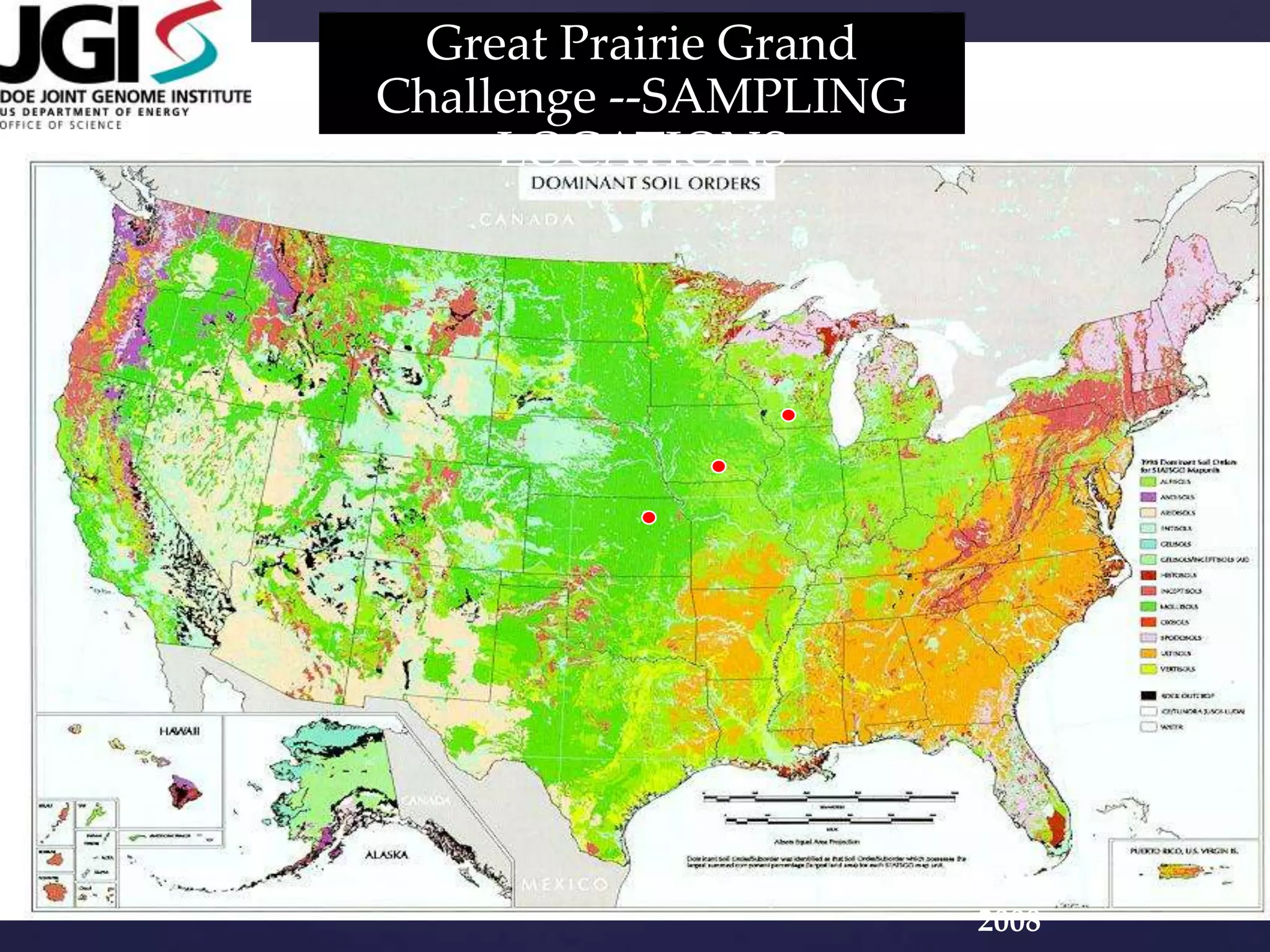
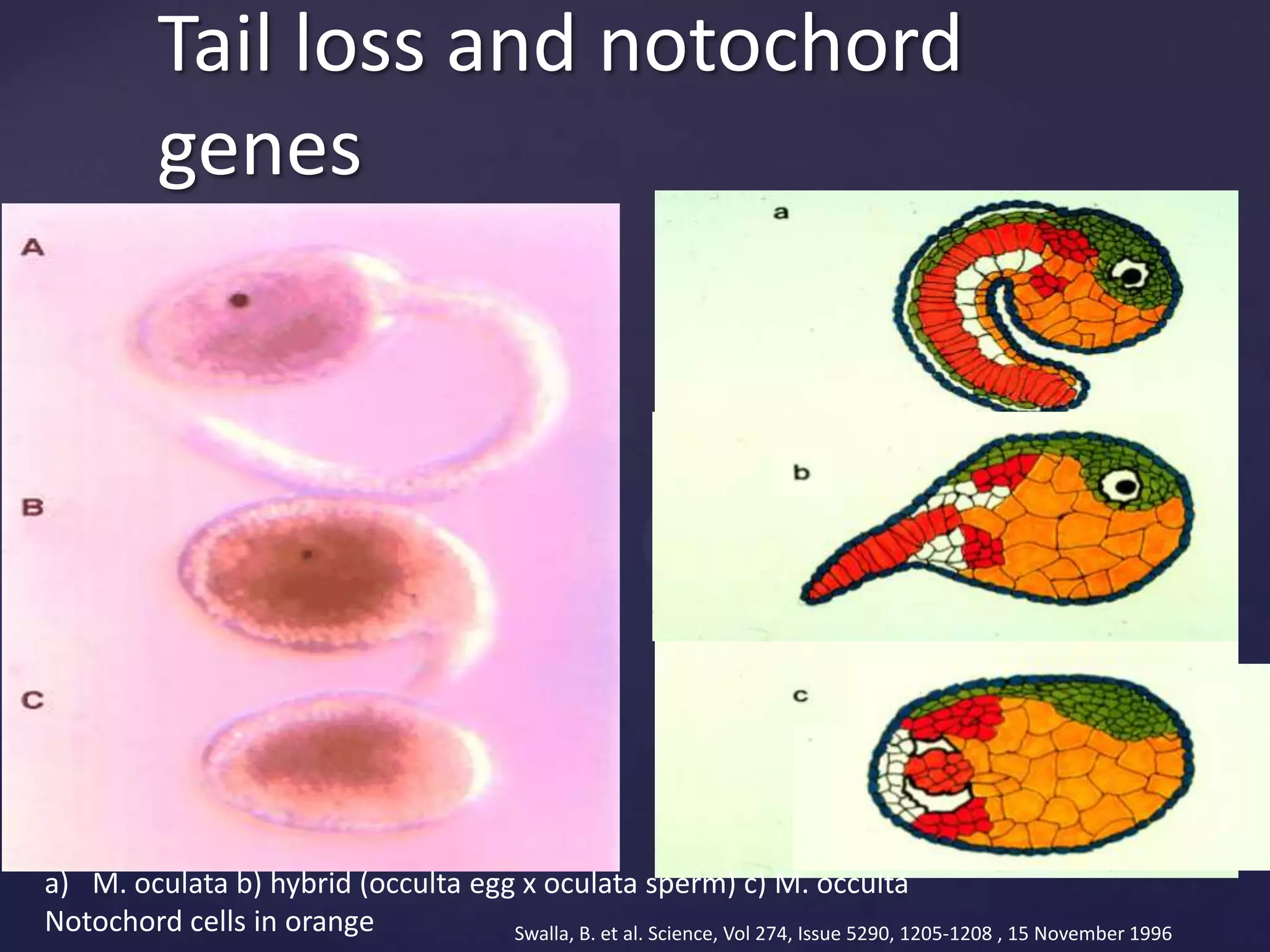
The document discusses how genomic sequencing is changing scientific research by providing three examples of how genome sequencing was used: to diagnose genetic conditions in fetuses non-invasively, to track the transmission of hospital-acquired infections, and to investigate global nutrient cycling in oceans by identifying previously unknown microbes. The author is an assistant professor who discusses his background and how cheap genome sequencing allows more genomes to be analyzed to generate data that provides unexpected results and interpretations.